Abstract
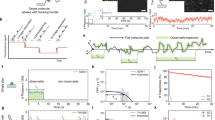
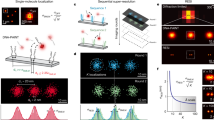
Single-molecule fluorescence resonance energy transfer (smFRET) is widely used to investigate dynamic (bio)molecular interactions occurring over distances of up to 10 nm. Recent advances in super-resolution methods have brought their spatiotemporal resolution closer towards the smFRET regime. Although these methods do not suffer from the spatial restrictions of FRET, they only visualize one emitter at a time, thus making it difficult to capture fast dynamics of the interactions. Here we describe two approaches to overcome this limitation in pulsed-interleaved MINFLUX (pMINFLUX) microscopy by using its intrinsic fluorescence lifetime information. First we combine pMINFLUX with smFRET, which enables tracking a FRET donor with nanometre precision while simultaneously determining its distance to a FRET acceptor, yielding the acceptor position by multilateration. Second, we developed pMINFLUX lifetime multiplexing—a method that simultaneously tracks two fluorophores with similar spectral properties but distinct fluorescence lifetimes—to extend co-localized tracking beyond the FRET range. We demonstrate applications on DNA origami systems as well as by imaging the paratopes of an antibody with precision better than 2 nm, paving the way for nanometre precise co-localized tracking for inter-dye distances between 4 nm and 100 nm, and closing the resolution gap between smFRET and co-tracking.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data are available from the corresponding author on reasonable request.
Code availability
Codes used in this study are available from the corresponding author on reasonable request.
References
Lerner, E. et al. Toward dynamic structural biology: two decades of single-molecule Förster resonance energy transfer. Science 359, eaan1133 (2018).
Robinson, A. & van Oijen, A. M. Bacterial replication, transcription and translation: mechanistic insights from single-molecule biochemical studies. Nat. Rev. Microbiol. 11, 303–315 (2013).
Stracy, M., Uphoff, S., Garza de Leon, F. & Kapanidis, A. N. In vivo single-molecule imaging of bacterial DNA replication, transcription, and repair. FEBS Lett. 588, 3585–3594 (2014).
Alhadid, Y. et al. Studying transcription initiation by RNA polymerase with diffusion-based single-molecule fluorescence. Protein Sci. 26, 1278–1290 (2017).
Blanchard, S. C. Single-molecule observations of ribosome function. Curr. Opin. Struct. Biol. 19, 103–109 (2009).
Munro, J. B., Vaiana, A., Sanbonmatsu, K. Y. & Blanchard, S. C. A new view of protein synthesis: mapping the free energy landscape of the ribosome using single-molecule FRET. Biopolymers 89, 565–577 (2008).
Gambin, Y. & Deniz, A. A. Multicolor single-molecule FRET to explore protein folding and binding. Mol. Biosyst. 6, 1540–1547 (2010).
Schuler, B. & Eaton, W. A. Protein folding studied by single-molecule FRET. Curr. Opin. Struct. Biol. 18, 16–26 (2008).
Sasmal, D. K., Pulido, L. E., Kasal, S. & Huang, J. Single-molecule fluorescence resonance energy transfer in molecular biology. Nanoscale 8, 19928–19944 (2016).
Weikl, T. R. & Paul, F. Conformational selection in protein binding and function. Protein Sci. 23, 1508–1518 (2014).
Sako, Y., Minoghchi, S. & Yanagida, T. Single-molecule imaging of EGFR signalling on the surface of living cells. Nat. Cell Biol. 2, 168–172 (2000).
Winckler, P. et al. Identification and super-resolution imaging of ligand-activated receptor dimers in live cells. Sci. Rep. 3, 2387 (2013).
Asher, W. B. et al. Single-molecule FRET imaging of GPCR dimers in living cells. Nat. Methods 18, 397–405 (2021).
Mieskes, F., Ploetz, E., Wehnekamp, F., Rat, V. & Lamb, D. C. Multicolor 3D orbital tracking. Small 19, e2204726 (2023).
Balzarotti, F. et al. Nanometer resolution imaging and tracking of fluorescent molecules with minimal photon fluxes. Science 355, 606–612 (2017).
Masullo, L. A. et al. Pulsed Interleaved MINFLUX. Nano Lett. 21, 840–846 (2021).
Gwosch, K. C. et al. MINFLUX nanoscopy delivers 3D multicolor nanometer resolution in cells. Nat. Methods 17, 217–224 (2020).
Weber, M. et al. MINSTED fluorescence localization and nanoscopy. Nat. Photonics 15, 361–366 (2021).
Masullo, L. A. et al. An alternative to MINFLUX that enables nanometer resolution in a confocal microscope. Light Sci. Appl. 11, 199 (2022).
Stefani, F. D. Tracking nanoscopic motion with minima of light. Nat. Photon. 17, 552–553 (2023).
Deguchi, T. et al. Direct observation of motor protein stepping in living cells using MINFLUX. Science 379, 1010–1015 (2023).
Eilers, Y., Ta, H., Gwosch, K. C., Balzarotti, F. & Hell, S. W. MINFLUX monitors rapid molecular jumps with superior spatiotemporal resolution. Proc. Natl Acad. Sci. USA 115, 6117–6122 (2018).
Ostersehlt, L. M. et al. DNA-PAINT MINFLUX nanoscopy. Nat. Methods 19, 1072–1075 (2022).
Zähringer, J. et al. Combining pMINFLUX, graphene energy transfer and DNA-PAINT for nanometer precise 3D super-resolution microscopy. Light Sci. Appl. 12, 70 (2023)
Betzig, E. Proposed method for molecular optical imaging. Opt. Lett. 20, 237–239 (1995).
Heilemann, M. et al. High-resolution colocalization of single dye molecules by fluorescence lifetime imaging microscopy. Anal. Chem. 74, 3511–3517 (2002).
Bückers, J., Wildanger, D., Vicidomini, G., Kastrup, L. & Hell, S. W. Simultaneous multi-lifetime multi-color STED imaging for colocalization analyses. Opt. Express 19, 3130–3143 (2011).
Schröder, T. et al. Shrinking gate fluorescence correlation spectroscopy yields equilibrium constants and separates photophysics from structural dynamics. Proc. Natl Acad. Sci. USA 120, e2211896120 (2023).
Mahalingam, M. et al. Structural mapping of divergent regions in the type 1 ryanodine receptor using fluorescence resonance energy transfer. Structure 22, 1322–1332 (2014).
Svensson, B. et al. FRET-based trilateration of probes bound within functional ryanodine receptors. Biophys. J. 107, 2037–2048 (2014).
Muschielok, A. et al. A nano-positioning system for macromolecular structural analysis. Nat. Methods 5, 965–971 (2008).
Muschielok, A. & Michaelis, J. Application of the nano-positioning system to the analysis of fluorescence resonance energy transfer networks. J. Phys. Chem. B 115, 11927–11937 (2011).
Qu, X., Wu, D., Mets, L. & Scherer, N. F. Nanometer-localized multiple single-molecule fluorescence microscopy. Proc. Natl Acad. Sci. USA 101, 11298–11303 (2004).
Ram, S., Ward, E. S. & Ober, R. J. Beyond Rayleigh’s criterion: a resolution measure with application to single-molecule microscopy. Proc. Natl Acad. Sci. USA 103, 4457–4462 (2006).
Gordon, M. P., Ha, T. & Selvin, P. R. Single-molecule high-resolution imaging with photobleaching. Proc. Natl Acad. Sci. USA 101, 6462–6465 (2004).
Pfeiffer, M. et al. Single antibody detection in a DNA origami nanoantenna. iScience 24, 103072 (2021).
Shaw, A. et al. Binding to nanopatterned antigens is dominated by the spatial tolerance of antibodies. Nat. Nanotechnol. 14, 184–190 (2019).
Zhang, P. et al. Capturing transient antibody conformations with DNA origami epitopes. Nat. Commun. 11, 3114 (2020).
Digman, M. A., Caiolfa, V. R., Zamai, M. & Gratton, E. The phasor approach to fluorescence lifetime imaging analysis. Biophys. J. 94, L14–L16 (2008).
Hedley, G. J. et al. Picosecond time-resolved photon antibunching measures nanoscale exciton motion and the true number of chromophores. Nat. Commun. 12, 1327 (2021).
Thiele, J. C. et al. Confocal fluorescence-lifetime single-molecule localization microscopy. ACS Nano 14, 14190–14200 (2020).
Oleksiievets, N. et al. Wide-field fluorescence lifetime imaging of single molecules. J. Phys. Chem. A 124, 3494–3500 (2020).
Slenders, E. & Vicidomini, G. ISM-FLUX: MINFLUX with an array detector. Phys. Rev. Res. 5, 023033 (2023).
Carter, N. J. & Cross, R. A. Mechanics of the kinesin step. Nature 435, 308–312 (2005).
Timney, B. L. et al. Simple rules for passive diffusion through the nuclear pore complex. J. Cell Biol. 215, 57–76 (2016).
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009).
Wagenbauer, K. F. et al. How we make DNA origami. Chembiochem 18, 1873–1885 (2017).
Fischer, S. et al. Shape and interhelical spacing of DNA origami nanostructures studied by small-angle X-ray scattering. Nano Lett. 16, 4282–4287 (2016).
Ke, Y. et al. Multilayer DNA origami packed on a square lattice. J. Am. Chem. Soc. 131, 15903–15908 (2009).
Nickels, P. C. et al. Molecular force spectroscopy with a DNA origami-based nanoscopic force clamp. Science 354, 305–307 (2016).
Wang, S. et al. Three-photon luminescence of gold nanorods and its applications for high contrast tissue and deep in vivo brain imaging. Theranostics 5, 251–266 (2015).
Cordes, T., Vogelsang, J. & Tinnefeld, P. On the mechanism of Trolox as antiblinking and antibleaching reagent. J. Am. Chem. Soc. 131, 5018–5019 (2009).
Acknowledgements
P.T. is grateful for support from the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation)—Project-ID 201269156—SFB 1032 Project A13 and funding by the Bavarian Ministry of Science and the Arts through the ONE MUNICH Project ‘Munich Multiscale Biofabrication’. P.T. is indebted to the DFG for funding via grant nos. 470075523 and 459594986, and Germany’s Excellence Strategy (grant no. EXC 2089/1-390776260). We thank P. Luna, P. Xaverl and T. Charly for their support with the measurements and data analysis, and A. Kardinal, H. Lipfert, S. Steger and M. Ehrl for laboratory upkeep.
Author information
Authors and Affiliations
Contributions
F.C., J.Z., T.S., F.S., F.D.S. and P.T developed the concept. F.C., J.Z. and J.B. designed and prepared samples. M.P. designed and prepared antibody samples. P.S. designed and prepared lipid bilayer samples. F.C. and J.Z. performed and analysed measurements. P.T. supervised the project. All authors have written, read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Photonics thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.44
Supplementary information
Supplementary Information
Supplementary Figs. 1–16 and Sections 1–14.
Supplementary Video 1
The time evolution of the localizations with the corresponding FRET radii and the subsequent localization of the FRET acceptor.
Supplementary Video 2
The time evolution of the co-localization of two pointer systems by lifetime multiplexing.
Supplementary Video 3
The time evolution of the orientation of the lipid raft measured by the co-localization of two fluorophores by lifetime multiplexing.
Supplementary Data 1
Supplementary Tables 2–9. Sequences of the DNA oligonucleotides used for DNA origami folding.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cole, F., Zähringer, J., Bohlen, J. et al. Super-resolved FRET and co-tracking in pMINFLUX. Nat. Photon. (2024). https://doi.org/10.1038/s41566-024-01384-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41566-024-01384-4