Abstract
The transport of polymers across nanoscale pores underpins many biological processes, such as the ejection of bacteriophage DNA into a host cell and the transfer of genes between bacteria. The movement of polymers into and out of confinement is also the basis for a wide range of sensing technologies used for single-molecule detection and sequencing. Acquiring an accurate understanding of the translocation dynamics is an essential step in the quantitative analysis of polymer structure, including the localization of binding sites or sequences. Here we use synthetic nanopores and nanostructured DNA molecules to directly measure the velocity profile of driven polymer translocation through synthetic nanopores. Our results reveal a two-stage behaviour in which the translocation initially slows with time before accelerating close to the end of the process. We also find distinct local velocity correlations as the DNA polymer chain passes through the nanopore. Brownian dynamics simulations show that the two-stage behaviour is associated with tension propagation, with correlations arising from the random-walk conformation in which the DNA begins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Source data are provided with this paper. Raw data of ionic current values for translocations together with a table summarizing all the nanopores used are available at https://doi.org/10.17863/CAM.69631.
Code availability
The code used for data collection and analysis and the code used for simulation analysis are available upon request from the corresponding author.
References
Pennisi, E. Search for pore-fection. Science 336, 534–537 (2012).
Restrepo-Pérez, L., Joo, C. & Dekker, C. Paving the way to single-molecule protein sequencing. Nat. Nanotechnol. 13, 786–796 (2018).
Marbach, S., Dean, D. S. & Bocquet, L. Transport and dispersion across wiggling nanopores. Nat. Phys. 14, 1108–1113 (2018).
Branton, D. et al. The potential and challenges of nanopore sequencing. Nat. Biotechnol. 26, 1146–1153 (2008).
Muthukumar, M. Polymer Translocation (CRC Press, 2009).
Ghosal, S., Sherwood, J. D. & Chang, H. C. Solid-state nanopore hydrodynamics and transport. Biomicrofluidics 13, 011301 (2019).
Palyulin, V. V., Ala-Nissila, T. & Metzler, R. Polymer translocation: the first two decades and the recent diversification. Soft Matter 10, 9016–9037 (2014).
Kasianowicz, J. J., Brandin, E., Branton, D. & Deamer, D. W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl Acad. Sci. USA 93, 13770–13773 (1996).
Storm, A. J. et al. Fast DNA translocation through a solid-state nanopore. Nano Lett. 5, 1193–1197 (2005).
Ghosal, S. Effect of salt concentration on the electrophoretic speed of a polyelectrolyte through a nanopore. Phys. Rev. Lett. 98, 238104 (2007).
Grosberg, A. Y., Nechaev, S., Tamm, M. & Vasilyev, O. How long does it take to pull an ideal polymer into a small hole? Phys. Rev. Lett. 96, 228105 (2006).
Sakaue, T. Nonequilibrium dynamics of polymer translocation and straightening. Phys. Rev. E 76, 021803 (2007).
Li, J. & Talaga, D. S. The distribution of DNA translocation times in solid-state nanopores. J. Phys. Condens. Matter 22, 454129 (2010).
Mihovilovic, M., Hagerty, N. & Stein, D. Statistics of DNA capture by a solid-state nanopore. Phys. Rev. Lett. 110, 028102 (2013).
Chen, P. et al. Probing single DNA molecule transport using fabricated nanopores. Nano Lett. 4, 2293–2298 (2004).
Carson, S., Wilson, J., Aksimentiev, A. & Wanunu, M. Smooth DNA transport through a narrowed pore geometry. Biophys. J. 107, 2381–2393 (2014).
Panja, D., Barkema, G. T. & Kolomeisky, A. B. Through the eye of the needle: recent advances in understanding biopolymer translocation. J. Phys. Condens. Matter 25, 413101 (2013).
Ikonen, T., Bhattacharya, A., Ala-Nissila, T. & Sung, W. Unifying model of driven polymer translocation. Phys. Rev. E 85, 051803 (2012).
Wanunu, M. Nanopores: a journey towards DNA sequencing. Phys. Life Rev. 9, 125–158 (2012).
Singer, A., Rapireddy, S., Ly, D. H. & Meller, A. Electronic barcoding of a viral gene at the single-molecule level. Nano Lett. 12, 1722–1728 (2012).
Plesa, C. et al. Velocity of DNA during translocation through a solid-state nanopore. Nano Lett. 15, 732–737 (2015).
Bell, N. A. W. & Keyser, U. F. Specific protein detection using designed DNA carriers and nanopores. J. Am. Chem. Soc. 137, 2035–2041 (2015).
Wanunu, M., Sutin, J., McNally, B., Chow, A. & Meller, A. DNA translocation governed by interactions with solid-state nanopores. Biophys. J. 95, 4716–4725 (2008).
Katkar, H. H. & Muthukumar, M. Role of non-equilibrium conformations on driven polymer translocation. J. Chem. Phys. 148, 024903 (2018).
Meller, A., Nivon, L. & Branton, D. Voltage-driven DNA translocations through a nanopore. Phys. Rev. Lett. 86, 3435–3438 (2001).
Muthukumar, M. Polymer translocation through a hole. J. Chem. Phys. 111, 10371–10374 (1999).
Sung, W. & Park, P. J. Polymer translocation through a pore in a membrane. Phys. Rev. Lett. 77, 783–786 (1996).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Bell, N. A. W. & Keyser, U. F. Digitally encoded DNA nanostructures for multiplexed, single-molecule protein sensing with nanopores. Nat. Nanotechnol. 11, 645–651 (2016).
Kwok, H., Briggs, K. & Tabard-Cossa, V. Nanopore fabrication by controlled dielectric breakdown. PLoS ONE 9, e92880 (2014).
Chen, K. et al. Ionic current-based mapping of short sequence motifs in single DNA molecules using solid-state nanopores. Nano Lett. 17, 5199–5205 (2017).
Saito, T. & Sakaue, T. Cis–trans dynamical asymmetry in driven polymer translocation. Phys. Rev. E 88, 042606 (2013).
Dubbeldam, J. L. A., Rostiashvili, V. G. & Vilgis, T. A. Driven translocation of a polymer: role of pore friction and crowding. J. Chem. Phys. 141, 124112 (2014).
Sarabadani, J. et al. Driven translocation of a semi-flexible polymer through a nanopore. Sci. Rep. 7, 7423 (2017).
Ghosal, S. Electrophoresis of a polyelectrolyte through a nanopore. Phys. Rev. E 74, 041901 (2006).
Dubbeldam, J. L. A., Rostiashvili, V. G., Milchev, A. & Vilgis, T. A. Driven translocation of a polymer: fluctuations at work. Phys. Rev. E 87, 032147 (2013).
de Haan, H. W., Sean, D. & Slater, G. W. Reducing the variance in the translocation times by prestretching the polymer. Phys. Rev. E 98, 022501 (2018).
Sarabadani, J., Ikonen, T. & Ala-Nissila, T. Iso-flux tension propagation theory of driven polymer translocation: the role of initial configurations. J. Chem. Phys. 141, 214907 (2014).
Saito, T. & Sakaue, T. Dynamical diagram and scaling in polymer driven translocation. Eur. Phys. J. E 34, 135 (2011).
Forrey, C. & Muthukumar, M. Langevin dynamics simulations of genome packing in bacteriophage. Biophys. J. 91, 25–41 (2006).
Forrey, C. & Muthukumar, M. Langevin dynamics simulations of ds-DNA translocation through synthetic nanopores. J. Chem. Phys. 127, 015102 (2007).
Sarabadani, J. & Ala-Nissila, T. Theory of pore-driven and end-pulled polymer translocation dynamics through a nanopore: an overview. J. Phys. Condens. Matter 30, 274002 (2018).
Lu, B., Albertorio, F., Hoogerheide, D. P. & Golovchenko, J. A. Origins and consequences of velocity fluctuations during DNA passage through a nanopore. Biophys. J. 101, 70–79 (2011).
Adhikari, R. & Bhattacharya, A. Driven translocation of a semi-flexible chain through a nanopore: a Brownian dynamics simulation study in two dimensions. J. Chem. Phys. 138, 204909 (2013).
Ikonen, T., Bhattacharya, A., Ala-Nissila, T. & Sung, W. Influence of non-universal effects on dynamical scaling in driven polymer translocation. J. Chem. Phys. 137, 085101 (2012).
Saito, T. & Sakaue, T. Process time distribution of driven polymer transport. Phys. Rev. E 85, 061803 (2012).
Briggs, K. et al. DNA translocations through nanopores under nanoscale preconfinement. Nano Lett. 18, 660–668 (2018).
Kumar Sharma, R., Agrawal, I., Dai, L., Doyle, P. S. & Garaj, S. Complex DNA knots detected with a nanopore sensor. Nat. Commun. 10, 4473 (2019).
Mondal, D. & Muthukumar, M. Stochastic resonance during a polymer translocation process. J. Chem. Phys. 144, 144901 (2016).
Sarabadani, J., Ikonen, T. & Ala-Nissila, T. Theory of polymer translocation through a flickering nanopore under an alternating driving force. J. Chem. Phys. 143, 074905 (2015).
Chen, K. et al. Digital data storage using DNA nanostructures and solid-state nanopores. Nano Lett. 19, 1210–1215 (2019).
Venkatesan, B. M. & Bashir, R. Nanopore sensors for nucleic acid analysis. Nat. Nanotechnol. 6, 615–624 (2011).
Bell, N. A. W., Muthukumar, M. & Keyser, U. F. Translocation frequency of double-stranded DNA through a solid-state nanopore. Phys. Rev. E 93, 022401 (2016).
Li, J., Gershow, M., Stein, D., Brandin, E. & Golovchenko, J. A. DNA molecules and configurations in a solid-state nanopore microscope. Nat. Mater. 2, 611–615 (2003).
Storm, A., Chen, J., Zandbergen, H. & Dekker, C. Translocation of double-strand DNA through a silicon oxide nanopore. Phys. Rev. E 71, 051903 (2005).
Acknowledgements
This work was supported by an ERC consolidator grant (no. 647144) for N.A.W.B., K.C. and U.F.K. and a National Institute of Health grant (no. 5R01HG002776-15) for I.J. and M.M. N.E. acknowledges funding from the EPSRC; Cambridge Trust; and Trinity Hall, Cambridge.
Author information
Authors and Affiliations
Contributions
N.A.W.B., K.C. and U.F.K. designed the experiments. K.C., N.E. and N.A.W.B. performed the experiments. M.M. and I.J. performed the simulations. M.M., I.J. and N.A.W.B. analysed the simulation results. The paper was written through contributions of all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Physics thanks Vincent Tabard-Cossa and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–34 and Tables 1–5.
Supplementary Video 1
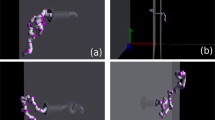
Fifteen example simulations for translocations through a membrane geometry nanopore with the position of the tension front highlighted in green. The two frames show orthogonal viewpoints of the 3D simulation.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Rights and permissions
About this article
Cite this article
Chen, K., Jou, I., Ermann, N. et al. Dynamics of driven polymer transport through a nanopore. Nat. Phys. 17, 1043–1049 (2021). https://doi.org/10.1038/s41567-021-01268-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41567-021-01268-2