Abstract
More than one-third of the world’s population is exposed to Plasmodium vivax malaria, mainly in Asia1. P. vivax preferentially invades reticulocytes (immature red blood cells)2,3,4. Previous work has identified 11 parasite proteins involved in reticulocyte invasion, including erythrocyte binding protein 2 (ref. 5) and the reticulocyte-binding proteins (PvRBPs)6,7,8,9,10. PvRBP2b binds to the transferrin receptor CD71 (ref. 11), which is selectively expressed on immature reticulocytes12. Here, we identified CD98 heavy chain (CD98), a heteromeric amino acid transporter from the SLC3 family (also known as SLCA2), as a reticulocyte-specific receptor for the PvRBP2a parasite ligand using mass spectrometry, flow cytometry, biochemical and parasite invasion assays. We characterized the expression level of CD98 at the surface of immature reticulocytes (CD71+) and identified an interaction between CD98 and PvRBP2a expressed at the merozoite surface. Our results identify CD98 as an additional host membrane protein, besides CD71, that is directly associated with P. vivax reticulocyte tropism. These findings highlight the potential of using PvRBP2a as a vaccine target against P. vivax malaria.
Similar content being viewed by others
Main
The development of malaria infection depends on the efficiency with which the invasive forms, merozoites, attach to and invade erythrocytes. Red blood cell (RBC) selection is not random, as the tropism towards erythrocytes of different ages varies with Plasmodium species and restrict zoonotic infections13,14. Furthermore, merozoite invasion occurs within minutes, and while free in the blood stream, these forms are particularly vulnerable to immune attack15. To take advantage of this brief window of opportunity, targeting the molecules implicated in RBC selection and invasion is attractive for vaccine development. This has prompted sustained efforts to unravel the complex sequence of events leading to erythrocyte invasion and to identify the parasite and host proteins that mediate them.
Knowledge of erythrocyte invasion has been mainly accrued for Plasmodium falciparum, a parasite that can invade all erythrocyte subsets, through detailed and elegant investigations made possible by the ability to culture this species in vitro and to generate genetic variants. This has led to the discovery of multiple combinations of parasite ligands and host receptors that mediate distinct invasion pathways, with the most recent PfRh5–basigin combination providing a particularly promising target for vaccine development16,17. In contrast, invasion by P. vivax, a species long known to have a strict predilection towards reticulocytes2,3,4, cannot be adequately continuously maintained in vitro and has consequently been far less studied. We have previously shown that the P. vivax merozoite preferentially targets reticulocytes expressing the immature reticulocyte marker CD71 (ref. 4). Gruszczyk et al.11 then identified CD71 (transferrin receptor 1) as a reticulocyte receptor recognized by the merozoite protein PvRBP2b. However, previous studies of different P. vivax strains from Brazil18 and Thailand19 have shown that merozoite invasion is trypsin-resistant. CD71 is trypsin sensitive (Fig. 1a), which suggests the possibility of an alternative receptor on reticulocytes for P. vivax merozoites. This hypothesis was recently supported by a study20 demonstrating that merozoite invasion could not be prevented by antibodies against CD71.
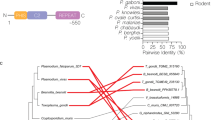
a, Schematic depicting the presence of an unknown P. vivax receptor involved in reticulocyte tropism (reticulocyte-specific receptor (RSR)). b, Comparison of protein abundance (expressed as relative fold-change) between CD71+ reticulocytes and CD71– erythrocytes measured by MS (Supplementary Table 1). The “R” in blue denotes trypsin resistance (see also Extended Data Fig. 2). c, Flow cytometry profile of CD98 and CD71 expression on TO–, TO-low and TO-high erythrocytes for three different human cord blood samples. The TO-high subset represents the most immature reticulocyte population compared with the TO-low subset (https://www.bdbiosciences.com/ds/is/tds/23-1789.pdf). d, CD98 and Demantin (Band 4.9) expression in CD71– and CD71+ erythrocytes (representative images are shown from an analysis consisting of 3 independent experiments, each time inspecting 40–50 cells), as recorded by immunofluorescence microscopy. Scale bar, 2 μm. e, Detection of CD98 by western blotting of ghost membranes derived from CD71– and CD71+ erythrocytes matured ex vivo for 20 and 40 h. Band 3 anion exchange protein served as the loading control for western blots. Representative images are shown, and the pattern of CD98 expression was consistent as observed from a total of eight independent experiments, using three batches of blood. f, Schematic representation of the CD98 structure (PDB access code: 6IRS) with the 4F2hc heavy-chain ectodomain of CD98 depicted as blue ribbons and its N-linked glycans as black sticks. The membrane-embedded light chain of CD98 (4F2lc) is depicted as yellow ribbons25.
To find potential targets that are differentially expressed by this susceptible cell population, we followed a strategy of a differential proteomics screen of CD71+ (susceptible) versus CD71– (non-susceptible) erythrocyte ghost membranes. Candidate reticulocyte-specific receptors were selected on the basis of the following two criteria: (1) abundant expression on CD71+ immature reticulocytes but not on CD71– normocytes and (2) resistance to trypsin treatment. The proteomics screen identified a number of proteins with increased expression on CD71+ reticulocytes21 (Fig. 1b), of which a subset was resistant to trypsin treatment as demonstrated by flow cytometry (Extended Data Fig. 1 and Supplementary Table 1). Two surface proteins displayed a significant fold-change reduction in their expression levels between CD71+ and CD71– erythrocytes, namely CD71 (~6-fold) and CD98 (~4-fold), while all others showed less than a 2-fold change (Fig. 1b). Of these two proteins, only CD98 proved resistant to trypsin treatment (Extended Data Fig. 1). The decrease in CD98 expression during cord blood erythrocyte maturation from CD71+ to CD71– erythrocytes was confirmed by careful immunophenotyping (Fig. 1c). The population of CD71+CD98+ reticulocytes represents a minor fraction of RBCs and reticulocytes in adult peripheral blood (Extended Data Fig. 2a). A decrease in CD98 expression during erythrocyte maturation was also confirmed by immunofluorescence (Fig. 1d) and western blotting (Fig. 1e) analyses of in vitro-matured CD71+ reticulocytes. CD98 (also known as SLC3A2 or 4F2hc) is the heavy chain component of the large neutral amino acid transporter complex22,23 that is peripherally expressed on the reticulocyte membrane24,25,26,27 (Fig. 1f).
To demonstrate the involvement of CD98 in the invasion of reticulocytes, we conducted invasion inhibition assays using different P. vivax clinical isolates28. Anti-CD98 antibodies significantly abrogated the invasion of P. vivax merozoites into CD71+ reticulocytes, with high levels of inhibition (~70%; Fig. 2). Conversely, antibodies directed against three other erythrocyte membrane proteins, CD240DCE (Rh), CD99 and CD147 (basigin), a known receptor of P. falciparum merozoite invasion16, had little to no effect on invasion efficiency (Fig. 2). All of the P. vivax clinical isolates from Thailand used in this study and previous studies10,28 were DARC dependent (>95% inhibition of invasion; Fig. 2). It is interesting to note that in the patient samples, CD98 could not be detected at the surface of the P. vivax-infected reticulocytes at the ring stage. This suggests that CD98 expression on the cell surface is lost soon after invasion of reticulocytes by P. vivax (Extended Data Fig. 2b,c), a phenomenon also seen for CD71 (ref. 4).
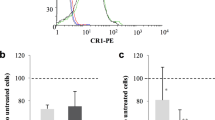
P. vivax invasion inhibition assay in the presence of anti-CD240DCE (monoclonal), anti-CD99 (monoclonal), anti-CD147 (monoclonal), anti-CD98 (polyclonal) and anti-CD98 (monoclonal) F(ab′)2 antibodies (final concentration of 25 μg ml–1). The anti-DARC antibody (monoclonal) was used as an inhibition positive-control, and a total of eight P. vivax clinical isolates were used. Inhibition values are represented as the mean ± s.d., Kruskal–Wallis test (P < 0.0001). NS, not significant.
To identify the corresponding parasite ligand that binds CD98, we generated a small pDisplay surface-expression library of selected P. vivax genes, such as the RBPs, which are potentially implicated in merozoite invasion (Extended Data Fig. 3a–c and Supplementary Table 2). The P. vivax antigen library, cloned in the pDisplay vector, consisted of 28 different constructs encoding for 7 different P. vivax proteins that were successfully transfected into HEK293 cells for cell surface expression. Binding assays using these cells with CD71+ immature versus CD71– mature erythrocytes identified fragments encoded by two genes, PvRBP2a (Fig. 3) and PvRBP2b (Supplementary Table 2), that mediated strong binding to immature reticulocytes. To determine which of these were the cognate target for CD98, anti-CD98 antibodies were used to block their binding. PvRBP2a binding was blocked by anti-CD98 antibodies (Fig. 3b,c). By contrast, the same anti-CD98 antibodies could not block PvRBP2b binding (Extended Data Fig. 3e), which was expected since PvRBP2b has been shown to bind to CD71 (ref. 11). PvRBP2a was also capable of binding to normocytes, but this interaction was not blocked by anti-CD98 antibodies (Extended Data Fig. 3f). These observations also indicate that the PvRBP2a protein can bind additional proteins on the surface of normocytes.
a, Binding of reticulocytes to 1,000 HEK293 cells transfected (efficiency > 20%) with different gene or gene fragment from P. vivax. Binding was assessed for HEK293 cells expressing a fragment of Duffy binding protein II (DBP) known to contain an erythrocyte-binding domain40 or different fragments of PvRPB2a (PvRBP2a23–767, PvRBP2a448–1090, PvRBP2a448–1314, PvRBP2a765–1314, PvRBP2a1315–1650 or PvRBP2a1539–1920), or MAEBL60–587. Specific binding numbers to transfected HEK293 cells were determined after subtraction of the number of reticulocytes or normocytes binding to non-transfected HEK293 cells (screening was done in duplicate). b, Binding assays of reticulocytes to HEK293 cells expressing PvRBP2a23–767 in the presence or absence of isotype control or anti-CD98 antibodies (concentrations). The binding of normocytes to HEK293 cells expressing PvRBP2a23–767 was specific since the addition of antibodies against CD98 did not abrogate binding to normocytes (Extended Data Fig. 3c). Values are expressed as the mean ± s.d., ANOVA followed by Dunnett post-test. c, Binding assays with reticulocytes in the presence of PvRBP2a23–767 fragment and antibodies against RBP2a (1C3 and 3A11 at 25 μg ml–1). Values are expressed as the mean ± s.d., ANOVA followed by Dunnett post-test. d, Inhibition of P. vivax invasion using two different mouse monoclonal anti-PvRBP2a (1C3 and 3A11) antibodies and one anti-DARC antibody with four different isolates tested in parallel. The inhibition was normalized to invasion efficiencies obtained without antibodies. Values are expressed as the mean ± s.d., Kruskal–Wallis test (P < 0.0001).
The highest level of binding with reticulocytes, more than twofold than that measured for the other constructs, was observed with HEK293 cells displaying PvRBP2a23–767. HEK293 cells transfected with constructs corresponding to other regions of PvRBP2a showed weaker but still significant binding (Fig. 3a, Extended Data Fig. 3b and Supplementary Table 2). Binding of cells transfected with the PvRBP2a23–767 construct to reticulocytes was abolished by the presence of two different clones of anti-PvRBP2a monoclonal antibodies (Fig. 3c and Extended Data Fig. 4). Inhibition of reticulocyte binding to HEK293 cells expressing the constructs at their surface by anti-PvRBP2a antibodies was stronger than the inhibition observed with the anti-CD98 antibodies that target the reticulocyte surface. This might be due to differences in the affinities of the antibodies used, thereby leading to variable inhibitory efficiency. In addition, the exact regions of interaction between PvRBP2a and CD98 are still unknown. Thus, the different antibodies, despite their inhibitory effect, may not directly recognize the actual sites of interactions and thus provide optimal inhibition. Notably, one of these two antibodies also significantly inhibited the invasion of reticulocytes by P. vivax isolates (Fig. 3d). The moderate levels of inhibition observed could be due to genetic polymorphisms of PvRBP2a across P. vivax isolates (Supplementary Table 3) and/or low affinity of the two anti-PvRBP2a clones against the native protein conformation(s).
To assess whether PvRBP2a and CD98 are direct binding partners, biochemical analyses were carried out. Immunoprecipitation of recombinant PvRBP2a160–1135 was able to co-immunoprecipitate recombinant monomeric CD98 protein, which indicates that they are direct binding partners (Fig. 4a). Similarly, direct binding interactions were observed between PvRBP2a and CD98 by both ELISA and biolayer interferometry (Fig. 4a–d). We found that monomeric CD98 interacted with PvRBP2a with a dissociation constant (KD) value of 12.3 ± 1.5 nM at 37 °C by ELISA using the method outlined in Friguet et al.29 (Fig. 4c,d) and 5.9 ± 1.5 nM at 37 °C by biolayer interferometry. Notably, this interaction is stronger than the first binding event of the canonical P. vivax invasion receptor, Duffy-binding protein (PvDBP) (KD of 2.2 μM)30, or the interaction between the P. falciparum invasion receptor PfRh4 and its cognate partner CR1 (KD of 2.9 μM)31 (Supplementary Table 4).
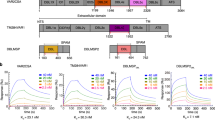
a, Recombinant His-tagged CD98 was incubated with PvBRP2a for 2 h at 37 °C. PvRBP2a was then immunoprecipitated with rabbit anti-PvRBP2a-bound protein A/G magnetic beads. Left: lane 1 shows 75 ng of recombinant PvRBP2a as a positive control, lane 2 shows successful immunoprecipitation (IP) of PvRBP2a. Right: lane 1 shows 75 ng of CD98 as a positive control, lane 2 shows successful immunoprecipitation of CD98 from PvRBP2a mixed with CD98, lanes 3 and 4 are controls using anti-PvRBP2a-bound beads incubated with CD98 without PvRBP2a, or beads alone, respectively. Protein ladder size inferred from the colorimetric images are indicated on chemiluminescent image panels (representative images from three independent experiments). b, Binding of PvRBP2a to CD98 or CD71 proteins coated on an ELISA plate. A total of 25 µg ml–1 PvRBP2a was added to plates pre-coated with 0–6 μg ml–1 of CD98 or CD71. Excess PvRBP2a was washed off, and bound PvRBP2a was detected using a rabbit anti-PvRBP2a antibody, followed by a goat anti-rabbit HRP-conjugated secondary antibody. Background-subtracted optical density values relative to an uncoated control are reported. Error bars indicate the standard deviation from two technical replicates within a representative experiment of two independent experiments. c, PvRBP2a and various concentrations of CD98 ([CD98initial]) were pre-incubated in solution for 2.5 h at 37 °C for binding to reach equilibrium. After the incubation time, the solution was applied to plates pre-coated with CD98 to detect the remaining unbound PvRBP2a concentration ([PvRBP2afinal]). Error bars indicate the standard deviation of three independent experiments. d, Klotz plot of the binding of PvRBP2a to CD98, where the KD was derived from the slope of the best-fit line29. Data from three independent experiments are shown. e, Left: representative binding sensorgram showing the binding of non-biotinylated PvRBP2a to 25 nM biotinylated CD98 in a dose-dependent manner. The binding of PvRBP2a to CD98 was detected using the Octet Red 96e biolayer interferometry system. Biotinylated CD98 was loaded onto the streptavidin (SA) sensor followed by dipping into wells containing the indicated concentration of PvRBP2a. Right: the real-time binding curves are shown as black lines while the red lines indicate the global fits generated using 1:1 Langmuir binding model.
The distinct tropism for different subsets of RBCs by Plasmodium parasites infecting humans as well as other mammalian hosts has held the interest of malariologists for more than 80 years. This was not for mere scientific curiosity, as the erythrocyte niche favoured by the parasite clearly has a major influence on the parasitological course and the clinical outcome of the infection. Restriction to reticulocytes could be considered a natural means to moderate parasite burden and the consequent pathology, allowing both host and parasite survival. Indeed, in humans, clinical severity is less frequent in P. vivax than in P. falciparum malaria, where the parasite invades all erythrocyte subsets. While the reticulocyte restriction has previously been postulated to be due to specific metabolic needs of P. vivax associated with a reticulocyte microenvironment32 and bone marrow localization33, we present compelling evidence that reticulocyte restriction is also caused by invasion receptor tropism. Specifically, CD98 plays an important role as a reticulocyte-specific host receptor, and PvRBP2a was identified as its corresponding parasite ligand. PvRBP2a is a member of a multigene family of 11 members6, some of which, like PvRBP2b, are involved in reticulocyte invasion by P. vivax34,35. Interestingly, PvRBP2a can induce a strong immunoglobulin G response36. It now becomes important to determine whether the PvRBP2a–CD98 interaction is essential for reticulocyte recognition for all P. vivax populations. Indeed, the occurrence of alternative invasion pathways, known for P. falciparum, has recently been raised for P. vivax when infections by this parasite, once considered strictly Duffy-dependent, were recorded in Duffy-negative individuals in Africa and South America37,38,39. PvRBP2a was also shown to bind to other unknown proteins present on normocytes and reticulocytes. Nonetheless, this was not sufficient to drive invasion of the mature RBCs. PvRBP2a–CD98 and PvRBP2b–CD71 can now be considered as two major ligand–receptor pairs implicated in the invasion of reticulocytes by P. vivax. The fact that one is trypsin-resistant and the other not, although apparently contradictory, might denote a combined interaction, a phenomenon of alternative invasion pathways or it might be due to a reliance on distinct receptors by different parasite strains. This can only be addressed by invasion inhibition assays in which identical parasite isolates (which are often polyclonal) are confronted head-to-head with the same sets of anti-CD71, anti-CD98 or anti-DARC antibodies. The current data only allow speculation on the nature of the contributions of CD98 versus that of CD71 in invasion, especially since these datasets relied on different host cell types for the invasion assays. It is interesting to note that indications for an alternative invasion pathway has been recently obtained using P. vivax isolates from India20. This, as well as the identification of other potential associations between PvRBPs and host receptors, merits further study, especially with respect to potential future vaccine development. Finally, the identification of reticulocyte-specific receptor–ligand pairs11 could help the development of continuous in vitro culture of P. vivax, for example, by expressing specific receptors on normocytes allowing invasion; the current lack of such a tool is severely hampering research on this parasite. Ultimately, this provides the community with a vaccine candidate that could contribute to the control and eventual eradication of this globally important pathogen.
Methods
Ethics statement
The clinical samples used in this study were collected from patients infected with P. vivax and from healthy donors (for the cord blood) attending the clinics of the Shoklo Malaria Research Unit (SMRU), Mae Sot, Thailand, under the following ethical guidelines in the approved protocols: OXTREC 45-09 and OXTREC 17–11 (University of Oxford, Centre for Clinical Vaccinology and Tropical Medicine, UK) and MUTM 2008–215 from the Ethics Committee of Faculty of Tropical Medicine, Mahidol University. Human cord blood and adult peripheral blood samples were collected under SingHealth CIRB 2019/2443- and 2017/2806-approved protocols, respectively, and written informed consent was obtained from participants.
CD98 cloning and expression in Sf-9 cells
The coding sequence of the soluble domain of CD98 (W218-A630) was cloned using the primers A 5′-TATCCACCTTTACTGTTAGGCCGCGTAGG-3′ and B 5′-TACTTCCAATCCATGTGGTGGCACACGGGC-3′, and the amplified fragment was purified before insertion into a pFastBac-LIC-BseR1 (Addgene) plasmid using a Gibson Assembly Cloning kit (NEB). Escherichia coli Top10 competent cells were transformed with the ligated plasmid (pFastBac-CD98) and spread on LB plates supplemented with ampicillin. E. coli DH10Bac competent cells were transformed with the purified pFastbac-CD98 plasmid. Extraction of the bacmid and virus packaging was done according to Bac-to-Bac (Invitrogen). For expression, Sf-9 cells were infected at a multiplicity of infection of 1 for 72 h at 27 °C under agitation. Cells were collected at 3,000 × g for 15 min, and the cell pellet was stored at −20 °C.
CD98 purification
Cells were resuspended in 20 mM Tris pH 7.5, 500 mM NaCl and 10 mM imidazole and lysed by sonication. After centrifugation at 20,000 × g for 45 min, the filtered supernatant was injected into a nickel affinity column (HisTrap 1 ml, GE Healthcare). After a washing step of 6% of 20 mM Tris-HCl pH 7.5, 500 mM NaCl and 500 mM imidazole (buffer B), the protein was eluted with 100% of buffer B and injected into a Superdex 75 Hiload 26/60 gel-filtration column (GE Healthcare) equilibrated in PBS. The protein fractions were pooled and concentrated to 19 mg ml–1 before being flash-frozen in liquid nitrogen and stored at −80 °C.
Reticulocyte enrichment
Enrichment of CD71+ reticulocytes was performed using the MACS system (Miltenyi). A total of 1–2 ml of blood at 50% haematocrit in PBS was passed through a LS column. The purity of the positive and negative fractions was monitored by flow cytometry using thiazole orange (TO), which stains RNA (the only nucleic acid species present in reticulocytes) and used for routine clinical quantitation of reticulocytes (https://www.bdbiosciences.com/ds/is/tds/23-1789.pdf). The purity of CD71+ cells was >80%.
Sample preparation and quantitative mass spectrometry
After MACS enrichment, samples were prepared and analysed on a mass spectrometer as described in Chu et al.21. Briefly, after isolation, both reticulocytes and normocytes were lysed using 0.02% saponin in PBS with protease inhibitors. Following this, the membrane and soluble fractions were separated by differential centrifugation and precipitated with ice-cold acetone. For the membrane fraction, the sample was reduced using 5 mM Tris 2-carboxy-ethyl phosphine hydrochloride for 3 h at 30 °C and then alkylated in the dark using 55 mM iodoacetamide for 1 h at room temperature (RT). In-solution digestion of different protein samples with trypsin (Promega) was carried out (at 10 ng ml–1 final concentration) at 37 °C for 16 h. These samples were desalted and then labelled with isobaric tags as described in Chu et al.21, with ~95% labelling efficiency. We labelled CD71– membrane samples with 116 isobaric tags and CD71+ membrane samples with 117 isobaric tags. These samples were further fractionated using a X-Bridge C18 column (Waters; 4.6 × 250 mm, 5 µm, 300 Å). Fractions obtained from this LC step were analysed using an Ultimate 3000 RSLC nano-HPLC system (Dionex) coupled to a QExactive Hybrid Quadrupole-Orbitrap mass spectrometer (Thermo Scientific).
Proteomics analysis
Database searching
All tandem mass spectrometry (MS/MS) samples were analysed using Mascot (Matrix Science, v.2.4.1). Mascot was set up to search the con_uni_human_20150401_20150623 database (unknown version, 180,822 entries) assuming the digestion enzyme trypsin. Mascot was searched with a fragment ion mass tolerance of 0.020 Da and a parent ion tolerance of 10.0 ppm. Carbamidomethyl of cysteine and iTRAQ4plex of lysine and the amino terminus were specified in Mascot as fixed modifications. Deamidated of asparagine and glutamine, oxidation of methionine and iTRAQ4plex of tyrosine were specified in Mascot as variable modifications.
Criteria for protein identification
Scaffold (v.Scaffold_4.4.3, Proteome Software) was used to validate MS/MS-based peptide and protein identifications. Peptide identifications were accepted if they could be established at greater than 95.0% probability by the Peptide Prophet algorithm41 with Scaffold delta-mass correction. Protein identifications were accepted if they could be established at greater than 99.0% probability and contained at least 2 identified peptides. Protein probabilities were assigned by the Protein Prophet algorithm42. The original MS dataset (as published by Chu et al.21) is available at Proteome Xchange Consortium via the PRIDE partner repository with the identifier PXD003851.
Flow cytometry phenotyping of cord blood samples and HEK293 cells expressing RBP2a23–767
Three Percoll-enriched reticulocyte samples from human cord blood were treated or not with trypsin (1 mg ml–1; Sigma)28. Packed cord blood (500 nl, 47.4 ± 14.5% CD71+, n = 3) was stained with different mouse anti-human marker antibodies followed by a F(ab′)2 anti-mouse immunoglobulin antibody coupled to eFluor 660 (eBioscience)12. The antibodies used in this study are described in Supplementary Table 5. Fifty thousand events were acquired on a LSR II flow cytometer (BD Biosciences) for each sample, and the data were analysed using FlowJo software (Three Star).
HEK293 cells expressing RBP2a23–767 were stained with two mouse anti-PvRBP2a monoclonal antibodies (1C3 and 3A11) at 25 μg ml–1 followed by a (Fab′)2 anti-mouse immunoglobulin antibody coupled to eFluor 660 (eBioscience) or with anti-Myc (Miltenyi Biotec) antibodies followed by a (Fab′)2 anti-rabbit immunoglobulin antibody coupled to Alexa 647 (Invitrogen).
Profiling CD98 expression on RBCs by western blotting
Immature reticulocytes (CD71+) were purified from cord blood using magnetic beads conjugated to anti-CD71 antibody using the MACS purification system (Miltenyi Biotec). Purified CD71+ cells were allowed to mature ex vivo in RPMI medium supplemented with 5% Albumax at 37 °C. Aliquots of 3 μl packed cells were collected at different time points (0, 20 and 40 h). Cells were incubated with 40 volumes of 5 mM sodium phosphate for 10 min on ice with occasional gentle mixing, which induces membrane breaks and release of the cell cytoplasmic contents. The lysate was then centrifuged at 16,000 × g for 10 min at 4 °C to sediment the RBC ghosts that were again washed twice in ice-cold 5 mM sodium phosphate.
Purified ghost membranes were treated with 0.25% Triton X-100/PBS. Membrane extracts corresponding to 10 μg total protein (estimated using a BCA assay) obtained at different time points during reticulocyte maturation, together with a normocyte membrane sample that was prepared alongside, were extracted in reduced Laemmli buffer, incubated at 37 °C for 10 min, then resolved by 10% SDS–PAGE and transferred to polyvinylidenedifluoride membrane. Samples were probed using anti-CD98 antibody (E-5: sc-376815, Santa Cruz Biotechnology) at 1:500 dilution in PBS followed by a mouse secondary antibody conjugated to horseradish peroxidase (HRP) and visualized by ECL detection (Thermo Fisher Scientific). An antibody against Band-3 anion exchanger was used for loading control.
Immunofluorescence assay
RBCs (CD71+ and CD71–) were fixed in 4% paraformaldehyde (Sigma Aldrich) and 0.0075% glutaraldehyde (Sigma Aldrich) in PBS for 30 min at RT. Subsequently, cells were washed in 1× PBS, quenched in 0.125 M glycine/PBS for 15 min at RT, washed again and permeabilized in 0.1% Triton X-100/PBS for 10 min at RT. After blocking for 1 h in blocking solution (3% BSA (w/v) in PBS), samples were incubated with rabbit anti-CD98 antibody (EPR3548 clone, Abcam) or rabbit anti-Dematin (Band 4.9) antibody (Thermo Fisher Scientific) both at a 1:100 dilution, for 1 h, followed by Alexa Fluor 546-conjugated anti-rabbit secondary antibodies (1:200; Invitrogen). Images were acquired with a Zeiss LSM 700 microscope (Carl Zeiss) using a ×63/1.4 oil DIC objective lens. Images were processed using LSM software Zen 2009 (Carl Zeiss).
P. vivax parasite collection and cryopreservation
P. vivax-infected blood samples were collected from patients with malaria receiving treatment at clinics run by the SMRU located at the north-western border of Thailand. The project was explained to all the patients before they provided informed consent before collection of blood by venepuncture. Whole blood (5 ml) was collected in lithium heparin collection tubes. These samples were cryopreserved in Glycerolyte 57 (Baxter) after leukocyte depletion using cellulose columns (Sigma, C6288)43. After thawing, the parasites present in the packed cells (1.5 ml per isolate) were cultured to the schizont stage in 12 ml of McCoy 5A medium supplemented with 2.4 g per litre d-glucose and 20% heat-inactivated human AB serum, in 5% O2 at 37.5 °C4.
P. vivax invasion assay
Invasion assays using freshly isolated P. vivax field samples were performed as previously described28. In some experiments, anti-human CD147 (BD Bioscience, clone HIM6), CD240DCE (AbD Serotec, clone BRIC 69), polyclonal CD98 (TransGenic), monoclonal CD98 (BD Bioscience, clone UM7F8) and anti-DARC (Fy6)44 F(ab′)2 antibodies were added. The antibodies were prepared using a Pierce Fab Micro Preparation kit (ThermoScientific Pierce) as previously described45. We chose to use F(ab′)2 antibodies to avoid steric hindrance or agglutination of erythrocytes. All antibodies were diluted in PBS and were azide- and glycerol-free. Concentrated mature schizont preparations were mixed with the CD71+ reticulocyte-enriched fraction and pre-incubated for 10 min with each antibody (final concentration of 25 μg ml–1)28. After 24 h of culture at 37 °C, blood thin smears were made and stained with Giemsa (Sigma-Aldrich). One to five thousand erythrocytes were counted per slide, and the lowest parasitaemia was 0.3% for the negative control (no monoclonal antibody) condition used to normalize the invasion assays.
The anti-human CD147 (BD Bioscience, clone HIM6) abrogated the invasion of three P. falciparum samples (one field isolate and two clones) with an efficiency of 87.6 ± 2.4% and 86.0 ± 5.4% in CD71– and CD17+ reticulocytes, respectively.
P. vivax antigen library
The P. vivax antigen library was designed and established as previously described for a P. falciparum library46 using P. vivax UMS203 as the template. Briefly, 28 gene fragments corresponding to 7 different P. vivax genes were amplified using PCR and either P. vivax genomic DNA or RNA as template, and cloned into the pDisplay vector (Invitrogen) (Supplementary Table 2). Cloning of PvRBP2a,b,c into expression vector codon-optimized DNA of PvRBP2a,b,c was purchased from Genscript. The resultant plasmids were then transfected into HEK293 cells using Lipofectamine 2000 (Invitrogen) for surface expression of the P. vivax antigens. The P. vivax antigens were tagged with a haemagglutinin (HA) short sequence at the N-terminal and a Myc short sequence at the carboxy-terminal part of the protein. These tags allow the assessment of transfection efficacy and expression level of the P. vivax antigen using an anti-HA (Sigma) or anti-Myc (Miltenyi Biotec) antibodies. Only HEK293 cells expressing P. vivax genes with transfection efficiency >20% were used. The PvRBP2a diversity is compiled in Supplementary Table 3.
Erythrocyte binding assay using the P. vivax antigen library
Adherent HEK293 cells transfected with P. vivax antigens were incubated with CD71+ or CD71– erythrocytes loaded with carboxy-fluorescein diacetate succinimidyl ester (CFSE) for 1 h at 37 °C under agitation. After five washing steps using PBS, cell binding was recorded using a confocal microscope. One thousand HEK293 cells were counted at ×40 magnification, and each CFSE-stained erythrocyte in contact with HEK293 cells were counted as a binding cell using the software IMARIS (Bitplane). For each experiment, the number of erythrocytes binding untransfected HEK293 cells was subtracted.
Reticulocyte binding inhibition and the competition assay were conducted by incubating the PvRBP2a-transfected HEK293 cells before adding reticulocytes with an isotype control anti-TNP (BD Bioscience, clone A111-3) and two different mouse anti-PvRBP2a antibodies (clone1C3 and 3A11, described below) or mouse anti-CD98 monocolonal antibody (BD Bioscience, clone UM7F8) and mouse anti-human CD147 (BD Bioscience, clone HIM6) at 25 μg ml–1 in PBS.
Anti-PvRBP2a mouse monoclonal antibody production
Anti-PvRBP2a monoclonal antibodies were produced at the Monoclonal Antibody Facility at the Walter and Eliza Hall Institute. BALB/c and C57BL/6 mice received three immunizations of recombinant PvRBP2a160–1135 purified as previously described35. At day 0, complete Freund adjuvant was mixed with the antigen into an emulsion and intraperitoneally injected. At days 30 and 60, the antigen was mixed with incomplete Freund adjuvant, and the resulting emulsion intraperitoneally injected. Serum samples were taken at day 72 and ELISAs were performed using the same recombinant protein. The mouse with the best response received a final injection of antigen in saline, and splenocytes were collected 3 days later. Spleen cells were fused with SP2/0 myeloma cells to generate hybridomas. Hybridomas were grown in hypoxanthine-aminopterine thymidine growth medium. ELISA was used to select hybridomas producing antibodies specific to PvRBP2a. Hybridomas were cloned by limiting dilution in multiwell plates. The subcloning supernatants were screened by ELISA. Two or more rounds of limiting dilution cloning were generally required before the hybridomas were deemed monoclonal. The antibodies were purified from monoclonal hybridoma supernatants with protein A sepharose.
ELISA-based binding assay between PvRBP2a and CD98
To examine the binding between PvRBP2a23–767 and CD98 recombinant protein, Maxisorp plates were coated with 3 µg ml–1 CD98 or BSA in 0.1 M sodium bicarbonate buffer (pH 9.6) overnight at 4 °C. Plates were then blocked with 5% BSA in 0.05% PBST (PBS buffer supplemented with Tween-20) for 1.5 h at 37 °C. PvRBP2a in 0.5% BSA/0.05% PBST was then added at varying concentrations from 0 nM to 50 nM and incubated for 1 h at 37 °C. To detect the quantity of bound PvRBP2a, rabbit anti-PvRBP2a antibody was added and incubated for 1 h at 37 °C, followed by a secondary goat anti-rabbit HRP-conjugated antibody, with a colorimetric TMB substrate-based readout.
ELISA-based solution binding K D between PvRBP2a and CD98
The KD of binding between PvRBP2a and CD98 was quantified using the method outlined in Friguet et al.29. PvRBP2a (25 nM) and CD98 (0 nM to 640 nM) were pre-incubated in solution for binding to occur. The duration of pre-incubation was optimized to 2.5 h at 37 °C to allow maximal time for the binding reaction to reach equilibrium while minimizing protein degradation. After the incubation time, the solution was applied to plates pre-coated with 3 µg ml–1 CD98 and blocked with 5% BSA/0.05% PBST, followed by secondary detection with rabbit anti-PvRBP2a and goat anti-rabbit HRP antibodies as described above. Resulting optical density values were transformed to molar PvRBP2a concentrations by normalization to a set of standards run in parallel. This gave the concentration of unbound PvRBP2a at equilibrium ([PvRBP2afinal]), assuming that the PvRBP2a already bound to solution-phase CD98 cannot bind to the plate-coated CD98. For each mixture of PvRBP2a and CD98, the concentration of PvRBP2a bound by CD98 in solution during the pre-incubation phase ([CD98:PvRBP2afinal]) was calculated as the difference between its concentration of unbound PvRBP2a ([PvRBP2afinal]) and the PvRBP2a concentration of a control well with the same input PvRBP2a concentration and pre-incubation conditions but which contained no CD98 ([PvRBP2ainitial]). The final concentration of CD98 at equilibrium, [CD98final], was calculated by subtracting the concentration of bound CD98 ([CD98:PvRBP2afinal]) at equilibrium from the initial known input concentration of CD98 ([CD98initial]). With these values, the KD, which for a 1:1 binding relationship is defined as [PvRBP2afinal][CD98final]/[CD98:PvRBP2afinal], can be calculated. Since the equation for KD can be rewritten as 1/[CD98:PvRBP2afinal] = KD × 1/([CD98final] × [PvRBP2ainitial]) + 1/[PvRBP2ainitial], which is in the linear equation format y = mx + b, the gradient of the best fit line when plotting 1/[CD98:PvRBP2afinal] against 1/([CD98final] × [PvRBP2ainitial]) shows the observed KD across varying ratios of input PvRBP2a and CD98 concentrations.
Kinetic binding assay using the Octet Red96e system
Binding sensorgrams were collected using eight-channel detection mode on the Octet Red 96e system. Fresh streptavidin sensors were used without any regeneration step. The streptavidin sensor, without biotinylated CD98 loading, was used as the reference. Kinetic binding assays were performed using Octet Data Acquisition (v.10.0.1.3) at 37 °C with the orbital shaking speed set at 1,000 r.p.m. The assay was carried out in PBS buffer supplemented with 0.05% Tween-20 and 1 mg ml–1 BSA (PBST/BSA). The biosensor was sequentially dipped into wells containing 200 μl each of the following solution: (1) baseline, PBST/BSA (60 s); (2) loading, 25 nM biotinylated CD98 (5 min); (3) washing, PBST/BSA (60 s); (4) blocking, 25 μM biocytin (5 min) to minimize nonspecific binding of PvRBP2a to the sensor; (5) washing, PBST/BSA (60 s); (6) association, varying concentrations of non-biotinylated P9 (30 min); and (7) dissociation, PBST/BSA (30 min). Binding sensorgrams were analysed using ForteBio’s Data Analysis (v.10.0). All sensorgrams were subtracted to the reference sensor and aligned to the baseline. The sensorgrams were globally fit to a 1:1 Langmuir binding model. Association (ka, M−1 s−1), dissociation (kd, s−1) and affinity constants (KD, nM) were calculated based on quadruplicate runs and represented as mean ± s.e.m.
Immunoprecipitation of full-length PvRBP2a
A total of 25 µg of PvRBP2a23–767 and His-tagged CD98 or His-tagged CD98 alone were incubated at 37 °C for 2 h in IP buffer 1 (50 mM Tris HCL pH 7.5, 1% Triton and 150 mM NaCl). In parallel, 100 µl of protein A/G magnetic beads (Pierce, 88802) were washed with IP buffer 1 and incubated at RT with 20 µg of anti-PvRBP2a11. Beads were then washed three times before incubating for 1 h at 37 °C with lysis buffer 1 alone or with lysis buffer 1 containing CD98 or CD98 and PvRBP2a23–767. Beads were then washed four times with IP buffer 2 (50 mM Tris HCL pH 7.5 (Gibco), 1% IGEPAL (Sigma), 0.5% sodium deoxycholate (Sigma) and 150 mM NaCl (Ambion)). This was followed by two washes with tube change in 50 mM Tris HCl to avoid nonspecific carry over, and elution in 1× Laemmli buffer (Bio-Rad) with β-mercaptoethanol (Sigma) and boiling for 5–10 min before western blotting was performed as described in Carissimo et al.47. Briefly, samples were deposited on 4–15% or 4–20% TGX mini-protean gels (Bio-Rad), with Spectra BR (ThermoFisher) used as a protein ladder, and ran in Bis-Tris buffer (Bio-Rad) at 120 V, and transferred to nitrocellulose membrane (Bio-Rad) using a semi-dry system and Bjerrum Schafer–Nielsen buffer (48 mM UltraPure Tris (Invitrogen), 38 mM glycine (Bio-Rad) and 20% (v/v) EMSURE methanol (EMD Millipore)). Membranes were then blocked in 5% (w/v) non-fat milk powder (Nacalai Tesque) in 1× Tris buffer saline (first base) with Tween 0.1% (v/v) (Sigma) (TBST) for 1 h before staining in blocking buffer. Antibodies used for staining were anti-His-HRP (Sigma Aldrich) at 1:10,000 dilution to detect His-tagged CD98, anti- PvRBP2a11 at 1:5,000 dilution and protein A/G-HRP (Pierce, 32490) at 1:15,000 dilution. Revelation was performed after membrane washing in 1× TBST using WesternBright ECL HRP substrate (Advansta), nand the signal was acquired as multichannel (chemiluminescence and colorimetric) on a ChemiDoc screen touch model 2017 (Bio-Rad). Non-saturated images were analysed using ImageLab 2.3 software (Bio-Rad) and exported in TIFF format. Arrows indicated the protein ladder were added to the chemiluminescent images and then the resulting image was copied and incorporated in Illustrator (v.16.0.0, Adobe) to produce the figure panel. No brightness or contrast correction was applied to the images. Untouched full-size membrane images of the merged channels are provided in the source data. Experiments were performed three times independently and representative results are presented.
Statistical analysis
Nonparametric Kruskal–Wallis test followed by Dunn’s post-test was used for the invasion inhibition assays (Fig. 2a), as the percentage of infected reticulocytes is not normally distributed28. For the difference in geometric mean (log-normalized data) fluorescence intensity (MFI), unpaired Student t-test was used (Extended Data Fig. 2c). D’Agostino’s K-squared test was used to determine the normal distribution of reticulocyte binding assay data (Extended Data Fig. 3d) and unpaired Student t-test for the comparison between transfected and non-transfected HEK293 cells assuming unequal variances of the data (Fig. 3b,c). One-way analysis of variance (ANOVA) with Geisser–Greenhouse correction (variance not equal) followed by Tukey post-test was used for the comparison of reticulocyte binding to HEK293-transfected cells with isotype control or anti-PvRBP2a antibodies (Fig. 3d). All statistical analyses used Graph Pad Prism (7.0).
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. All other data are available from the corresponding authors upon reasonable request.
References
Confronting Plasmodium vivax malaria. WHO/HTM/GMP/2015.3 (World Health Orgainization, 2015).
Hegner, R. Relative frequency of ring-stage plasmodia in reticulocytes and mature erythrocytes in man an monkey. Am. J. Trop. Med. Hyg. 27, 690–718 (1938).
Mons, B., Croon, J. J., van der Star, W. & van der Kaay, H. J. Erythrocytic schizogony and invasion of Plasmodium vivax in vitro. Int. J. Parasitol. 18, 307–311 (1988).
Malleret, B. et al. Plasmodium vivax: restricted tropism and rapid remodeling of CD71-positive reticulocytes. Blood 125, 1314–1324 (2015).
Hester, J. et al. De novo assembly of a field isolate genome reveals novel Plasmodium vivax erythrocyte invasion genes. PLoS Negl. Trop. Dis. 7, e2569 (2013).
Carlton, J. M. et al. Comparative genomics of the neglected human malaria parasite Plasmodium vivax. Nature 455, 757–763 (2008).
Han, J. H. et al. Identification of a reticulocyte-specific binding domain of Plasmodium vivax reticulocyte-binding protein 1 that is homologous to the PfRh4 erythrocyte-binding domain. Sci. Rep. 6, 26993 (2016).
Gupta, S. et al. Targeting a reticulocyte binding protein and Duffy binding protein to inhibit reticulocyte invasion by Plasmodium vivax. Sci. Rep. 8, 10511 (2018).
Ntumngia, F. B. et al. Identification and immunological characterization of the ligand domain of Plasmodium vivax reticulocyte binding protein 1a. J. Infect. Dis. 218, 1110–1118 (2018).
Chim-Ong, A. et al. The blood stage antigen RBP2-P1 of Plasmodium vivax binds reticulocytes and is a target of naturally acquired immunity. Infect. Immun. 88, e00616–e00619 (2020).
Gruszczyk, J. et al. Transferrin receptor 1 is a reticulocyte-specific receptor for Plasmodium vivax. Science 359, 48–55 (2018).
Malleret, B. et al. Significant biochemical, biophysical and metabolic diversity in circulating human cord blood reticulocytes. PLoS ONE 8, e76062 (2013).
Proto, W. R. et al. Adaptation of Plasmodium falciparum to humans involved the loss of an ape-specific erythrocyte invasion ligand. Nat. Commun. 10, 4512 (2019).
Kosaisavee, V. et al. Strict tropism for CD71+/CD234+ human reticulocytes limits the zoonotic potential of Plasmodium cynomolgi. Blood 130, 1357–1363 (2017).
Wright, G. J. & Rayner, J. C. Plasmodium falciparum erythrocyte invasion: combining function with immune evasion. PLoS Pathog. 10, e1003943 (2014).
Crosnier, C. et al. Basigin is a receptor essential for erythrocyte invasion by Plasmodium falciparum. Nature 480, 534–537 (2011).
Douglas, A. D. et al. Neutralization of Plasmodium falciparum merozoites by antibodies against PfRH5. J. Immunol. 192, 245–258 (2014).
Barnwell, J. W., Nichols, M. E. & Rubinstein, P. In vitro evaluation of the role of the Duffy blood group in erythrocyte invasion by Plasmodium vivax. J. Exp. Med. 169, 1795–1802 (1989).
Malleret, B., Renia, L. & Russell, B. The unhealthy attraction of Plasmodium vivax to reticulocytes expressing transferrin receptor 1 (CD71). Int. J. Parasitol. 47, 379–383 (2017).
Kanjee, U. et al. Plasmodium vivax strains use alternative pathways for invasion. J. Infect. Dis. 223, 1817–1821 (2020).
Chu, T. T. T. et al. Quantitative mass spectrometry of human reticulocytes reveal proteome-wide modifications during maturation. Br. J. Haematol. 180, 118–133 (2018).
Boado, R. J., Li, J. Y., Nagaya, M., Zhang, C. & Pardridge, W. M. Selective expression of the large neutral amino acid transporter at the blood–brain barrier. Proc. Natl Acad. Sci. USA 96, 12079–12084 (1999).
Segawa, H. et al. Identification and functional characterization of a Na+-independent neutral amino acid transporter with broad substrate selectivity. J. Biol. Chem. 274, 19745–19751 (1999).
Fort, J. et al. The structure of human 4F2hc ectodomain provides a model for homodimerization and electrostatic interaction with plasma membrane. J. Biol. Chem. 282, 31444–31452 (2007).
Yan, R., Zhao, X., Lei, J. & Zhou, Q. Structure of the human LAT1-4F2hc heteromeric amino acid transporter complex. Nature 568, 127–130 (2019).
Chiduza, G. N. et al. LAT1 (SLC7A5) and CD98hc (SLC3A2) complex dynamics revealed by single-particle cryo-EM. Acta Crystallogr. D Struct. Biol. 75, 660–669 (2019).
Lee, Y. et al. Cryo-EM structure of the human l-type amino acid transporter 1 in complex with glycoprotein CD98hc. Nat. Struct. Mol. Biol. 26, 510–517 (2019).
Russell, B. et al. A reliable ex vivo invasion assay of human reticulocytes by Plasmodium vivax. Blood 118, e74–e81 (2011).
Friguet, B., Chaffotte, A. F., Djavadi-Ohaniance, L. & Goldberg, M. E. Measurements of the true affinity constant in solution of antigen–antibody complexes by enzyme-linked immunosorbent assay. J. Immunol. Methods 77, 305–319 (1985).
Batchelor, J. D. et al. Red blood cell invasion by Plasmodium vivax: structural basis for DBP engagement of DARC. PLoS Pathog. 10, e1003869 (2014).
Tham, W. H. et al. Complement receptor 1 is the host erythrocyte receptor for Plasmodium falciparum PfRh4 invasion ligand. Proc. Natl Acad. Sci. USA 107, 17327–17332 (2010).
Srivastava, A. et al. Host reticulocytes provide metabolic reservoirs that can be exploited by malaria parasites. PLoS Pathog. 11, e1004882 (2015).
Obaldia, N. III et al. Bone marrow is a major parasite reservoir in Plasmodium vivax infection. mBio 9, e00625-18 (2018).
Galinski, M. R., Medina, C. C., Ingravallo, P. & Barnwell, J. W. A reticulocyte-binding protein complex of Plasmodium vivax merozoites. Cell 69, 1213–1226 (1992).
Gruszczyk, J. et al. Structurally conserved erythrocyte-binding domain in Plasmodium provides a versatile scaffold for alternate receptor engagement. Proc. Natl Acad. Sci. USA 113, E191–E200 (2016).
Franca, C. T. et al. Plasmodium vivax reticulocyte binding proteins are key targets of naturally acquired immunity in young Papua New Guinean children. PLoS Negl. Trop. Dis. 10, e0005014 (2016).
Ryan, J. R. et al. Evidence for transmission of Plasmodium vivax among a duffy antigen negative population in Western Kenya. Am. J. Trop. Med. Hyg. 75, 575–581 (2006).
Menard, D. et al. Plasmodium vivax clinical malaria is commonly observed in Duffy-negative Malagasy people. Proc. Natl Acad. Sci. USA 107, 5967–5971 (2010).
Cavasini, C. E. et al. Plasmodium vivax infection among Duffy antigen-negative individuals from the Brazilian Amazon region: an exception? Trans. R. Soc. Trop. Med. Hyg. 101, 1042–1044 (2007).
Chitnis, C. E. & Miller, L. H. Identification of the erythrocyte binding domains of Plasmodium vivax and Plasmodium knowlesi proteins involved in erythrocyte invasion. J. Exp. Med. 180, 497–506 (1994).
Keller, A., Nesvizhskii, A. I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Nesvizhskii, A. I., Keller, A., Kolker, E. & Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 75, 4646–4658 (2003).
Sriprawat, K. et al. Effective and cheap removal of leukocytes and platelets from Plasmodium vivax infected blood. Malar. J. 8, 115 (2009).
Wasniowska, K. et al. Structural characterization of the epitope recognized by the new anti-Fy6 monoclonal antibody NaM 185-2C3. Transfus. Med 12, 205–211 (2002).
Lee, W. C. et al. Glycophorin C (CD236R) mediates vivax malaria parasite rosetting to normocytes. Blood 123, e100–e109 (2014).
Peng, K. et al. Breadth of humoral response and antigenic targets of sporozoite-inhibitory antibodies associated with sterile protection induced by controlled human malaria infection. Cell Microbiol. 18, 1739–1750 (2016).
Carissimo, G. et al. VCP/p97 Is a proviral host factor for replication of chikungunya virus and other alphaviruses. Front. Microbiol. 10, 2236 (2019).
Acknowledgements
We are indebted to C. Chu, R. McGready and the staff of the Mae Sot Malaria Clinic and the clinics associated with SMRU (Tak Province, Thailand) and the patients attending these clinics. We thank M. Mauduit for help at the beginning of the project. We thank the SIgN flow cytometry platform (supported by a grant from the National Research Foundation, Immunomonitoring Service Platform ISP) (NRF2017_SISFP09)). B.R. and B.M. were funded by the Singapore National Medical Research Council (NMRC/CBRG/0047/2013). B.M. was also funded by the Agency for Science, Technology and Research (A*STAR, Singapore) Young Investigator Grant (BMRC YIG grant no: 13/1/16/YA/009), core funds to SIgN from A*STAR, NUHS Start-up grant (NUHSRO/2018/006/SU/01) and MOE Tier 1 (NUHSRO/2018/094/T1/SEED-NOV/04). L.R. was supported by a Singapore National Medical Research Council IRG grant (NMRC/OFIRG/0065/2018), by a Singapore Immunology Network core research grant and by the Horizontal Programme on Infectious Diseases under A*STAR. SMRU is supported by The Wellcome Trust of Great Britain as part of the Oxford Tropical Medicine Research Programme of Wellcome Trust–Mahidol University. R.C. acknowledges funding support through the following grants: T1MOE1702 (MOE Tier 1 Grant through the Singapore University of Technology & Design) and RGUOO180301 (Marsden Grant Sub-award through the University of Otago). W.-H.T. was funded by the Australian Research Council Future Fellowship. Duke-NUS Medical School efforts were supported by Singapore’s Health and Biomedical Sciences (HBMS) Industry Alignment Fund Pre-Positioning (IAF-PP) grant H18/01/a0/018, administered by A*STAR.
Author information
Authors and Affiliations
Contributions
B.M. and B.R. carried out the phenotyping characterization of the erythrocytes and the antibody validations for the P. vivax invasion assays. G.C., S.W.H., R.S., V.K. and A.S.M.O. developed the P. vivax library and performed the erythrocyte binding assays. T.T.T.C., A.S. and R.C. performed and analysed the MS data. J.G. and W.-H.T. carried out construct design and protein purification for the PvRBP2a recombinant proteins and the anti-PvRBP2a monoclonal antibodies. Y.C. developed the anti-Duffy antibodies. J.K.Y.C., Y.F. and F.N. were in charge of the management of clinical data. A.E.S., J. Lin, J. Lescar, G.C. and L.F.P.N. managed the biochemistry aspects of the project. S.M.-S. managed the protein modelling of the interaction between CD98 and PvRPB2a. W.N., M.Z.T. and A.-M.C. managed the Octet experiments for CD98 and PvRPB2a interaction measurements. Overall project management was carried out by B.M., B.R. and L.R. The manuscript was prepared by B.M., G.S., B.R. and L.R.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Tuan Tran, Sanjeeva Srivastava and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Reticulocyte phenotyping.
Trypsin resistance profile of different markers expressed at the surface of cord blood reticulocytes. The black and yellow histograms represent the level of expression before and after trypsin treatment respectively. The trypsin sensitive proteins are annotated in red and the resistant ones in black. The isotype antibody used as control is represented in grey. We used human cord blood samples pre-enriched with reticulocytes and that thus still harbour some normocytes, hence the double peak observed for some proteins such as CD71 and CD98.
Extended Data Fig. 2 Flow cytometry analysis of adult peripheral blood, cord blood and P. vivax isolates.
a, Flow cytometry profile of CD98 and CD71 expression on thiazole orange (TO) negative, low and high erythrocytes for three different human adult peripheral blood samples (forward scatter (FSC) for x-axis and side scatter (SSC) for y-axis). The TO high subset represents the most immature reticulocyte population compared to low subset. b, Flow cytometry histogram of CD98 expression (right) at the surface of P. vivax rings, early infected reticulocyte forms, gated on Hoechst-positive cells (left), side scatter (SSC) for y-axis. The infected cells positive for Hoechst (a DNA stain) are in blue and the uninfected red blood cells (uRBC), which do not contain DNA, are in red. c, Comparison of delta of geometric mean fluorescence intensity (MFI) between immature reticulocytes from three cord blood samples and three vivax infected patients (ring stage), Values are expressed as mean ± SD, unpaired Student t-test.
Extended Data Fig. 3 Binding assay of erythrocytes to HEK cells expressing P. vivax genes.
a, Schematic description of the development of the library and its use in the erythrocyte binding assay: (1) cloning of P. vivax genes in the pDisplay plasmid, (2) transfection of HEK cells, (3) expression of the protein containing MYC and HA tags at the surface of HEK cells and (5) binding assay with erythrocytes. b, Schematic representation of full-length PvRBP2a and of its recombinant protein fragments used in this study (right). Signal peptide (SP), transmembrane helix (TM) and nucleotide-binding domain (green) are indicated. The ticks indicated for nonsynonymous SNPs with alternate allele frequency >20%. c, Representative images from 3 independent experiments of binding of reticulocytes (CD71 + ) or normocytes (CD71-) loaded with the fluorescent dye CFSE to HEK cells expressing the PvRBP2a 1315-1650 (negative control), Duffy binding protein II (DBPII) fragment or PvRBP2a23-767 in the presence or absence of anti-CD98 antibodies. The scale represents 50 μm. d, The binding of reticulocytes to HEK transfected with the PvRBP2a23–767 fragment was done in octuplicate (eight independent experiments) and was shown to follow a normal distribution as determine by the D’Agostino’s K-squared test, and differed significantly from binding to non-transfected HEK cells, Values are expressed as mean ± SD, Student t-test. e, Binding assays of reticulocytes to HEK cells expressing PvRBP2b481-1530 in the presence or absence of anti-CD98 antibodies. (three independent experiments). Values are expressed as mean ± SD, no significant differences were observed. e, Binding assays of normocytes to HEK cells expressing PvRBP2a23-767 in the presence or absence of anti-CD98 antibodies (three independent experiments). Values are expressed as mean ± SD, no significant differences were observed.
Extended Data Fig. 4 Antigen specificity of anti-PvRBP2a antibodies.
Mouse 1C3 and 3A11 mAbs recognize specifically the PvRBP2a fragment, PvRBP2a23–767, expressed by HEK cells when tested by flow cytometry. As positive control, rabbit anti-myc antibodies were used. Secondary anti-mouse IgG antibodies coupled with eFluor 660 or mouse anti-rabbit immunoglobulin coupled with Alexa 674 were used as negative control.
Supplementary information
Supplementary Tables
Combined PDF with Supplementary Tables 1–5.
Source data
Source Data Fig. 1
Raw data.
Source Data Fig. 1
Unprocessed western blot.
Source Data Fig. 2
Raw data.
Source Data Fig. 3
Raw data.
Source Data Fig. 4
Raw data.
Source Data Fig. 4
Unprocessed immunoprecipitation gel.
Source Data Extended Data Fig. 2
Raw data.
Source Data Extended Data Fig. 3
Raw data.
Rights and permissions
About this article
Cite this article
Malleret, B., El Sahili, A., Tay, M.Z. et al. Plasmodium vivax binds host CD98hc (SLC3A2) to enter immature red blood cells. Nat Microbiol 6, 991–999 (2021). https://doi.org/10.1038/s41564-021-00939-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-021-00939-3
This article is cited by
-
Plasmodium vivax merozoite-specific thrombospondin-related anonymous protein (PvMTRAP) interacts with human CD36, suggesting a novel ligand–receptor interaction for reticulocyte invasion
Parasites & Vectors (2023)
-
Development of a Plasmodium vivax biobank for functional ex vivo assays
Malaria Journal (2023)
-
Systems biology of malaria explored with nonhuman primates
Malaria Journal (2022)
-
N-glycosylation is crucial for trafficking and stability of SLC3A2 (CD98)
Scientific Reports (2022)
-
The genome of the zoonotic malaria parasite Plasmodium simium reveals adaptations to host switching
BMC Biology (2021)