Abstract
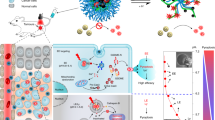
Plasma membrane rupture is a promising strategy for drug-resistant cancer treatment, but its application is limited by the low tumour selectivity of membranolytic molecules. Here we report the design of ‘proton transistor’ nanodetergents that can convert the subtle pH perturbation signals of tumour tissues into sharp transition signals of membranolytic activity for selective cancer therapy. Our top-performing ‘proton transistor’ nanodetergent, P(C6-Bn20), can achieve a >32-fold change in cytotoxicity with a 0.1 pH input signal. At physiological pH, P(C6-Bn20) self-assembles into neutral nanoparticles with inactive membranolytic blocks shielded by poly(ethylene glycol) shells, exhibiting low toxicity. At tumour acidity, a sharp transition in its protonation state induces a morphological transformation and an activation of the membranolytic blocks, and the cation–π interaction facilitates the insertion of benzyl groups-containing hydrophobic domains into the cell membranes, resulting in potent membranolytic activity. P(C6-Bn20) is well tolerated in mice and shows high anti-tumour efficacy in various mouse tumour models.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All relevant data of this study are available within the paper and its Supplementary Information files. Full gels of the SDS–PAGE image in Fig. 3i are provided as Source data with this paper. The raw and analysed datasets generated during the study are available for research purposes from the corresponding authors on reasonable request.
Change history
12 May 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41565-022-01139-8
References
Hancock, R. E. W. & Sahl, H. G. Antimicrobial and host-defense peptides as new anti-infective therapeutic strategies. Nat. Biotechnol. 24, 1551–1557 (2006).
Nederberg, F. et al. Biodegradable nanostructures with selective lysis of microbial membranes. Nat. Chem. 3, 409–414 (2011).
Trambas, C. M. & Griffiths, G. M. Delivering the kiss of death. Nat. Immunol. 4, 399–403 (2003).
Leung, C. et al. Real-time visualization of perforin nanopore assembly. Nat. Nanotechnol. 12, 467–473 (2017).
Fjell, C. D., Hiss, J. A., Hancock, R. E. W. & Schneider, G. Designing antimicrobial peptides: form follows function. Nat. Rev. Drug Discov. 11, 37–51 (2012).
Park, N. H. et al. Addressing drug resistance in cancer with macromolecular chemotherapeutic agents. J. Am. Chem. Soc. 140, 4244–4252 (2018).
Magana, M. et al. The value of antimicrobial peptides in the age of resistance. Lancet Infect. Dis. 20, e216–e230 (2020).
Shen, W. et al. Antineoplastic drug-free anticancer strategy enabled by host-defense-peptides-mimicking synthetic polypeptides. Adv. Mater. 32, 2001108 (2020).
Mookherjee, N., Anderson, M. A., Haagsman, H. P. & Davidson, D. J. Antimicrobial host defence peptides: functions and clinical potential. Nat. Rev. Drug Discov. 19, 311–332 (2020).
Voskoboinik, I., Whisstock, J. C. & Trapani, J. A. Perforin and granzymes: function, dysfunction and human pathology. Nat. Rev. Immunol. 15, 388–400 (2015).
Tew, G. N., Scott, R. W., Klein, M. L. & Degrado, W. F. De novo design of antimicrobial polymers, foldamers, and small molecules: from discovery to practical applications. Acc. Chem. Res. 43, 30–39 (2010).
Palermo, E. F., Lienkamp, K., Gillies, E. R. & Ragogna, P. J. Antibacterial activity of polymers: discussions on the nature of amphiphilic balance. Angew. Chem. Int. Ed. Engl. 58, 3690–3693 (2019).
Liang, Y. B., Zhang, X. S., Yuan, Y. L., Bao, Y. & Xiong, M. H. Role and modulation of the secondary structure of antimicrobial peptides to improve selectivity. Biomater. Sci. 8, 6858–6866 (2020).
Mishra, B., Narayana, J. L., Lushnikova, T., Wang, X. Q. & Wang, G. S. Low cationicity is important for systemic in vivo efficacy of database-derived peptides against drug-resistant Gram-positive pathogens. Proc. Natl Acad. Sci. USA 116, 13517–13522 (2019).
Chin, W. et al. A macromolecular approach to eradicate multidrug resistant bacterial infections while mitigating drug resistance onset. Nat. Commun. 9, 917 (2018).
de Breij, A. et al. The antimicrobial peptide SAAP-148 combats drug-resistant bacteria and biofilms. Sci. Transl. Med. 10, eaan4044 (2018).
Xiong, M. et al. Helical antimicrobial polypeptides with radial amphiphilicity. Proc. Natl Acad. Sci. USA 112, 13155–13160 (2015).
Mowery, B. P., Lindner, A. H., Weisblum, B., Stahl, S. S. & Gellman, S. H. Structure–activity relationships among random nylon-3 copolymers that mimic antibacterial host-defense peptides. J. Am. Chem. Soc. 131, 9735–9745 (2009).
Fan, Y. et al. A biomimetic peptide recognizes and traps bacteria in vivo as human defensin-6. Sci. Adv. 6, eaaz4767 (2020).
Xiong, M. H. et al. Bacteria-assisted activation of antimicrobial polypeptides by a random-coil to helix transition. Angew. Chem. Int. Ed. 56, 10826–10829 (2017).
Xiong, M. H. et al. Selective killing of Helicobacter pylori with pH-responsive helix–coil conformation transitionable antimicrobial polypeptides. Proc. Natl Acad. Sci. USA 114, 12675–12680 (2017).
Qi, G. B., Zhang, D., Liu, F. H., Qiao, Z. Y. & Wang, H. An “on-site transformation” strategy for treatment of bacterial infection. Adv. Mater. 29, 1703461 (2017).
Wang, Y. G. et al. A nanoparticle-based strategy for the imaging of a broad range of tumours by nonlinear amplification of microenvironment signals. Nat. Mater. 13, 204–212 (2014).
Zhao, T. et al. A transistor-like pH nanoprobe for tumour detection and image-guided surgery. Nat. Biomed. Eng. 1, 0006 (2017).
Webb, B. A., Chimenti, M., Jacobson, M. P. & Barber, D. L. Dysregulated pH: a perfect storm for cancer progression. Nat. Rev. Cancer 11, 671–677 (2011).
Anderson, M., Moshnikova, A., Engelman, D. M., Reshetnyak, Y. K. & Andreev, O. A. Probe for the measurement of cell surface pH in vivo and ex vivo. Proc. Natl Acad. Sci. USA 113, 8177–8181 (2016).
Gerweck, L. E. & Seetharaman, K. Cellular pH gradient in tumor versus normal tissue: potential exploitation for the treatment of cancer. Cancer Res. 56, 1194–1198 (1996).
Feng, L. Z., Dong, Z. L., Tao, D. L., Zhang, Y. C. & Liu, Z. The acidic tumor microenvironment: a target for smart cancer nano-theranostics. Natl Sci. Rev. 5, 269–286 (2018).
Sun, L. et al. MiR-200b and miR-15b regulate chemotherapy-induced epithelial-mesenchymal transition in human tongue cancer cells by targeting BMI1. Oncogene 31, 432–445 (2012).
Meyer, E. A., Castellano, R. K. & Diederich, F. Interactions with aromatic rings in chemical and biological recognition. Angew. Chem. Int. Ed. 42, 1210–1250 (2003).
Dougherty, D. A. Cation-π interactions in chemistry and biology: a new view of benzene, Phe, Tyr, and Trp. Science 271, 163–168 (1996).
Li, Y. et al. Chaotropic-anion-induced supramolecular self-assembly of ionic polymeric micelles. Angew. Chem. Int. Ed. 53, 8074–8078 (2014).
Riedl, S. et al. In search of a novel target—phosphatidylserine exposed by non-apoptotic tumor cells and metastases of malignancies with poor treatment efficacy. Biochim. Biophys. Acta Biomembr. 1808, 2638–2645 (2011).
Birge, R. B. et al. Phosphatidylserine is a global immunosuppressive signal in efferocytosis, infectious disease, and cancer. Cell Death Differ. 23, 962–978 (2016).
Ma, X. P. et al. Ultra-pH-sensitive nanoprobe library with broad pH tunability and fluorescence emissions. J. Am. Chem. Soc. 136, 11085–11092 (2014).
Li, H. J. et al. Smart superstructures with ultrahigh pH-sensitivity for targeting acidic tumor microenvironment: instantaneous size switching and improved tumor penetration. ACS Nano 10, 6753–6761 (2016).
Acknowledgements
We thank G. Yao (Nanfang Hospital) for providing the patient sample. We thank J. Li (Sun Yat-sen Memorial Hospital, Sun Yat-sen University) for providing the cisplatin-resistant human tongue cancer cell line CAL27/DDP. This work was supported by National Key R&D Program of China grant no. 2017YFA0205600 (J.W.); the National Natural Science Foundation of China grants no. 51633008 (J.W.), no. 52073102 (M.X.), no. 32171319 (Y.B.), no. 31971297 (Y.B.), no. U1801252 (M.X.) and no. 32000949 (M.L.); the Program for Guangdong Introducing Innovative and Entrepreneurial Teams grant no. 2017ZT07S054 (J.W.); Guangdong Provincial Pearl River Talents Program grants no. 2019QN01Y090 (M.X.) and no. 2017GC010713 (J.W.); the Key Research and Development Program of Guangzhou grant no. 202007020002 (M.X.); the Natural Science Foundation of Guangdong Province grants no. 2019A1515110617 (M.L.) and no. 2021A1515012492 (M.L.); the China Postdoctoral Science Foundation grant no. 2019M652909 (M.L.); and Guangdong Science and Technology Department grants no. 2020B1212060018 (Y.B.) and no. 2020B1212030004 (Y.B.).
Author information
Authors and Affiliations
Contributions
M.L., L.H. and Weinan Zhang designed and performed the majority of the experiments, analysed the data and wrote the manuscript. X.W. and Y.Z. assisted with polymer synthesis and characterization. Y.G. performed the GUV study. L.W. helped with the cryo-TEM observation. S.X. performed and analysed the simulation studies, and wrote the manuscript. Wenbin Zhang helped with the PDX model. Y.-J.Z. provided the PDX model. Y.B., M.X. and J.W. supervised all of the experiments, analysed the data and revised the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
M.L., L.H., X.W., Y.B., M.X. and J.W. have submitted a patent application (PCT/CN2021/102758) related to this study. The other authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Cameron Alexander, Yiyan Yang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The proton transistor-like properties of P(C6-Rx) copolymers.
a, PD as a function of pH for P(C6-Rx) copolymers when R was methyl (Me), ethyl (E), butyl (Bu), hexyl (H), iso-octyl (IO), tetradecyl (T). b, Merged images of Panc02GFP/mCherry cells incubated with P(C6-Rx) copolymers (160 μg ml-1) at different pH conditions for 2 h by a HCAS. The red fluorescence represents the fluorescent protein mCherry, and the green fluorescence represents MyrPalm-GFP. The scale bar is 40 μm. The cytotoxicity of P(C6-Bnx) (c) and P(C6-R20) (d) copolymers (160 μg ml-1) against Panc02 cells at different pH conditions after 2 h incubation. In c, d, n = 3 biologically independent samples, and the data are presented as the mean ± s.d. The representative images in b were shown from two independent experiments.
Extended Data Fig. 2 The cytotoxicity of P(C6-Rx) at pH 7.4 or 6.8.
The concentration dependent cytotoxicity of pTNTs against Panc02 cells after 24 h incubation at pH 7.4 (a) and after 4 h incubation at pH 6.8 (b). c, The concentration-dependent cytotoxicity of P(C6-Bn20) against normal cell lines at pH 7.4 after 24 h of incubation. d, The concentration-dependent cytotoxicity of P(C6-Bn20) against a variety of cancer cells at pH 6.8 after 4 h incubation. In c, d, n = 3 biologically independent samples, and the data are presented as the mean ± s.d.
Extended Data Fig. 3 The chemical structures, pH dependent PD and cytotoxicity of P(C6-Ph20) and P(C6-NPh20).
Illustration of chemical structure of P(C6-Ph20) (a) and P(C6-NPh20) (b). c, The PD as a function of pH for P(C6-Ph20) and P(C6-NPh20). The concentration dependent cytotoxicity of P(C6-Ph20) and P(C6-NPh20) against Panc02 cells after 24 h incubation at pH 7.4 (d) and after 4 h incubation at pH 6.8 (e). In d, e, n = 3 biologically independent samples, and the data are presented as the mean ± s.d.
Extended Data Fig. 4 Atomic molecular dynamics simulation studies.
The recorded force of pulling a P(C6-Bn20)l (a), P(C6-Bn20)h (b), P(C6-H20)l (c) and P(C6-H20)h (d) molecule away from an aggregated particle of 8 molecules as a function of the distance between the selected molecule and the mass centre of the other seven molecules. Time evolution of the interactions between P(C6-Bn20)l (e), P(C6-Bn20)h (f), P(C6-H20)l (g) and P(C6-H20)h (h) molecules with bilayer membranes (a mixed PC/PS bilayer with a fixed ratio of 3:1). The recorded force of pulling a molecule of P(C6-Bn20)h (i) and P(C6-H20)h (j) towards the bilayer membrane as a function of the height between the molecule and the bilayer centre. In a-d, i, j, n ≥ 10 independent experiments, and the data are shown as the mean ± s.d.
Extended Data Fig. 5 The NMR spectra of P(C6-Bn20) and P(C6-NPh20) at different pH conditions.
a, 1H NMR spectra of P(C6-Bn20) at pH 7.4, 7.0, 6.8 and 6.5. The diffusion ordered NMR spectra of P(C6-Bn20) at pH 7.4 (b) and 6.8 (c) in D2O. d, 1H NMR spectra of P(C6-NPh20) at pH 7.4, 7.0, 6.8 and 6.5. The diffusion ordered NMR spectra of P(C6-NPh20) at pH 7.4 (e) and 6.8 (f) in D2O.
Extended Data Fig. 6 The cytotoxicity of P(C6-Bn20) at pH 6.8 when incubated at 4 °C, with inhibitors, or with FBS.
a, The concentration-dependent cytotoxicity of P(C6-Bn20) against Panc02 cells when incubated at 37 °C or 4 °C at pH 7.4 or 6.8. b, The concentration-dependent cytotoxicity of P(C6-Bn20) against Panc02 cells at pH 6.8 after the pretreatment with methyl-β-cyclodextrin (M-β-C, a caveolae-dependent endocytosis inhibitor, 50 μM), wortmannin (Wor, a macropinocytosis inhibitor, 50 nM), deoxyglucose (Deo, an energy-dependent endocytosis inhibitor, 50 mM), or chlorpromazine (Chl, a clathrin-dependent endocytosis inhibitor, 10 μg ml-1) for 0.5 h. c, The concentration-dependent cytotoxicity of P(C6-Bn20) against Panc02 cells at pH 6.8 after the pretreatment with VX-765 (a caspase 1/4 inhibitor, 40 µM), Z-VAD (a pan-caspase inhibitor, 40 µM), ferrostatin (Fer, a ferroptosis inhibitor, 1 µM) or Nec-1 (a necrosis inhibitor, 100 µM) for 0.5 h. d, The diameter of P(C6-Bn20) (1.0 mg ml-1) as a function of time when incubated in PBS with 10% FBS at 37 °C under gentle stirring. e, The concentration-dependent cytotoxicity of P(C6-Bn20) against Panc02 cells at pH 6.8 when preincubated with or without 10% FBS for 24 h at pH 7.4. In a-e, n = 3 biologically independent samples, and the data are presented as the mean ± s.d.
Extended Data Fig. 7 The anti-tumour efficacy of P(C6-Bn20) against CT26 colon tumour model and B16-F10 melanoma model.
a, Treatment schedule for the murine CT26 colon tumour model. BALB/c mice were inoculated with CT26 cells on day 0. The tumour-bearing mice were then treated with PBS or P(C6-Bn20) (50 mg kg-1) daily from day 15 to 20. b, The tumour growth inhibition study of CT26 tumours. Images (c) and weight (d) of excised CT26 tumours after treatment. e, The weight of mice during the treatments. f, Treatment schedule for the murine B16-F10 melanoma model. C57BL/6 mice were inoculated with B16-F10 cells on day 0. The tumour-bearing mice were then treated with PBS or P(C6-Bn20) (50 mg kg-1) daily from day 6 to 10. g, The tumour growth inhibition study of B16-F10 tumours. Images (h) and weight (i) of excised B16-F10 tumours after treatment. j, The weight of mice during the treatments. In b, d, e, g, i, j, data are shown as the mean ± s.e.m. (n = 7 independent mice in b, d, e; n = 6 independent mice in g, i, j). Statistical significance was determined by unpaired two-tailed Student’s t-test.
Extended Data Fig. 8 The pharmacokinetic behaviours, distribution, and in-vivo toxicity of P(C6-Bn20).
a, The pharmacokinetic behaviours of cy5 labelled P(C6-Bn20) following intravenous (i.v.) injection at a dose of 50 mg kg-1 into healthy ICR mice. b, The biodistribution of cy5 labelled P(C6-Bn20) after i.v. injection into Panc02 tumour-bearing C57BL/6 mice. c, H&E staining of normal tissues, including the heart, liver, spleen, lung, and kidney 24 h after one i.v. injection of PBS or P(C6-Bn20) (50 mg kg-1). The scale bar is 200 μm. d, The schedule for the evaluation of toxicity of P(C6-Bn20). PBS or P(C6-Bn20) (50 mg kg-1) was intravenously administrated every other day for 4 injections. The mice were sacrificed and serum samples were collected at 7, 14 and 28 days after the last injection for the detection of the concentrations of AST (e), ALT (f), Urea (g) and UA (h). In a, data are shown as the mean ± s.d. (n = 5 independent mice). In b, e-h, data are shown as the mean ± s.e.m. (n = 3 independent mice). Statistical significance was determined by unpaired two-tailed Student’s t-test. In c, the representative images from three independent mice were shown.
Supplementary information
Supplementary Information
Additional materials and methods and Supplementary Table 1, Schemes 1 and 2, and Fig. 1.
Supplementary Video 1
The distribution of Bn groups in the aggregation of P(C6-Bn20)l.
Supplementary Video 2
The distribution of Bn groups in the aggregation of P(C6-Bn20)h.
Supplementary Video 3
The distribution of H groups in the aggregation of P(C6-H20)l.
Supplementary Video 4
The distribution of H groups in the aggregation of P(C6-H20)h.
Supplementary Video 5
Video of pulling a P(C6-Bn20)l molecule away from an aggregated particle of 8 molecules.
Supplementary Video 6
Video of pulling a P(C6-Bn20)h molecule away from an aggregated particle of 8 molecules.
Supplementary Video 7
Video of pulling a P(C6-H20)l molecule away from an aggregated particle of 8 molecules.
Supplementary Video 8
Video of pulling a P(C6-H20)h molecule away from an aggregated particle of 8 molecules.
Supplementary Video 9
Video of pulling P(C6-Bn20)h through the membrane until complete polymer insertion.
Supplementary Video 10
Video of pulling P(C6-H20)h through the membrane until complete polymer insertion.
Supplementary Video 11
Video of the interactions between the P(C6-Bn20)-cy5 and Panc02GFP/mCherry cells at pH 7.4.
Supplementary Video 12
Video of the interactions between the P(C6-Bn20)-cy5 and Panc02GFP/mCherry cells at pH 6.8.
Supplementary Video 13
Video of Panc02 cells incubated with P(C6-Bn20)-cy5, Annexin V-FITC and PI at pH 6.8.
Source data
Source Data Fig. 1
Unprocessed gels for Fig. 3i.
Rights and permissions
About this article
Cite this article
Liu, M., Huang, L., Zhang, W. et al. A transistor-like pH-sensitive nanodetergent for selective cancer therapy. Nat. Nanotechnol. 17, 541–551 (2022). https://doi.org/10.1038/s41565-022-01085-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-022-01085-5
This article is cited by
-
Mesoporous nanoperforators as membranolytic agents via nano- and molecular-scale multi-patterning
Nature Communications (2024)
-
A cytotoxic T cell inspired oncolytic nanosystem promotes lytic cell death by lipid peroxidation and elicits antitumor immune responses
Nature Communications (2023)
-
Recent advances of bioresponsive polymeric nanomedicine for cancer therapy
Nano Research (2023)