Abstract
Owing to advances in radiotherapy, the physical properties of radiation can be optimized to enable individualized treatment; however, optimization is rarely based on biological properties and, therefore, treatments are generally planned with the assumption that all tumours respond similarly to radiation. Radiation affects multiple cellular pathways, including DNA damage, hypoxia, proliferation, stem cell phenotype and immune response. In this Review, we summarize the effect of these pathways on tumour responses to radiotherapy and the current state of research on genomic classifiers designed to exploit these variations to inform treatment decisions. We also discuss whether advances in genomics have generated evidence that could be practice changing and whether advances in genomics are now ready to be used to guide the delivery of radiotherapy alone or in combination.
Key points
-
The biological effects of radiation are mediated by a complex network of signalling pathways, and assessing these effects can help to classify tumours as radiosensitive or radioresistant.
-
Advances in genomics could be used to guide the delivery of radiotherapy alone and in combination; the commercialization of genomic-based tools will be important to drive their implementation.
-
Developments from the past 20 years have enabled unprecedented high-throughput analyses and rapid profiling of RNA and DNA that are currently embedded in clinical laboratories.
-
The community needs to learn from past failures to translate radiobiology biomarkers into the clinic and improve collaborations, focusing on standardizing methods and performing multicentre validation.
-
RNA-based signatures are among the most advanced tools currently available: a ten-gene signature of cellular radiosensitivity should be available soon, and hypoxia-related signatures need further testing in clinical trials.
-
The development of DNA-based signatures is currently in progress, and areas that need further study include the role of somatic mutations in DNA damage response genes that affect radiosensitivity.
Similar content being viewed by others
Introduction
Radiotherapy is a key treatment modality in oncology that contributes to ~40% of cures1, and its importance is underpinned by the fact that the technology for delivering radiation is continually improving. Contemporary volumetric arc techniques, supported by 3D2 or 4D3 image guidance, produce continuous beams of radiation and enable highly conformal tumour doses4,5,6. Although certain technical aspects of radiotherapy (such as the position of the multileaf collimators, target volume delineation and application of dose constraints to target volumes and vital structures at risk) are optimized for each patient, all patients with a given cancer type are prescribed the same radiation schedule and dose. Furthermore, radiation doses to target tumour volumes can be compromised because dose prescriptions are based on empirical dose constraints to non-malignant tissues that were defined decades ago.
Contemporary patient selection for radiotherapy regimens is based on factors related to the patient (for example, age, performance status or presence of comorbidity) and the tumour (such as size, stage and histological subtype); the use of information related to tumour biology is limited. For example, the methylation status of MGMT, routinely tested in patients with glioblastoma, is a predictive biomarker of response to radiotherapy, but currently this information does not influence radiotherapy dose prescription7. By contrast, genomic information is currently used to guide decisions on the use of systemic therapies (for example, the Oncotype DX tool is used to select patients with breast cancer for adjuvant chemotherapy)8. In this regard, radiation oncology lags behind despite the fact that not all patients benefit from radiotherapy in the definitive (in which patients might receive other therapies, such as surgery) or adjuvant (in which the use of radiation could be avoided in some patients) settings. Identifying the genomic features associated with response to radiation to personalize the use of radiotherapy would not only reduce toxicities and improve quality of life in long-term cancer survivors but also enable individualized dose prescription.
The development of tests to predict tumour radiosensitivity that would enable the delivery of biologically optimized radiotherapy is currently an area of considerable research interest. This research is important because it would guide the rational use of radiotherapy over surgery for cancers in which both modalities are equally effective (such as some prostate and bladder cancers) and avoid its use in settings in which patients are unlikely to derive any benefit (such as low-risk breast cancers). Other potential benefits include individualization of fractionation schedules on the basis of predicted radioresponsiveness and, importantly, personalization of combination treatments (with novel radiosensitizers, cytotoxic agents and/or immunotherapies). In this Review, we summarize the mechanism of action of photon radiotherapy to highlight key genes and pathways that mediate its effects, describe the profiling approaches used to identify the radioresponsive genome and discuss whether the currently available evidence is practice changing and ready for clinical implementation.
Mechanism of action of radiotherapy
The effects of radiotherapy are mediated by a complex network of signalling pathways. Radiation acts in an indiscriminate manner by causing excitations and ionizations of atoms in the tissues to which beams are targeted to deposit the maximum amount of energy. Most molecules quickly revert to their natural state, but unrestored ionizations produce free radicals that damage tissue. DNA is considered the most important target of radiation and can be damaged directly or indirectly via ionization of surrounding water molecules. The spectrum of DNA lesions includes (in increasing order of lethality and decreasing order of frequency): base damage, single-strand breaks (SSBs), double-strand breaks (DSBs) and clustered DNA lesions (complex arrangements of SSBs and DSBs). Such lesions activate inter-related biological pathways, the complexity of which contrasts with the more-linear biological pathways that mediate the effects of most other anticancer agents (for example, the antimetabolite 5-fluorouracil). Radiation-induced damage initiates cellular pathways with various downstream consequences that include, among others, slowing cell cycle progression to maximize time for repair and attempting DNA damage repair (DDR), both leading to cell survival or death (Fig. 1).
Radiation damages tissues via two main mechanisms of action: direct damage to DNA and indirect damage via the formation of reactive oxygen species (ROS). a, A spectrum of lesions can be directly inflicted upon DNA, including single-strand breaks (SSBs) and double-strand breaks (DSBs). SSBs trigger base excision repair (BER) whereas DSBs are repaired by non-homologous end-joining (NHEJ), theta-mediated end-joining (TMEJ) or high-fidelity homologous recombination. Errors in DNA repair can lead to genomic instability. When DNA repair mechanisms are defective, several cytotoxic pathways can be activated, leading to apoptosis or permanent cell cycle arrest. EGFR-mediated signalling leads to activation of the RAS–RAF–MEK–ERK pathway, which promotes DNA repair and prevents apoptosis. b, Indirect damage mediated by the production of ROS results in promotion of apoptosis and release of pro-inflammatory cytokines that lead to the recruitment of inflammatory cells. CDK, cyclin-dependent kinase; CtIP, DNA endonuclease RBBP8; DAMP, damage-associated molecular pattern; DNA-PK, DNA-dependent protein kinase catalytic subunit; MRN, MRE11, RAD50 and nibrin; PARP, poly(ADP-ribose) polymerase; PNK, bifunctional polynucleotide phosphatase/kinase PNK; pol, polymerase; PRR, pattern-recognition receptor; XRCC, DNA repair protein XRCC.
Radiation exerts lethal and sublethal effects. Misrepaired sublethal effects can lead to genomic instability and increase the risk of further, radiation-induced malignancy. When cells undergo mitosis, unrepaired or misrepaired DNA damage produces chromosomal aberrations that can be asymmetrical (such as dicentric fragments that lead to mitotic catastrophe, which is lethal) or symmetrical (such as translocations that lead to genomic instability). In addition to direct DNA damage, radiation-induced reactive oxygen species (ROS) can promote apoptosis or senescence (depending on the extent of cell stress)9, stimulate proliferation by mimicking the binding of ligands to cell-surface receptors10,11 and co-ordinate signalling through inflammatory cytokines (such as IL-1 and TNF)12. Cell proliferation is important for repopulation of non-malignant tissues; however, the repopulation of tumours between radiation fractions can be detrimental. Tumour repopulation does not confer radioresistance to very short treatments (for example, stereotactic body radiation therapy), but can cause resistance to radiotherapy schedules that exceed 3–4 weeks. Radiation also has immunomodulatory effects that include the promotion of innate and adaptive immune responses13. Immune responses are implicated in possible abscopal effects (that is, the regression of a non-irradiated lesion after radiotherapy). Radiation can also have bystander effects on nearby cells by leading to the release of soluble factors that promote survival and proliferation and thus, radioresistance14. Sublethal radiation effects can activate the transcription of genes that support not only cell survival, DNA repair and proliferation, but also a cancer stem cell (CSC) phenotype. These signalling pathways have a protective effect on cancer cells by reducing cytotoxicity and promoting radioresistance11. Finally, no summary of the mechanism of action of radiation is complete without mentioning the importance of oxygen levels and hypoxia, which reduces radiosensitivity. In essence, radiation clearly has pleiotropic effects that we summarize in a simplified manner. All these pathways affect radiosensitivity and thus, the genes involved in them are candidate components of the radioresponsive tumour genome, which upon identification can aid the delivery of biologically driven precision radiotherapy.
DNA repair
SSBs are repaired by base excision repair following their recognition by poly(ADP-ribose) polymerase (PARP) and the subsequent recruitment of DNA polymerase β, AP endonuclease 1, DNA repair protein XRCC1, bifunctional polynucleotide phosphatase/kinase PNK, Flap endonuclease 1 and, finally, DNA ligases I and III15. DSBs are broadly repaired by either non-homologous end-joining (NHEJ), which can be error prone, or high-fidelity homologous recombination repair16. NHEJ, the dominant pathway for DSB repair, begins with the binding of the helicase XRCC5–XRCC6 dimer to DNA ends followed by the recruitment of DNA-dependent protein kinase catalytic subunit (DNA-PK), end processing by artemis and PNK followed by religation by DNA ligase IV17. In theta-mediated end-joining (TMEJ), an alternative form of NHEJ, the MRN complex (which involves the nucleases MRE11, RAD50 and nibrin) and DNA endonuclease RBBP8 (commonly known as CtIP) perform end resection, and PARP and FANCD2 recruit DNA polymerase θ, which has helicase and polymerase domains. Finally, end ligation is performed by DNA ligases I and III18. In homologous recombination repair, the slowest yet most accurate mechanism of DSB repair, CtIP and the MRN complex mediate end resection followed by coating of DNA single strands by RAD51 before locating and invading homologous regions on sister chromatids. Both strands are then used as a template for DNA synthesis by DNA polymerases (mainly polymerase δ)19.
Cell response and proliferation
The products of radiation-induced DNA repair can activate cytotoxic pathways to induce, among other processes, apoptosis or permanent cell cycle arrest. In addition to its role in DNA repair, the MRN complex recruits ATM, which activates p53 either directly or via stepwise phosphorylation through the ATM–CHEK2–p53 pathway20. Activated p53 induces the transcription of BBC3 and PMAIP1 to activate the pro-apoptotic protein BAX and the formation of a complex at the mitochondrial membrane that results in caspase activation and mitochondria-dependent apoptosis (also known as intrinsic apoptosis)21. p53 activation also contributes to cell cycle arrest, along with phosphorylation of the ATR–CHK1–cyclin-dependent kinase (CDK) 2 and WEE1–CDK1 pathways22. EGFR also has a role in the cellular response to radiotherapy: EGFR expression contributes to radioresistance by activating the RAS–RAF–MEK–ERK pathway23, which promotes homologous recombination repair and evasion of apoptosis.
Hypoxia signalling
The importance of hypoxia and its effects on radiosensitivity have been well described in numerous reviews24,25. Hypoxia-inducible factors (HIFs), the unfolded protein response (UPR) and mTOR all act in an integrated way, influencing both each other and common downstream signalling pathways that affect multiple processes involved in the response to hypoxia26. Hypoxia also causes replication stress (for example, stalled replication forks) and the accumulation of single-stranded DNA that activates DDR pathways27. Furthermore, hypoxia induces genomic instability that can drive cancer progression and radioresistance and is not targetable using hypoxia-specific radiosensitizers28,29.
HIFs are key oxygen sensors and the most widely studied mediators of responses to hypoxia. Under physiological conditions, HIF1α is produced continuously and has a half-life of <5 minutes30. As oxygen tension decreases, HIF1α stabilizes and forms heterodimers with HIF1β that translocate to the nucleus and bind to hypoxia-response elements in target gene promoters to stimulate transcription. Some of these genes are involved in metabolism (for example, GLUT1), control of intracellular acidity (CA9) and stimulation of angiogenesis (VEGF). Other members of the HIF family include EPAS1 and HIF3A, encoding HIF2α and HIF3α, respectively, which also heterodimerize with HIF1β but are not expressed in all cell types31.
Whereas HIF responses are induced at intermediate levels of hypoxia (<2% oxygen), the UPR is activated at lower levels of oxygen (<0.2%). The UPR restores endoplasmic and mitochondrial homeostasis to promote survival or, if not possible, induce cell death27. The master regulator of this response is HSPA5, which binds to and inactivates sensor proteins (including PERK, IRE1 and ATF4) under normoxic conditions but dissociates under hypoxia-induced endoplasmic reticulum stress. Following dissociation, PERK phosphorylation of the translation factor eIF2α inhibits protein synthesis, IRE1 maintains cellular functions (including those of the endoplasmic reticulum) that either promote cell survival or initiate apoptosis, and ATF4 induces the transcription of genes involved in apoptosis, autophagy and amino acid metabolism. Adaptive responses to hypoxia are mediated in part through a decrease in the kinase activity of the mTORC1 complex, a master regulator of protein translation32.
Hypoxia affects other aspects of the tumour microenvironment (TME) that are involved in radiosensitivity, such as lactate metabolism. In hypoxia, pyruvate is preferentially converted into lactic acid in a reaction catalysed by lactate dehydrogenase (LDH), which leads to acidification of the cytoplasm. Lactate efflux via monocarboxylate transporters (MCTs) is important to prevent this acidification and maintain cellular homeostasis. Strategies that target lactate metabolism (LDH and MCT1–4) can modulate radiosensitivity and therefore, have been tested in combination with radiation in preclinical studies and are being explored in early-phase clinical trials33.
Stem cell phenotype
Tumours contain varying proportions of CSCs that are pluripotent and inherently radioresistant, and their properties have been extensively reviewed elsewhere34. CSCs can be identified in tumours on the basis of their expression of cell-surface proteins including CD133, CD44, CD24, SLC3A2 and aldehyde dehydrogenase34,35. Studies in preclinical models have shown that radiation preferentially eliminates differentiated tumour cells while sparing CSCs. In addition, radiation upregulates SOX2, OCT4, KLF4 and NANOG, the transcription factors that lead to differentiated tumour cells acquiring a CSC phenotype36. Slowly proliferating and senescent CSCs can be ‘awakened’ by radiation, which stimulates TME-dependent pathways through which tumour cells acquire plasticity, or the ability to fluctuate between a differentiated and CSC phenotype, resulting in cell cycle re-entry and metastasis37,38.
By contrast with differentiated tumour cells, radioresistance is characterized in CSCs by reduced accumulation of radiation-induced DNA damage39, increased capacity for DDR via CHK1/2 (ref.40) and increased activation of anti-apoptotic signalling pathways41,42. Furthermore, CSCs also have lower levels of ROS and tend to overexpress ROS scavengers39, which limit the extent of ROS-dependent damage, and have genetic features suggestive of chromosomal instability43. CSCs are strongly associated with tumour hypoxia; indeed, hypoxia-associated factors, such as HIF1α, contribute to the development and maintenance of a CSC phenotype44.
Immune response
Tumour interactions with the host immune system are complex, and the avoidance of antitumour immunity is one of the hallmarks of cancer45. The TME is typically immunosuppressive, owing to the presence of immune cell types such as tumour-associated macrophages (TAMs) and regulatory T (Treg) cells12, with persistent pro-inflammatory features maintained by the presence of certain cytokines. The proportion of TAMs and Treg cells tends to increase after radiotherapy owing to their higher radioresistance compared with other immune cells in the TME46. Radiotherapy exerts both immunosuppressive and immunostimulatory effects on the TME47 (Fig. 2) and the contribution of immune pathways to its mechanism of action has been reviewed in detail elsewhere48,49,50.
Radiation has both immunostimulatory and immunosuppressive effects. a, Immunostimulatory effects are initiated when cell death from radiotherapy-induced DNA damage activates cGAS–STING pathway signalling, resulting in the generation of a range of damage-associated molecular patterns (DAMPs) that include calretinin, ATP and HMGB1 proteins. Calretinin binds to dendritic cells (DCs), generating an ‘eat-me’ signal that results in antigen processing and maturation of DCs and other antigen-presenting cells (APCs). APCs then present these antigens to CD8+ cytotoxic T cells, recruited via the interferon type I pathway, leading to antigen-specific T cell killing. b, Immunosuppressive effects are mediated by factors that include reactive oxygen species (ROS), HIF1α and PD-L1. ROS leads to the release of TGFβ, which inhibits antigen-specific T cell killing, as do other immune cells such as regulatory T (Treg) cells, cancer-associated fibroblasts (CAFs), tumour-associated macrophages (TAMs) and myeloid-derived suppressor cells (MDSCs). RLR, RIG I-like receptor.
Immunogenic cell death following radiation-induced DNA damage stimulates several processes, including recruitment of circulating immune cells, activation of inflammatory cytokine signalling (via the cGAS–STING pathway), generation of a range of damage-associated molecular patterns (DAMPs; including calretinin, ATP and HMGB1 proteins) and activation of pattern-recognition receptors (including Toll-like receptors and c-type lectin receptors)12. This response leads to antigen generation and presentation via antigen-presenting cells (for example, dendritic cells) and subsequent activation of CD8+ cytotoxic T cells and antigen-specific cell cytotoxicity50,51. Several immune cell types in the TME promote antitumour immunity (such as CD3+ T cells and CD45RO+ memory T cells52), and cytotoxic CD8+ T cells are particularly important in this regard; indeed, high intratumoural densities of CD8+ T cells are associated with favourable outcomes following radiotherapy in multiple cancer types12,53,54.
The immunosuppressive effects of radiotherapy inhibit antigen-specific cell cytotoxicity: the production of ROS after radiation leads to the release of TGFβ, which inhibits effector T cell function. Radiation-induced upregulation of HIF1α increases the production of CXCL12, VEGF and FOXP3, which activate Treg cells, myeloid-derived suppressor cells and cancer-associated fibroblasts55. Increases in the numbers of these cells promotes the secretion of pro-inflammatory cytokines and chemokines, and the polarization of macrophages and monocytes from a tumour-suppressive (M1) to a tumour-promoting (M2) phenotype49. Signalling through PD-1–PD-L1 and CTLA4 attenuates the immune responses elicited by radiotherapy56.
Measuring radiosensitivity
Many biological factors affect tumour radiosensitivity, including cell-intrinsic radiosensitivity, hypoxia, the ability to undergo accelerated repopulation and enrichment in CSC numbers; immune responses are emerging as another important factor. The terms radiosensitivity and radioresponsiveness are both used to refer to the overall effect of multiple factors — in this Review we use radiosensitivity to refer to both the overall effect and cell-intrinsic radiosensitivity. Both terms are loosely defined as the relative susceptibility of cells, tissues or organisms to the effects of ionizing radiation. Tumour radiosensitivity is typically assessed in terms of the classical ‘four Rs’ of radiotherapy: repair of DNA damage, reassortment (or the redistribution of cells previously in radioresistant phases into more-radiosensitive phases after each radiation fraction), tumour repopulation and re-oxygenation57. For example, human papillomavirus (HPV)-associated carcinomas of the oropharynx or uterine cervix are considered to be radiosensitive owing to an impaired DDR, a tendency towards being in radiosensitive phases of the cell cycle (G2/M arrest), low rates of tumour repopulation (as a result of low numbers of CSCs) and generally a non-hypoxic TME58. Cell-intrinsic radiosensitivity is genetically determined, underpinned by alterations in genes involved in DDR. For example, an individual with homozygous mutations in ATM has an approximately threefold increase in radiosensitivity above the average at the cell, tissue and organism levels. This extreme cell-intrinsic radiosensitivity also translates into increased tumour radiosensitivity. Mutations in TP53 can increase cellular radioresistance59, but radiotherapy should be avoided in individuals with germline homozygous TP53 mutations owing to radiosusceptibility (a high predisposition to radiation-induced cancers) and a resultant increased risk of second cancers60.
Some traditional pathology techniques remain valid methods of gauging tumour radiosensitivity. For example, haematoxylin and eosin staining can be used to identify radiosensitive (for example, seminomas) or radioresistant (for example, gliomas) tumours and immunohistochemistry (IHC) of tumour p16 is used as a surrogate biomarker of HPV positivity (and thus, radiosensitivity)61. More-advanced pathology techniques (such as DNA methylome profiling), are currently used for tumour classification62 but not yet to guide radiotherapy dose prescription in the clinic.
Tumour cell-intrinsic radiosensitivity
Interest in measuring tumour cell-intrinsic radiosensitivity stems from the seminal work of Fertil and Malaise in the 1980s63. These researchers analysed 101 published radiation survival curves (produced using clonogenic assays) for cell lines derived from various human tumour types and found that parameters that reflect the initial cellular response to radiation (such as surviving fraction at 2 Gy (SF2)) are the measures that better correlate with the clinical radiosensitivity of different types of human tumour (classified empirically on the basis of clinical experience)64. Strong evidence from other preclinical studies also supports the fact that cell-intrinsic radiosensitivity is a highly important component of local tumour control65. This initial evidence generated interest in measuring cellular radiosensitivity using functional assays to predict radioresponsiveness66. Several studies were conducted that had inconsistent findings67. Although some interest in identifying better functional assays to measure the radiosensitivity of tumour cells (such as ex vivo γH2AX assays68) remains, contemporary research generally involves exploration of biomarkers at the protein, RNA and gene levels using samples obtained from diagnostic biopsies.
Protein biomarkers
Many studies have assessed the expression of proteins involved in DDR in tumours as potential biomarkers of radiosensitivity69,70,71. The results of such studies continue to be published, although lack of coordinated effort and validation work hampers progress in this area. Here, we illustrate the challenge of using MRE11 as a biomarker, but an extensive review of all studies of DDR proteins is beyond the scope of this Review.
In two retrospective cohorts of patients with bladder cancer, high levels of tumour MRE11 expression on IHC were associated with improved cancer-specific survival in patients who received radiotherapy but not in those who underwent surgery72,73. This finding seems biologically counterintuitive (given that MRE11 contributes to DNA DSB repair and thus, radioresistance), although MRE11 expression might also correlate with genomic instability. These findings were not replicated in a subsequent multicentre study that attempted to validate an MRE11 IHC assay, perhaps owing to poor intercentre scoring agreement74. In a retrospective cohort of 55 patients with rectal cancer who received neoadjuvant radiotherapy, the presence of high levels of MRN on IHC in resection specimens was associated with inferior overall survival (OS) on multivariable analysis (HR 4.2, 95% CI 0.97–18.19; P = 0.045)69. In a series of 224 patients with early-stage breast cancer, weak staining for MRN proteins was associated with adverse pathological features, such as oestrogen receptor negativity and poor tumour differentiation. Furthermore, patients with moderate or strong expression of proteins from the MRN complex on IHC derived more benefit from adjuvant radiotherapy than those with low MRN expression (RR 0.27, 95% CI 0.10–0.72; P = 0.009)75. The same group assessed the expression of BRCA1/2 and RAD51 in the same cohort: patients with limited expression of these proteins on IHC had increased local recurrence (RR 3.2, 95% CI 1.48–6.88; P = 0.003) and derived increased benefit from radiotherapy (RR 0.31, 95% CI 0.14–0.70; P = 0.005) compared with those with high levels of expression76. Overall, these contrasting findings might reflect differences in the prognostic and predictive value of proteins from the MRN complex in various cancer types and illustrate some of the challenges associated with developing an IHC-based test for clinical decision-making.
RNA signatures
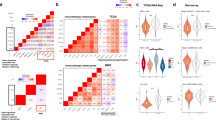
Whereas IHC approaches have rarely been adopted in routine clinical use, an increasing number of RNA-based classifiers have now been validated and commercialized, possibly because RNA assays are more sensitive and the results are more reproducible between laboratories. The sequencing of the human genome and the development of microarray technology for high-throughput RNA profiling have driven an interest in identifying signatures to measure tumour cell-intrinsic radiosensitivity.
Torres-Roca and collaborators77,78,79 performed transcriptomic and expression analyses to develop the radiosensitivity index (RSI), a ten-gene signature to predict tumour cell-intrinsic radiosensitivity measured by SF2. RSI has been assessed in retrospective cohorts of patients across various cancer types, with evidence suggesting that it predicts tumour radiosensitivity (Table 1). The signature was subsequently marketed as CvergenX, a molecular diagnostic for radiation therapy. RSI has been integrated with the radiation dose delivered to derive the genomic-assisted radiotherapy dose (GARD). A model was developed on more than 8,000 tumour samples from more than 20 tumour types, and GARD was derived using a linear quadratic equation that correlates cell survival and delivered dose and each patient’s RSI and their target radiation dose and fractionation schedule80. As with RSI, GARD has been assessed in several retrospective cohorts of patients with cancer, but has not been validated in prospective studies (Table 2). GARD is commercially available as part of CvergenX.
Many other gene signatures have been developed to predict tumour radiosensitivity (Table 3), particularly for breast and prostate cancers, although these are currently not available commercially. In breast cancer, Speers et al.81 derived a 51-gene signature by irradiating multiple cell lines and correlating SF2 values with gene expression data. The signature outperformed conventionally used clinical features on multivariable analysis (HR 2.78, 95% CI 1.68–4.59; P = 0.00007) in an independent validation data set. Investigators from Denmark used resection samples from 191 patients enrolled in a randomized trial of post-mastectomy radiotherapy to develop a seven-gene signature associated with locoregional tumour control (LRC). Internal validation using samples from a further 112 patients demonstrated that radiotherapy improves LRC in patients with high-risk disease (57% versus 12% with no radiotherapy at 20 years; P < 0.0001) but not in those with low-risk disease (8% versus 9% at 20 years; P = 0.93)82. Perhaps the most highly developed genomic signature of radiosensitivity in breast cancer is the ARTIC classifier, which was derived from three publicly available patient cohorts and incorporates 27 genes and the patient’s age. The classifier was validated in the SweBCG91-RT trial cohort (1,178 patients randomly assigned to radiotherapy versus no radiotherapy following surgery): patients with low ARTIC scores (dichotomized as below the 75th percentile) who received radiotherapy had a reduced risk of locoregional relapse (LRR) (HR 0.33, 95% CI 0.21–0.52; P < 0.01) compared with those who did not receive radiotherapy. Furthermore, among patients who received radiotherapy, high ARTIC scores (indicative of radioresistance) were associated with increased rates of LRR (23.8% versus 9.2%; P < 0.01), suggesting that the classifier can be used to identify groups of patients who might need radiotherapy intensification. In the same study, eight previously published gene signatures were tested on the validation data set, and none had a statistically significant interaction with radiotherapy, that is, they did not have predictive value83. Other gene expression assays based on quantitative reverse transcriptase PCR or RNA microarrays have been developed primarily to inform decisions on adjuvant chemotherapy based on the risk of distant metastases, including OncotypeDX, MammaPrint and Prosigna84,85. Establishing whether these assays predict benefit from adjuvant radiotherapy is an area of interest: retrospective studies have shown positive correlation between assay results and LRC, although the results of ongoing trials, such as IDEA (NCT02400190), PRECISION (NCT02653755), EXPERT (NCT02889874) and TAILOR RT (NCT03488693) are awaited86.
Several signatures have been developed to predict benefit from adjuvant radiotherapy in men undergoing surgery for prostate cancer. PORTOS is a signature with 24 genes involved in DDR that was developed retrospectively using data from five studies (n = 196) that compared adjuvant radiotherapy with no radiotherapy and for which array samples and information on relevant clinicopathological factors were available. In patients with a high PORTOS score, adjuvant radiotherapy was associated with a reduced incidence of distant metastases (HR 0.12, 95% CI 0.03–0.41; P < 0.0001). This benefit was confirmed in a validation cohort (n = 330) pooled from four independent studies (HR 0.15, 95% CI 0.04–0.60; P = 0.002)87. Decipher is a 22-gene classifier developed using a retrospective, single-institution data set (n = 639) of patients who underwent prostatectomy to predict for risk of early clinical metastases after rising prostate-specific antigen (PSA). Decipher has been validated in multiple studies, of which the latest include a retrospective prostatectomy cohort from the NRG Oncology/RTOG 9601 randomized trial, in which this classifier demonstrated prognostic value for risk of distant metastasis on multivariable analysis (HR 1.17, 95% CI 1.05–1.32; P = 0.06)88, and a data set from the SAKK 09/10 dose-escalation trial. Decipher scores were prognostic in the latter cohort, although disappointingly, high versus low scores did not predict benefit from dose escalation89.
DNA-based methods
Various studies have assessed whether mutations in candidate DDR genes are significantly associated with radiotherapy outcomes. A well-studied example of the use of genomics to guide radiotherapy dose prescription is that of homozygous germline ATM mutations in patients with ataxia telangiectasia. Radiotherapy is generally avoided in these patients because their high clinical and cellular radiosensitivity typically results in exceptionally severe toxicities that can lead to death during or shortly after treatment90. In these patients, radiotherapy has been used in situations of dire clinical need, and disease control can be achieved with doses of approximately one-third of those prescribed in typical schedules91,92 (Supplementary Table 1). Other known rare genetic syndromes associated with homozygous germline mutations in genes involved in DDR that lead to increased cellular and clinical radiosensitivity, risk of toxicity and radiosusceptibility have been reviewed elsewhere93. Although exposure to therapeutic radiation is avoided in patients with these rare syndromes, any tumours that they might develop are likely to be radiosensitive because these genetic alterations are associated with hypersensitivity. As is often the case in biology, not all situations can be classified as being in a fixed status, and homozygous mutations in TP53 generally lead to both radioresistance and radiosusceptibility.
No evidence exists that patients harbouring heterozygous germline ATM mutations have radiosensitive tumours. However, a growing interest exists in evaluating multigene panels at the germline level, which is increasingly feasible because improved affordability is enabling more centres to perform routine genetic testing. For example, in a study of 286 patients with breast cancer, 25% were identified as having germline pathogenic variants: 12.6% had mutations in BRCA1/2 and, among non-BRCA1/2 variants, the most commonly detected mutations affected ATM, CHEK2, PALB2, CDH1, TP53 and PTEN94. OS and other end points were similar in patients with and without pathogenic variants, suggesting that studies focused on somatic rather than germline alterations might be more fruitful.
In another study, researchers reported that a patient with locally advanced tongue carcinoma and a somatic frameshift ATM mutation had an exceptional response to palliative radiotherapy. Subsequently, a further eight patients across multiple cancer types were identified who had somatic truncating ATM mutations and experienced a durable response following palliative radiotherapy95. In a larger study of 357 patients with advanced-stage cancer across multiple subtypes harbouring a somatic ATM mutation, patients with a loss-of-function mutation had lower cumulative rates of tumour progression at 2 years following radiotherapy than those with a variant of unknown significance (HR 0.51, 95% CI 0.34–0.77; P = 0.01)96.
In a study of copy number variations in DDR genes, gain-of-function mutations in NBN (encoding nibrin) were detected in 14% of 139 prostate cancer specimens and were associated with reduced biochemical relapse-free survival after radiotherapy on multivariable analysis (HR 3.28, 95% CI 1.56–6.89; P = 0.0017)97. Somatic mutations in RAD51 can also confer radiosensitivity. RAD51G172T polymorphisms, identified by genotyping 193 cervical carcinomas from patients receiving chemoradiotherapy, were associated with improved OS on multivariable analysis (HR 0.37, 95% CI 0.18–0.77; P = 0.008)98. PTEN mutations have also been associated with increased radiosensitivity in preclinical models99. For example, a 39-year-old patient with Cowden syndrome, caused by a germline PTEN mutation, developed total hair loss during her first week of whole-brain radiotherapy (<10 Gy)100.
Other somatic mutations are associated with radioresistance. A meta-analysis of 168 reports involving 18,766 patients with colorectal cancer found that TP53 mutations are associated with poor OS after radiotherapy (RR 1.31, 95% CI 1.19–1.45)101. Studies in patients with breast102, head and neck103 and oesophageal104,105 cancers, gliomas106 and sarcomas107 have also described an association between somatic mutations in TP53 and a poor response to radiotherapy. Other studies have identified somatic mutations in non-DDR genes associated with resistance to radiotherapy. For example, mutations in KEAP1 and NFE2L2 are frequent in non-small-cell lung cancer (NSCLC) and are associated with marked radioresistance. Irradiation of squamous lung cancer cell lines demonstrated that activating NFE2L2 mutations are associated with radioresistance108. Jeong et al.109 genotyped 42 NSCLC specimens and found significantly higher rates of cumulative LRR after chemoradiotherapy in patients harbouring KEAP1 or NFE2L2 mutations than in those with wild-type tumours (70% versus 18%; P < 0.003). NFE2L2 mutations have also been detected in head and neck squamous cell carcinomas (HNSCCs). In a study genotyping 20 early-stage glottic larynx tumours, these were classified as radioresistant (n = 8) or radiosensitive (n = 12) on the basis of clinical response, and higher rates of NFE2L2 mutations were found in radioresistant than in radiosensitive tumours (P < 0.05)110. Finally, gene sequencing of 18 cervical cancers demonstrated that the presence of simultaneous mutations in KRAS and SMAD4 is associated with LRR after radiotherapy. Subsequent analysis of genomic data from more than 50,000 cervical cancers demonstrated a trend towards co-occurrence of KRAS and SMAD4 mutations in this cancer type, and preclinical experiments suggested that cells harbouring these mutations are preferentially killed by carbon ion therapy111.
In the past few years, whole-genome sequencing approaches have been used in supervised analyses to identify somatic mutations associated with radiosensitivity. Researchers performed next-generation sequencing on tumour samples from 97 patients with various solid malignancies and found that 23%, 5% and 6% of patients harboured missense, nonsense or frameshift mutations, respectively, in BRCA1/2, ATM or both. The presence of these mutations was associated with low LRR 1 year after radiotherapy (0% versus 47%; P = 0.008)112. In another study, sequencing of up to 468 cancer-associated genes in samples from 110 locally advanced NSCLCs revealed that loss-of-function mutations in KMT2C (12% of patients) and KMT2D (9%) were independently associated with inferior OS after (chemo)radiotherapy (HR 13.4 and 7.0, respectively; P < 0.001 both). Somatic mutations in AKT2 (2% of patients) were associated with inferior LRC (HR 9.0; P = 0.007). Conversely, deleterious mutations in a panel of 38 genes involved in DDR (0–3%) were associated with improved LRC compared with missense mutations or wild-type DDR genes (P = 0.049)113. Researchers from the same institution performed next-generation sequencing on resection specimens from 89 patients who underwent surgery for NSCLC, followed by post-operative radiotherapy. Sixteen patients had deleterious mutations in genes previously associated with radioresistance (STK11, KEAP1, NFE2L2 and PIK3CA) and had significantly higher rates of LRR 2 years after radiotherapy than those without such mutations (60% versus 11%; P < 0.001). Conversely, for the 15 patients with mutations only in genes involved in DDR (ARID1A, ATM, POLE, PMS2 and TP53BP1), 2-year LRR was 0% versus 22% in patients without such mutations (P = 0.048). Of note, in patients treated with surgery and no radiotherapy (n = 19), the 2-year LRC was 57% regardless of whether or not they had mutations in radioresistance genes or in DDR genes (P = 0.99). Furthermore, the authors reported that a higher tumour mutational burden (TMB) was independently associated with LRC on multivariable analysis (HR 0.86, 95% 0.77–0.97; P = 0.01)114. This finding is plausible given that, in a study of patients with breast cancer, higher TMB (≥20 mutations/megabase (mut/Mb)) was correlated with an increased frequency of mutations in DDR genes (P = 0.043)115.
Tumour hypoxia
The development of methods for measuring tumour hypoxia has been an active area of radiotherapy research since the 1960s. Several available approaches include direct measurement of oxygen tension (for example, using the Eppendorf histograph device), exogenous markers (such as pimonidazole), cell-intrinsic biomarkers (for example, CA9), imaging (with MRI or PET) and gene signatures. Given that this area of research has been comprehensively reviewed elsewhere24, here, we focus on methods that are closest to clinical application.
Numerous studies using these various methods have shown that highly hypoxic tumours have a poor response to radiotherapy. Given that the most hypoxic tumours are also the most aggressive, the prognosis is generally poor after surgery and thus, avoidance of radiotherapy is not an option. Similarly, hypoxia confers resistance to most types of chemotherapy. Fortunately, a high level of preclinical and clinical evidence indicates that targeting hypoxia concomitantly with radiotherapy improves patient outcomes. Several hypoxia-targeting agents (such as nimorazole and carbogen–nicotinamide) are already available, and their development continues to be an active area of research116. In terms of biological personalization of radiotherapy, measuring hypoxia can help to identify patients who could receive hypoxia-targeting agents, because robust evidence indicates that only patients with the most hypoxic tumours benefit from them. The best developed area in this regard is that of gene signatures, which have been extensively validated and shown to predict benefit from hypoxia-targeting agents117,118. The key advantages of using signatures are that they can be evaluated in samples obtained from diagnostic biopsies, with no need for additional interventions, and that the methods for assessing RNA expression are very well developed and widely available for clinical application. The currently available signatures are listed elsewhere24, and additional signatures continue to be developed, some of which are progressing to commercialization119. A comparison of 32 hypoxia-related signatures identified commonalities and differences117; most signatures are tailored for specific tumour types and, although no single gene is included in all signatures, overlap can occur117. The most reliable signatures are those validated in multiple cohorts that have also been shown to predict benefit from hypoxia-targeting agents concomitant with radiotherapy24. Disappointingly, however, the preliminary report of the EORTC 1219–DAHANCA 29 trial showed no benefit from the addition of the hypoxia sensitizer nimorazole to accelerated (total dose given over a shorter period of time compared with standard radiotherapy) chemoradiotherapy in a cohort of patients with p16– HNSCC, or in the subgroup of those with hypoxic tumours, who were identified using a 15-gene signature that was previously shown to predict benefit from having nimorazole with radiotherapy120.
The other area with potential for clinical application is that using imaging to identify hypoxic subvolumes on which to perform dose escalation. Most research involves PET and MRI and has been reviewed elsewhere121,122. Despite a paucity of large-cohort validation studies and prospective assessment and a lack of standardization of imaging approaches, this area of research has the potential to be used for biological optimization of radiotherapy. In a phase II trial with results reported in 2022, patients with HNSCC underwent pretreatment PET scanning with the hypoxia tracer [18F]fluoromisonidazole. Those with hypoxic tumours were randomly allocated to receive standard radiotherapy (70 Gy in 35 fractions) or dose escalation (77 Gy in 35 fractions) delivered to the hypoxic subvolume. This study confirmed the prognostic value of hypoxia PET and dose escalation, improving the 5-year LRC by 25% (P = 0.15). Unfortunately, the study also had slow accrual and closed early, possibly owing to the complexity of the design123.
Proliferation and repopulation ability
Accelerated tumour repopulation is the process whereby tumour cells that survive fractionated radiotherapy proliferate more quickly during instead of before treatment124. The process occurs 3–4 weeks after the start of radiotherapy as cell death and re-oxygenation enable surviving radioresistant CSCs (which had been previously growth-suppressed owing to crowding and nutrient deprivation) to proliferate more quickly125.
Biomarkers of proliferation that are easily detectable on IHC have been studied, and Ki-67 (expressed by proliferating cells) is probably the most widely used. A secondary analysis of the Medical Research Council’s Continuous Hyperfractionated Accelerated Radiotherapy (CHART) randomized trial using hierarchical clustering identified a group of HNSCCs negative for p53 and the anti-apoptotic protein BCL-2, and with high levels of CD31 and low levels of Ki-67 on IHC. Patients with these tumours derived a large local tumour control benefit (HR 0.14; P = 0.004) from the CHART schedule (54 Gy in 36 fractions over 12 consecutive days) compared with conventionally fractionated radiotherapy, likely owing to their tumours having a high propensity for accelerated repopulation126. Given these results, a classifier associated with the ability to undergo accelerated repopulation could potentially guide the choice of fractionation schedule. Subsequently, several studies showed that high tumour EGFR expression detected by IHC can predict benefit from accelerated versus conventional radiotherapy (Supplementary Table 2). Although several randomized trials have reported this finding, research on EGFR as a biomarker has not progressed, probably owing to reduced interest in studying modified fractionation versus novel combinations. However, in our opinion this area remains very relevant given the increasing use of hypofractionation in radiotherapy.
In the past few years, and owing to the ready availability of high-throughput databases linked to survival outcomes, interest has emerged in the development of classifiers associated with tumour radiosensitivity. Genes involved in proliferation, cell cycle regulation and the DDR are often included in these signatures. In a study using The Cancer Genome Atlas (TCGA) database investigators analysed transcriptomic data from 1,664 patients across 15 cancer types treated with radiotherapy. The top 100 genes identified as differentially expressed in radiotherapy responders versus non-responders were used to build a gene classifier. These genes were involved in cell proliferation, migration, invasion, epithelial-to-mesenchymal transition and DDR pathways127. In another study, researchers analysed TCGA data from 1,016 patients with solid tumours (HNSCC, cervical cancer and breast cancer) who received radiotherapy. Fifty hallmark gene sets from the Molecular Signatures Database were analysed and divided into eight categories defined by a shared biological or functional process. Three of them (termed radiobiological, metabolism and proliferation) were prognostic for OS in the three cancer types128.
Stem cell properties
The identification of CSCs is hindered by intertumour and intratumour heterogeneity, and the lack of specific biomarkers129. Surrogate biomarkers include the enzyme aldehyde dehydrogenase (ALDH; elevated in CSCs), colony-forming ability, in vivo tumorigenic ability and various cell-surface biomarkers. Nonetheless, although multiple cell-surface biomarkers have been identified (such as CD44 and CD133), no single biomarker is specific or applicable to multiple cancer types130 and thus, no clinically useful CSC biomarker is available. The development of therapeutic approaches to directly overcome CSC-associated radioresistance has been hampered by factors that include the plasticity of CSCs, and a single CSC-targeted approach does not currently exist. Nevertheless, novel PET imaging techniques have been demonstrated to enable the identification of CD133+ tumour subvolumes, raising the possibility of future trials of dose escalation to such areas131. Other radiation-specific approaches include the use of charged particle therapy; indeed, in preclinical studies, charged particles can overcome CSC radioresistance via induction of complex DNA damage, inhibition of cell cycle progression and lack of oxygen dependence132. In preclinical studies, inhibition of Notch signalling (involved in CSC maintenance) using γ-secretase inhibitors rendered glioma cells radiosensitive133. Several Notch inhibitors are being tested as single agents in early-phase clinical trials; disappointingly, a trial of the inhibitor R04929097 in combination with whole-brain or stereotactic radiotherapy in patients with brain metastases was terminated owing to slow accrual (NCT01217411). Future trials testing Notch inhibitors in combination with radiotherapy are needed134. Other drug-based approaches include the addition of the disulfiram–copper complex, which eliminated CSC stemness (determined with various functional assays), to radiotherapy in a xenograft model of chondrosarcoma135. EGFR activation has been shown to induce a CSC phenotype136, a process that might partially explain the previously discussed observation that tumours with high levels of EGFR benefit from accelerated radiotherapy versus conventional fractionation (Supplementary Table 2).
In terms of genomic classifiers, a few studies have tried to assess the relative importance of those involved in CSC properties. A study of biomarkers of response to chemoradiotherapy in patients with HNSCC identified various biomarkers associated with OS outcomes, among which were two CSC biomarkers (CD44 and SLC3A2)137. Moreover, the study found additive value in combining them with EGFR and immune biomarkers in multifactorial analyses. Overall, although the number of CSCs is accepted to have an important role in radiosensitivity, the research in this area is not sufficiently advanced in terms of clinical utility.
Immune biomarkers
The development of biomarkers of radiation-induced immunity and the identification of patients who could benefit from radiotherapy–immunotherapy combinations are two areas of interest. Several immune-related gene signatures associated with radioresponse have been developed. For example, transcriptome-wide expression profiling of 136 muscle-invasive bladder cancer samples identified several signatures, among which one related to T cell activation was associated with improved disease-specific survival in patients receiving chemoradiotherapy (HR 0.30, 95% CI 0.14–0.65; P = 0.002) but not in those receiving neoadjuvant chemotherapy and/or surgery138. TMB predicts benefit from immune checkpoint inhibitors (ICIs) in the setting of metastatic disease139 and might also predict benefit from radiotherapy. However, in 101 patients with locally advanced HNSCC receiving chemoradiotherapy, a high TMB (≥5.1 mut/Mb) was associated with inferior OS compared with a low TMB on multivariable analysis (HR 1.79, 95% CI 1.02–3.14; P = 0.04)140. Other potential biomarkers related to immune status include sex, with evidence indicating that female patients have increased radiosensitivity141, and the composition of the gut microbiota, with studies describing that the presence of certain intestinal bacterial species augments immune responses following radiation142.
Some evidence indicates that irradiation of the primary site can improve outcomes in patients with metastatic disease by promoting antitumour immunity, although the mechanism underlying this effect is unclear. In a prespecified subgroup analysis of the STAMPEDE trial, patients with low-volume metastatic disease receiving radical-dose radiotherapy to the prostate had improved OS compared with those receiving standard-of-care systemic therapy (81% versus 73% at 3 years; HR 0.68, 95% CI 0.52–0.90; P = 0.007)143. Similarly, in a phase III trial involving patients with nasopharyngeal carcinomas, de novo metastatic disease and a response to induction chemotherapy, the addition to chemotherapy of subsequent radical radiotherapy to the head and neck improved OS (76.4% versus 54.5% at 2 years with chemotherapy alone; HR 0.42, 95% CI 0.23–0.77; P = 0.004)144.
Many studies of radiotherapy–immunotherapy combinations in patients with metastatic disease are ongoing, with the aim of assessing for synergy and/or the abscopal effect145. In the curative setting, the use of adjuvant ICIs after chemoradiotherapy in NSCLC146 and surgery for oesophageal cancer147 has proved successful; however, trials of radiotherapy with concurrent ICIs have not demonstrated benefit142. For example, in the JAVELIN Head and Neck 100 trial involving patients with HNSCC the addition of the anti-PD-L1 antibody avelumab to chemoradiation did not improve progression-free survival (PFS) or OS148, perhaps owing to detrimental effects of elective nodal irradiation on the immune system149. Many trials continue in this area, some with a particular interest in the development of predictive biomarkers, identification of optimal radiotherapy fractionation schedules (such as hypofractionated versus conventional fractionation) and the addition of DDR inhibitors150. Tumour PD-L1 status is a potential biomarker of benefit from the addition of immunotherapy to radiotherapy. In a planned subgroup analysis of JAVELIN Head and Neck 100, only patients with tumours classified as PD-L1-high (with PD-L1 expression on ≥25% of tumour cells) derived a PFS benefit from avelumab plus chemoradiation at 2 years, although the confidence intervals were wide (HR 0.59, 95% CI 0.28–1.22)148. In PACIFIC, patients derived a PFS benefit from the anti-PD-1 antibody durvalumab after chemoradiotherapy (HR 0.55, 95% CI 0.45–0.68), which was more pronounced in those with PD-L1-high tumours (HR 0.41, 95% CI 0.26–0.65)146. The pretreatment absolute lymphocyte count has prognostic value as a biomarker of immune function and also predicts for benefit from the addition of radiotherapy to chemotherapy in oropharyngeal carcinoma151 and to immunotherapy in various solid tumour types152. Although this area of investigation is currently very active, at the time of writing none of the above immune biomarkers has been translated into clinical practice153.
Clinical potential
Although our Review focuses on tumour radiosensitivity, we want to highlight the interest in measuring the radiosensitivity of non-malignant tissue to personalize radiotherapy dose prescriptions. Ideally, in the future both individual and tumour radiosensitivity would be measured routinely in all patients to maximize the probability of survival and minimize the risks of toxicity, although a relationship between the two has not been established in most patients. An exception is that of individuals harbouring homozygous mutations in ATM, in whom measurements of individual cellular radiosensitivity underpin the use of reduced tumour doses that achieve LRC without the excessive toxicity associated with full-dose radiotherapy (Supplementary Table 1). This example illustrates how biology-guided precision approaches can be used in clinical practice, although these mutations are rare.
The measurement of cell-intrinsic radiosensitivity in non-malignant tissues is important for future prediction of individual risk, not only of radiotherapy-associated toxicities but also of developing second malignancies following radiological exposure154. In the past, this area has been hindered by a lack of collaborative initiatives, with too many small, underpowered studies and lack of validation in independent cohorts. The Radiogenomics Consortium was established in 2010 to promote a large collaborative framework to facilitate the identification of genetic variants associated with radiotherapy-related toxicities155. For example, the Consortium performed a meta-analysis of six genome-wide association studies using data from ~4,000 men with prostate cancer from six centres and identified three single-nucleotide polymorphisms associated with increased risk of toxicity following radiotherapy156. Of course, genetic testing alone is unlikely to predict risk of toxicity with the specificity required to modify doses and thus, ongoing work is focused on developing predictive models that incorporate dose (individual doses to organs at risk), patient-related factors (such as age and smoking history) and genetic variants157. Once sufficiently validated, these models could be used clinically to individualize treatments, for example, recommending modification of dose constraints to non-malignant tissue, avoidance of radiotherapy or dose increases in patients with radioresistant tumours. Advantages of using genetic testing instead of other methods for measuring individual radiosensitivity include the requirement of only a single blood test (although the use of other biospecimens, such as saliva, is becoming increasingly feasible) and the availability of genetic testing equipment in clinical laboratories. Although genetic testing in this setting holds promise, the blood-based tests being commercialized to predict tolerance to radiotherapy for patients with prostate or breast cancers also include a functional assay of radiosensitivity along with patient factors. These tests were validated in multicentre prospective clinical studies158 and are currently recommended by the French Society of Radiation Oncology and thus, can be used to deliver precision radiotherapy in France. As discussed, an RSI-based approach has also been commercialized as one of the first tools to deliver genomics-guided radiation oncology. This test is likely to become available soon in the USA. Some hypoxia signatures are also progressing towards commercialization.
In terms of potential benefit, prior modelling studies suggest that the probability of tumour control can be substantially improved by prescribing radiotherapy doses guided by the results of cellular radiosensitivity assays159. An obvious application of radiotherapy personalization is dose escalation to radioresistant tumours. In general, attempts to perform dose escalation in unselected patients have been unsuccessful, resulting in increased toxicity and with little evidence of increased tumour control. Dose painting, or the delivery of an increased radiation dose to resistant tumour subvolumes, is possibly a more promising avenue. For example, in the previously described trial of dose escalation to hypoxic subvolumes for patients with locally advanced HNSCC, there was a trend towards improved 5-year LRC, although this trial was closed early owing to poor accrual and thus, was underpowered123.
A further area of potential patient benefit is dose de-escalation to reduce the risks of toxicity in patients with radiosensitive tumours. For example, the observation that HPV positivity in oropharyngeal carcinomas is associated with radiosensitivity led to trials of de-escalation in this setting. Unfortunately, the first three randomized de-escalation trials showed a clear detriment in OS when cisplatin was omitted or substituted160. This example highlights the potential dangers of implementing changes in clinical practice before high-level evidence is obtained160. Clearly, we are not yet ready to de-escalate radiotherapy regimens in patients with oropharyngeal carcinomas on the basis of HPV positivity alone.
Future directions
Robust biomarkers of tumour radiosensitivity are urgently required for patients to benefit from precision radiotherapy161. This area of research has yielded many discoveries but little validation has been made to assess transferability between centres. The latter is important because different centres can use different fractionation schedules and combination treatments. The radiotherapy community needs to focus more on validation studies that involve international collaborative groups — in this regard, initiatives such as the Radiogenomics Consortium show the advantages of increased study power. We need biomarkers that rise above the noise, that is, that overcome heterogeneity between cohorts. Funders should and are now supporting validation and clinical application over discovery.
Clinical delivery of personalized radiotherapy will happen earlier in certain cancer subtypes. For example, strategies to reduce radiotoxicity are ongoing in HPV-associated oropharyngeal carcinoma, in which the survival outcomes of patients receiving chemoradiotherapy are generally excellent but treatment-related late toxicities are a particular concern, despite the failure of de-escalation trials160. Several groups have proposed a radical reduction in adjuvant radiotherapy doses (for example, from 60 Gy to 30 Gy), either as a blanket approach162 or guided by functional PET imaging163. Although the data obtained thus far are promising, results from larger phase II (such as NCT03323463) and phase III (NCT02908477) trials are required. Other examples include breast and prostate cancer, in which gene signatures can identify patients with radiosensitive tumours who might have excellent OS outcomes without receiving conventional radiotherapy schedules, although supporting evidence from clinical trials is needed.
Importantly, the complexity of the biological pathways associated with tumour radiosensitivity has made it clear that biomarkers should be evaluated alongside other known prognostic factors. For example, given that increasing tumour volume (a poor prognostic factor for cancers treated with radiotherapy such as those of the head and neck35,164 and lungs165) is correlated with other factors (such as hypoxia and CSC count166), biomarker validation studies require multivariable analyses that include tumour volume, to avoid confounding. In the setting of HNSCC, the German Cancer Consortium–Radiation Oncology Group (DKTK-ROG) has described correlations between multiple factors, including tumour p16 status, hypoxia, tumour volume and CSC, and survival35,167,168. This group identified biomarker signatures of radioresponse that are currently being assessed in an ongoing prospective observational study (NCT02059668) and will be validated in a planned interventional trial169.
When developing biomarkers, the dynamic changes evident in tumours during radiotherapy require attention. For example, assessment of tumour hypoxia on 18F-PET–CT in HNSCC has been shown to be less predictive of LRC when it is performed before chemoradiotherapy than when it is performed mid-treatment170. Measuring a biomarker on a pretreatment diagnostic biopsy sample or image is an attractive approach to guide management, but in reality little is known about the optimal times for assessing certain biomarkers (such as immune or unresolved hypoxia biomarkers).
Pharmaceutical companies are driving initiatives to identify potential novel radiosensitizers in what has become a crowded area of research. Given that the risk of toxicities limits the number of drugs that can be combined with radiation, in our view the major need in this area is to identify biomarkers of benefit from novel interventions that will enable more complex biomarker-driven trials with multiple targeted interventions.
Conclusions
In this Review we posit that advances in genomics are now ready to guide the delivery of radiotherapy, and that commercialization of genomic-based tools is important to drive implementation119. Developments from the past 20 years have enabled unprecedented high-throughput analyses at the RNA and DNA levels. The ability to extract nucleic acids from routine clinical biopsy-derived samples continues to improve and the techniques for rapid RNA and DNA profiling are embedded in clinical laboratories.
We need to learn the lessons from past failures to translate radiobiology biomarkers into the clinic. Many years of developing approaches for measuring hypoxia have been compromised by the inability to work collaboratively to transfer methods across laboratories, standardize and carry out the required validation work. Well-designed trials of biomarker-driven personalized radiotherapy are required to demonstrate benefit. The use of genomic tools to inform clinical decisions is proving difficult, and to date, most technical radiotherapy innovations have been applied empirically without robust evidence of therapeutic benefit.
The number of RNA-based gene signatures used in oncology has steadily increased over time. We now have well-validated signatures of radiosensitivity and hypoxia that could be used at present. Both types of signature are promising because they can be used for straightforward targeted interventions, including the avoidance of radiotherapy in patients with low-risk breast cancer and delivery of hypoxia-targeting agents, respectively. Although the EORTC 1219–DAHANCA 29 trial did not validate a 15-gene signature as a predictor of benefit from hypoxia-targeting agents120, we argue the need for extensive validation and multinational collaboration to ensure the robustness of biomarkers. Funders need to recognize the importance of biomarker assay development and support adequately sized clinical trials.
The development of DNA-based signatures lags behind that of those based on RNA, but the increasing affordability of analysis tools means that more progress can be achieved. Current evidence suggests that somatic mutations in key genes involved in the DDR confer sensitivity to radiotherapy. In this Review, we highlight the potential of genomics-guided radiotherapy and call for all stakeholders to recognize this area as a research priority.
References
Ringborg, U. et al. The Swedish Council on Technology Assessment in Health Care (SBU) systematic overview of radiotherapy for cancer including a prospective survey of radiotherapy practice in Sweden 2001 – Summary and conclusions. Acta Oncol. 42, 357–365 (2003).
Mackie, T. R. et al. Image guidance for precise conformal radiotherapy. Int. J. Radiat. Oncol. Biol. Phys. 56, 89–105 (2003).
Li, G. et al. Advances in 4D medical imaging and 4D radiation therapy. Technol. Cancer Res. Treat. 7, 67–81 (2008).
Otto, K. Volumetric modulated arc therapy: IMRT in a single gantry arc. Med. Phys. 35, 310–317 (2008).
Teoh, M., Clark, C. H., Wood, K., Whitaker, S. & Nisbet, A. Volumetric modulated arc therapy: a review of current literature and clinical use in practice. Br. J. Radiol. 84, 967–996 (2011).
Yu, C. X. Intensity-modulated arc therapy with dynamic multileaf collimation: an alternative to tomotherapy. Phys. Med. Biol. 40, 1435–1449 (1995).
Rivera, A. L. et al. MGMT promoter methylation is predictive of response to radiotherapy and prognostic in the absence of adjuvant alkylating chemotherapy for glioblastoma. Neuro Oncol. 12, 116–121 (2010).
Rouzier, R. et al. Multigene assays and molecular markers in breast cancer: systematic review of health economic analyses. Breast Cancer Res. Treat. 139, 621–637 (2013).
Redza-Dutordoir, M. & Averill-Bates, D. A. Activation of apoptosis signalling pathways by reactive oxygen species. Biochim. Biophys. Acta Mol. Cell Res. 1863, 2977–2992 (2016).
Maier, P., Hartmann, L., Wenz, F. & Herskind, C. Cellular pathways in response to ionizing radiation and their targetability for tumor radiosensitization. Int. J. Mol. Sci. 17, 102 (2016).
Ouellette, M. M., Zhou, S. & Yan, Y. Cell signaling pathways that promote radioresistance of cancer cells. Diagnostics 12, 656 (2022).
Barker, H. E., Paget, J. T. E., Khan, A. A. & Harrington, K. J. The tumour microenvironment after radiotherapy: mechanisms of resistance and recurrence. Nat. Rev. Cancer 15, 409–425 (2015).
Kumari, S. et al. Immunomodulatory effects of radiotherapy. Int. J. Mol. Sci. 21, 8151 (2020).
Ghasemi, Z. et al. Fractionated radiation promotes proliferation and radioresistance in bystander A549 cells but not in bystander HT29 cells. Life Sci. 257, 118087 (2020).
Chalmers, A. J. & Carruthers, R. D. Radiobiology summaries: DNA damage and repair. Clin. Oncol. 33, 275–278 (2021).
Feng, W., Smith, C. M., Simpson, D. A. & Gupta, G. P. Targeting non-homologous and alternative end joining repair to enhance cancer radiosensitivity. Semin. Radiat. Oncol. 32, 29–41 (2022).
Chang, H. H. Y., Pannunzio, N. R., Adachi, N. & Lieber, M. R. Non-homologous DNA end joining and alternative pathways to double-strand break repair. Nat. Rev. Mol. Cell Biol. 18, 495–506 (2017).
Wyatt, D. W. et al. Essential roles for polymerase θ-mediated end joining in the repair of chromosome breaks. Mol. Cell 63, 662–673 (2016).
Li, X. & Heyer, W. D. Homologous recombination in DNA repair and DNA damage tolerance. Cell Res. 18, 99–113 (2008).
Weber, A. M. & Ryan, A. J. ATM and ATR as therapeutic targets in cancer. Pharmacol. Ther. 149, 124–138 (2015).
Aubrey, B. J., Kelly, G. L., Janic, A., Herold, M. J. & Strasser, A. How does p53 induce apoptosis and how does this relate to p53-mediated tumour suppression. Cell Death Differ. 25, 104–113 (2018).
Engeland, K. Cell cycle arrest through indirect transcriptional repression by p53: I have a DREAM. Cell Death Differ. 25, 114–132 (2018).
Baumann, M. et al. EGFR-targeted anti-cancer drugs in radiotherapy: preclinical evaluation of mechanisms. Radiother. Oncol. 83, 238–248 (2007).
Thiruthaneeswaran, N. et al. Lost in application: measuring hypoxia for radiotherapy optimisation. Eur. J. Cancer 148, 260–276 (2021).
West, C. M. & Slevin, F. Tumour hypoxia. Clin. Oncol. 31, 595–599 (2019).
Wouters, B. G. & Koritzinsky, M. Hypoxia signalling through mTOR and the unfolded protein response in cancer. Nat. Rev. Cancer 8, 851–864 (2008).
Bolland, H., Ma, T. S., Ramlee, S., Ramadan, K. & Hammond, E. M. Links between the unfolded protein response and the DNA damage response in hypoxia: a systematic review. Biochem. Soc. Trans. 49, 1251–1263 (2021).
Yaromina, A. et al. Radiobiological hypoxia, histological parameters of tumour microenvironment and local tumour control after fractionated irradiation. Radiother. Oncol. 96, 116–122 (2010).
Tang, M., Bolderson, E., O’Byrne, K. J. & Richard, D. J. Tumor hypoxia drives genomic instability. Front. Cell Dev. Biol. 9, 626229 (2021).
Semenza, G. L. Heritable disorders of oxygen sensing. Am. J. Med. Genet. A 185, 3334–3339 (2021).
Semenza, G. L. The genomics and genetics of oxygen homeostasis. Annu. Rev. Genomics Hum. Genet. 21, 183–206. (2020).
Vadysirisack, D. & Ellisen, L. W. mTOR activity under hypoxia. Methods Mol. Biol. 821, 44–58 (2012).
Liu, K. X., Everdell, E., Pal, S., Haas-Kogan, D. A. & Milligan, M. G. Harnessing lactate metabolism for radiosensitization. Front. Oncol. 11, 672339 (2021).
Peitzsch, C., Kurth, I., Ebert, N., Dubrovska, A. & Baumann, M. Cancer stem cells in radiation response: current views and future perspectives in radiation oncology. Int. J. Radiat. Biol. 95, 900–911 (2019).
Linge, A. et al. HPV status, cancer stem cell marker expression, hypoxia gene signatures and tumour volume identify good prognosis subgroups in patients with HNSCC after primary radiochemotherapy: a multicentre retrospective study of the German Cancer Consortium Radiation. Radiother. Oncol. 121, 364–373 (2016).
Vlashi, E. & Pajonk, F. Cancer stem cells, cancer cell plasticity and radiation therapy. Semin. Cancer Biol. 31, 28–35 (2015).
Lee, S. Y. et al. Induction of metastasis, cancer stem cell phenotype, and oncogenic metabolism in cancer cells by ionizing radiation. Mol. Cancer 16, 10 (2017).
Quail, D. F., Taylor, M. J. & Postovit, L. M. Microenvironmental regulation of cancer stem cell phenotypes. Curr. Stem Cell Res. Ther. 7, 197–216 (2012).
Diehn, M. et al. Association of reactive oxygen species levels and radioresistance in cancer stem cells. Nature 458, 780–783 (2009).
Bao, S. et al. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature 444, 756–760 (2006).
Peitzsch, C., Kurth, I., Kunz-Schughart, L., Baumann, M. & Dubrovska, A. Discovery of the cancer stem cell related determinants of radioresistance. Radiother. Oncol. 108, 378–387 (2013).
Zhong, J. T. et al. GLUT-1 siRNA enhances radiosensitization of laryngeal cancer stem cells via enhanced DNA damage, cell cycle redistribution, and promotion of apoptosis in vitro and in vivo. Onco Targets Ther. 12, 9129–9142 (2019).
Kaseb, H. O., Fohrer-Ting, H. F., Lewis, D. W., Lagasse, E. & Gollin, S. Identification, expansion and characterization of cancer cells with stem cell properties from head and neck squamous cell carcinomas. Exp. Cell Res. 348, 75–86 (2016).
Emami Nejad, A. et al. The role of hypoxia in the tumor microenvironment and development of cancer stem cell: a novel approach to developing treatment. Cancer Cell Int. 21, 1–26 (2021).
Hanahan, D. Hallmarks of cancer: new dimensions. Cancer Discov. 12, 31–46 (2022).
Kachikwu, E. L. et al. Radiation enhances regulatory T cell representation. Int. J. Radiat. Oncol. Biol. Phys. 81, 1128–1135 (2011).
Honeychurch, J. & Illidge, T. M. The influence of radiation in the context of developing combination immunotherapies in cancer. Ther. Adv. Vaccines Immunother. 5, 115–122 (2017).
Weichselbaum, R. R., Liang, H., Deng, L. & Fu, Y. X. Radiotherapy and immunotherapy: a beneficial liaison? Nat. Rev. Clin. Oncol. 14, 365–379 (2017).
Colton, M., Cheadle, E. J., Honeychurch, J. & Illidge, T. M. Reprogramming the tumour microenvironment by radiotherapy: implications for radiotherapy and immunotherapy combinations. Radiat. Oncol. 15, 254 (2020).
De Martino, M., Daviaud, C. & Vanpouille-Box, C. Radiotherapy: an immune response modifier for immuno-oncology. Semin. Immunol. 52, 101474 (2021).
Krombach, J. et al. Priming anti-tumor immunity by radiotherapy: dying tumor cell-derived DAMPs trigger endothelial cell activation and recruitment of myeloid cells. Oncoimmunology 8, e1523097 (2018).
Fridman, W. H., Pagès, F., Saut̀s-Fridman, C. & Galon, J. The immune contexture in human tumours: Impact on clinical outcome. Nat. Rev. Cancer 12, 298–306 (2012).
Anitei, M. G. et al. Prognostic and predictive values of the immunoscore in patients with rectal cancer. Clin. Cancer Res. 20, 1891–1899 (2014).
Gupta, A. et al. Radiotherapy promotes tumor-specific effector CD8+ T cells via dendritic cell activation. J. Immunol. 189, 558–566 (2012).
Vanpouille-Box, C. et al. TGFβ is a master regulator of radiation therapy-induced anti- tumor immunity. Cancer Res. 75, 2232–2242 (2015).
Dovedi, S. J. et al. Acquired resistance to fractionated radiotherapy can be overcome by concurrent PD-L1 blockade. Cancer Res. 74, 5458–5468 (2014).
Withers HR. The Four R’s of Radiotherapy. vol. 5 (Academic Press, 1975).
Spiotto, M. T. et al. Biology of the radio- and chemo-responsiveness in HPV malignancies. Semin. Radiat. Oncol. 31, 274–285 (2021).
Lee, J. M. & Bernstein, A. p53 mutations increase resistance to ionizing radiation (y radiation/DNA damage/transgenic mice/cardnogenesis). Proc. Natl Acad. Sci. USA 90, 5742–5746 (1993).
Thariat, J. et al. Avoidance or adaptation of radiotherapy in patients with cancer with Li-Fraumeni and heritable TP53-related cancer syndromes. Lancet Oncol. 22, e562–e574 (2021).
Avril, D. et al. Biomarkers of radioresistance in head and neck squamous cell carcinomas. Int. J. Radiat. Biol. https://doi.org/10.1080/09553002.2022.2110301 (2022).
Louis, D. N. et al. The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol. 23, 1231–1251 (2021).
Fertil, B. & Malaise, E. P. Inherent cellular radiosensitivity as a basic concept for human tumor radiotherapy. Int. J. Radiat. Oncol. Biol. Phys. 7, 621–629 (1981).
Fertil, B. & Malaise, E. P. Intrinsic radiosensitivity of human cell lines is correlated with radioresponsiveness of human tumors: analysis of 101 published survival curves. Int. J. Radiat. Oncol. Biol. Phys. 11, 1699–1707 (1985).
Bristow, R. G. & Hill, R. P. Comparison between in vitro radiosensitivity and in vivo radioresponse in murine tumor cell lines. II: in vivo radioresponse following fractionated treatment and in vitro/in vivo correlations. Int. J. Radiat. Oncol. Biol. Phys. 18, 331–345 (1990).
Deacon, J., Peckham, M. J. & Steel, G. G. The radioresponsiveness of human tumours and the initial slope ofthe cell survival curve. Radiother. Oncol. 2, 317–323 (1984).
Dale, RG; Jones B. Radiobiological Modelling in Radiation Oncology (BIR, 2007).
De-Colle, C. et al. Ex vivo γH2AX radiation sensitivity assay in prostate cancer: inter-patient and intra-patient heterogeneity. Radiother. Oncol. 124, 386–394 (2017).
Ho, V. et al. Overexpression of the MRE11-RAD50-NBS1 (MRN) complex in rectal cancer correlates with poor response to neoadjuvant radiotherapy and prognosis. BMC Cancer 18, 869 (2018).
Hasegawa, T. et al. Ku70-Expression prognostiziert Ergebnisse der Strahlentherapie beim Prostatakarzinom. Strahlentherapie Und Onkol. 193, 29–37 (2017).
Wilson, C. R. et al. Expression of Ku70 correlates with survival in carcinoma of the cervix. Br. J. Cancer 83, 1702–1706 (2000).
Choudhury, A. et al. MRE11 expression is predictive of cause-specific survival following radical radiotherapy for muscle-invasive bladder cancer. Cancer Res. 70, 7017–7026 (2010).
Laurberg, J. R. et al. Expression of TIP60 (tat-interactive protein) and MRE11 (meiotic recombination 11 homolog) predict treatment-specific outcome of localised invasive bladder cancer. BJU Int. 110, E1228–E1236 (2012).
Walker, A. K. et al. MRE11 as a predictive biomarker of outcome after radiation therapy in bladder cancer. Int. J. Radiat. Oncol. Biol. Phys. 104, 809–818 (2019).
Söderlund, K. et al. Intact Mre11/Rad50/Nbs1 complex predicts good response to radiotherapy in early breast cancer. Int. J. Radiat. Oncol. Biol. Phys. 68, 50–58 (2007).
Söderlund, K., Skoog, L., Fornander, T. & Askmalm, M. S. The BRCA1/BRCA2/Rad51 complex is a prognostic and predictive factor in early breast cancer. Radiother. Oncol. 84, 242–251 (2007).
Torres-Roca, J. F. et al. Prediction of radiation sensitivity using a gene expression classifier. Cancer Res. 65, 7169–7176 (2005).
Eschrich, S. A. et al. A gene expression model of intrinsic tumor radiosensitivity: prediction of response and prognosis after chemoradiation. Int. J. Radiat. Oncol. Biol. Phys. 75, 489–496 (2009).
Eschrich, S. et al. Systems biology modeling of the radiation sensitivity network: a biomarker discovery platform. Int. J. Radiat. Oncol. Biol. Phys. 75, 497–505 (2009).
Scott, J. G. et al. A genome-based model for adjusting radiotherapy dose (GARD): a retrospective, cohort-based study. Lancet Oncol. 18, 202–211 (2017).
Speers, C. et al. Development and validation of a novel radiosensitivity signature in human breast cancer. Clin. Cancer Res. 21, 3667–3677 (2015).
Tramm, T. et al. Development and validation of a gene profile predicting benefit of postmastectomy radiotherapy in patients with high-risk breast cancer: a study of gene expression in the DBCG82bc cohort. Clin. Cancer Res. 20, 5272–5280 (2014).
Sjöström, M. et al. Clinicogenomic radiotherapy classifier predicting the need for intensified locoregional treatment after breast-conserving surgery for early-stage breast cancer. J. Clin. Oncol. 37, 3340–3349 (2019).
Sparano, J. A. et al. Adjuvant chemotherapy guided by a 21-gene expression assay in breast cancer. N. Engl. J. Med. 379, 111–121 (2018).
Cardoso, F. et al. 70-Gene signature as an aid to treatment decisions in early-stage breast cancer. N. Engl. J. Med. 375, 717–729 (2016).
Krug, D. et al. Commercially available gene expression assays as predictive tools for adjuvant radiotherapy? A critical review. Breast Care 15, 118–126 (2020).
Zhao, S. G. et al. Development and validation of a 24-gene predictor of response to postoperative radiotherapy in prostate cancer: a matched, retrospective analysis. Lancet Oncol. 17, 1612–1620 (2016).
Feng, F. Y. et al. Validation of a 22-gene genomic classifier in patients with recurrent prostate cancer: an ancillary study of the NRG/RTOG 9601 randomized clinical trial. JAMA Oncol. 7, 544–552 (2021).
Pra, A. D. et al. Validation of the Decipher genomic classifier in patients receiving salvage radiotherapy without hormone therapy after radical prostatectomy–an ancillary study of the SAKK 09/10 randomized clinical trial 5. Ann. Oncol. 33, 950–958 (2022).
Pollard, J. M. & Gatti, R. Clinical radiation sensitivity with DNA repair disorders: an overview. Int. J. Radiat. Oncol. Biol. Phys. 74, 1323–1331 (2009).
DeWire, M. D. et al. Radiation therapy and adjuvant chemotherapy in a patient with a malignant glioneuronal tumor and underlying ataxia telangiectasia: a case report and review of the literature. J. Clin. Oncol. 31, e12–e14 (2013).
Hart, R. M., Kimler, B. F., Evans, R. G. & Park, C. H. Radiotherapeutic management of medulloblastoma in a pediatric patient with ataxia telangiectasia. Int. J. Radiat. Oncol. Biol. Phys. 13, 1237–1240 (1987).
Bergom, C. et al. The implications of genetic testing on radiation therapy decisions: a guide for radiation oncologists. Int. J. Radiat. Oncol. Biol. Phys. 105, 698–712 (2019).
Chapman, B. V. et al. Breast radiation therapy-related treatment outcomes in patients with or without germline mutations on multigene panel testing. Int. J. Radiat. Oncol. Biol. Phys. 112, 437–444 (2022).
Ma, J. et al. Genomic analysis of exceptional responders to radiotherapy reveals somatic mutations in ATM. Oncotarget 8, 10312–10323 (2017).
Pitter, K. L. et al. Pathogenic ATM mutations in cancer and a genetic basis for radiotherapeutic efficacy. J. Natl Cancer Inst. 113, 266–273 (2021).
Berlin, A. et al. NBN gain is predictive for adverse outcome following image-guided radiotherapy for localized prostate cancer. Oncotarget 5, 11081–11090 (2014).
Nogueira, A. et al. Role of the RAD51 G172T polymorphism in the clinical outcome of cervical cancer patients under concomitant chemoradiotherapy. Gene 504, 279–283 (2012).
Bassi, C. et al. Nuclear PTEN controls DNA repair and sensitivity to genotoxic stress. Science 341, 395–399 (2013).
Tatebe, K., Chmura, S. J. & Connell, P. P. Elevated radiotherapy toxicity in the setting of germline PTEN mutation. Pract. Radiat. Oncol. 9, 492–495 (2019).
Munro, A. J., Lain, S. & Lane, D. P. p53 abnormalities and outcomes in colorectal cancer: a systematic review. Br. J. Cancer 92, 434–444 (2005).
Jameel, J. K. A., Rao, V. S. R., Cawkwell, L. & Drew, P. J. Radioresistance in carcinoma of the breast. Breast 13, 452–460 (2004).
Pacelli, F. P. et al. Radioresistance in head and neck squamous cell carcinoma: biological bases and therapeutic implications. Head. Neck 37, 763–770 (2015).
Ribeiro, U. et al. p53 sequence analysis predicts treatment response and outcome of patients with esophageal carcinoma. Cancer 83, 7–18 (1998).
Makino, T. et al. p53 mutation status predicts pathological response to chemoradiotherapy in locally advanced esophageal cancer. Ann. Surg. Oncol. 17, 804–811 (2010).
Werbrouck, C. et al. TP53 pathway alterations drive radioresistance in diffuse intrinsic pontine gliomas (DIPG). Clin. Cancer Res. 25, 6788–6800 (2019).
Casey, D. L. et al. TP53 mutations increase radioresistance in rhabdomyosarcoma and Ewing sarcoma. Br. J. Cancer 125, 576–581 (2021).
Abazeed, M. E. et al. Integrative radiogenomic profiling of squamous cell lung cancer. Cancer Res. 73, 6289–6298 (2013).
Jeong, Y. et al. Role of KEAP1/NRF2 and TP53 mutations in lung squamous cell carcinoma development and radiotherapy response prediction. Cancer Discov. 7, 86–101 (2017).
Sheth, S. et al. Correlation of alterations in the KEAP1/CUL3/NFE2L2 pathway with radiation failure in larynx squamous cell carcinoma. Laryngoscope Investig. Otolaryngol. 6, 699–707 (2021).
Oike, T. et al. Mutation analysis of radioresistant early-stage cervical cancer. Int. J. Mol. Sci. 23, 51–12 (2021).
Kim, K. H. et al. Increased radiosensitivity of solid tumors harboring ATM and BRCA1/2 mutations. Cancer Res. Treat. 54, 54–64 (2022).
Luo, L. Y. et al. Genomic analyses for predictors of response to chemoradiation in stage III non-small cell lung cancer. Adv. Radiat. Oncol. 6, 100615 (2021).
Shaverdian, N. et al. Effects of tumor mutational burden and gene alterations associated with radiation response on outcomes of postoperative radiation therapy in non-small cell lung cancer. Int. J. Radiat. Oncol. Biol. Phys. 113, 335–344 (2022).
Mei, P. et al. High tumor mutation burden is associated with DNA damage repair gene mutation in breast carcinomas. Diagn. Pathol. 15, 50 (2020).
Singleton, D. C., Macann, A. & Wilson, W. R. Therapeutic targeting of the hypoxic tumour microenvironment. Nat. Rev. Clin. Oncol. 18, 751–772 (2021).
Harris, B. H. L., Barberis, A., West, C. M. L. & Buffa, F. M. Gene expression signatures as biomarkers of tumour hypoxia. Clin. Oncol. 27, 547–560 (2015).
Yang, L. & West, C. Hypoxia gene expression signatures as predictive biomarkers for personalising radiotherapy. Br. J. Radiol. 92, 20180036 (2019).
Irlam, J. & West, C. M. L. Time to rethink commercialisation: spin out or lose out? Clin. Oncol. 34, 439–441 (2022).
Grégoire, V. et al. OC-0278 Accelerated CH-RT with/without nimorazole for p16- HNSCC: the randomized DAHANCA 29-EORTC 1219 trial. Radiother. Oncol. 161, S187–S188 (2021).
Busk, M., Overgaard, J. & Horsman, M. R. Imaging of tumor hypoxia for radiotherapy: current status and future directions. Semin. Nucl. Med. 50, 562–583 (2020).
O’Connor, J. P. B., Robinson, S. P. & Waterton, J. C. Imaging tumour hypoxia with oxygen-enhanced MRI and BOLD MRI. Br. J. Radiol. 92, 20180642 (2019).
Welz, S. et al. Dose escalation to hypoxic subvolumes in head and neck cancer: a randomized phase II study using dynamic [18F]FMISO PET/CT. Radiother. Oncol. 171, 30–36 (2022).
Kim, J. J. & Tannock, I. F. Repopulation of cancer cells during therapy: an important cause of treatment failure. Nat. Rev. Cancer 5, 516–525 (2005).
Withers, H. R., Taylor, J. M. G. & Maciejewski, B. The hazard of accelerated tumor clonogen repopulation during radiotherapy. Acta Oncol. 27, 131–146 (1988).
Buffa, F. M. et al. Molecular marker profiles predict locoregional control of head and neck squamous cell carcinoma in a randomized trial of continuous hyperfractionated accelerated radiotherapy. Clin. Cancer Res. 10, 3745–3754 (2004).
Xu, Y. et al. Prediction of response to radiotherapy by characterizing the transcriptomic features in clinical tumor samples across 15 cancer types. Comput. Intell. Neurosci. 2022, 5443709 (2022).
de Mey, S., Dufait, I. & De Ridder, M. Radioresistance of human cancers: clinical implications of genetic expression signatures. Front. Oncol. 11, 761901 (2021).
Baumann, M. et al. Radiation oncology in the era of precision medicine. Nat. Rev. Cancer 16, 234–249 (2016).
Arnold, C. R., Mangesius, J., Skvortsova, I. I. & Ganswindt, U. The role of cancer stem cells in radiation resistance. Front. Oncol. 10, 164 (2020).
Gaedicke, S. et al. Noninvasive positron emission tomography and fluorescence imaging of CD133+ tumor stem cells. Proc. Natl Acad. Sci. USA 111, E692–E701 (2014).
Ghaffari, H., Beik, J., Talebi, A., Mahdavi, S. R. & Abdollahi, H. New physical approaches to treat cancer stem cells: a review. Clin. Transl. Oncol. 20, 1502–1521 (2018).
Wang, J. et al. Notch promotes radioresistance of glioma stem cells. Stem Cell 28, 17–28 (2010).
Thippu Jayaprakash, K. & Michael, A. Notch inhibition: a promising strategy to improve radiosensitivity and curability of radiotherapy. Clin. Oncol. 33, e44–e49 (2021).
Wang, K. et al. Targeting cancer stem cells by disulfiram and copper sensitizes radioresistant chondrosarcoma to radiation. Cancer Lett. 505, 37–48 (2021).
Abhold, E. L. et al. EGFR kinase promotes acquisition of stem cell-like properties: a potential therapeutic target in head and neck squamous cell carcinoma stem cells. PLoS ONE 7, e32459 (2012).
van der Heijden, M. et al. Biological determinants of chemo-radiotherapy response in HPV-negative head and neck cancer: a multicentric external validation. Front. Oncol. 9, 1470 (2020).
Efstathiou, J. A. et al. Impact of immune and stromal infiltration on outcomes following bladder-sparing trimodality therapy for muscle-invasive bladder cancer. Eur. Urol. 76, 59–68 (2019).
Yarchoan, M., Hopkins, A. & Jaffee, E. M. Tumor mutational burden and response rate to PD-1 inhibition. N. Engl. J. Med. 377, 2500–2501 (2017).
Eder, T. et al. Interference of tumour mutational burden with outcome of patients with head and neck cancer treated with definitive chemoradiation: a multicentre retrospective study of the German Cancer Consortium Radiation Oncology Group. Eur. J. Cancer 116, 67–76 (2019).
De Courcy, L., Bezak, E. & Marcu, L. G. Gender-dependent radiotherapy: the next step in personalised medicine? Crit. Rev. Oncol. Hematol. 147, 102881 (2020).
Pointer, K. B., Pitroda, S. P. & Weichselbaum, R. R. Radiotherapy and immunotherapy: open questions and future strategies. Trends Cancer 8, 9–20 (2021).
Parker, C. C. et al. Radiotherapy to the primary tumour for newly diagnosed, metastatic prostate cancer (STAMPEDE): a randomised controlled phase 3 trial. Lancet 392, 2353–2366 (2018).
You, R. et al. Efficacy and safety of locoregional radiotherapy with chemotherapy vs chemotherapy alone in de novo metastatic nasopharyngeal carcinoma: a multicenter phase 3 randomized clinical trial. JAMA Oncol. 6, 1345–1352 (2020).
Spina, C. S. & Drake, C. G. Mechanisms of immune modulation by radiation. Semin. Radiat. Oncol. 31, 205–216 (2021).
Antonia, S. J. et al. Durvalumab after chemoradiotherapy in stage III non–small-cell lung cancer. N. Engl. J. Med. 377, 1919–1929 (2017).
Kelly, R. J. et al. Adjuvant nivolumab in resected esophageal or gastroesophageal junction cancer. N. Engl. J. Med. 384, 1191–1203 (2021).
Lee, N. Y. et al. Avelumab plus standard-of-care chemoradiotherapy versus chemoradiotherapy alone in patients with locally advanced squamous cell carcinoma of the head and neck: a randomised, double-blind, placebo-controlled, multicentre, phase 3 trial. Lancet Oncol. 22, 450–462 (2021).
Marciscano, A. E. et al. Elective nodal irradiation attenuates the combinatorial efficacy of stereotactic radiation therapy and immunotherapy. Clin. Cancer Res. 24, 5058–5071 (2018).
McLaughlin, M. et al. Inflammatory microenvironment remodelling by tumour cells after radiotherapy. Nat. Rev. Cancer 20, 203–217 (2020).
Price, J. M. et al. Pretreatment lymphocyte count predicts benefit from concurrent chemotherapy with radiotherapy in oropharyngeal cancer. J. Clin. Oncol. 40, 2203–2212 (2022).
Chen, D. et al. Absolute lymphocyte count predicts abscopal responses and outcomes in patients receiving combined immunotherapy and radiation therapy: analysis of 3 phase 1/2 trials. Int. J. Radiat. Oncol. Biol. Phys. 108, 196–203 (2020).
Ngwa, W. et al. Using immunotherapy to boost the abscopal effect. Nat. Rev. Cancer 18, 313–322 (2018).
Hall, J. et al. Ionizing radiation biomarkers in epidemiological studies – an update. Mutat. Res. Rev. Mutat. Res. 771, 59–84 (2017).
West, C. & Rosenstein, B. S. Establishment of a radiogenomics consortium. Int. J. Radiat. Oncol. Biol. Phys. 76, 1295–1296 (2010).
Kerns, S. L. et al. Radiogenomics consortium genome-wide association study meta-analysis of late toxicity after prostate cancer radiotherapy. J. Natl Cancer Inst. 112, 179–190 (2020).
Barnett, G. C. et al. Incorporating genetic biomarkers into predictive models of normal tissue toxicity. Clin. Oncol. 27, 579–587 (2015).
Azria, D. et al. Radiation-induced CD8 T-lymphocyte apoptosis as a predictor of breast fibrosis after radiotherapy: results of the prospective multicenter French trial. EBioMedicine 2, 1965–1973 (2015).
Mackay, R. I. & Hendry, J. H. The modelled benefits of individualizing radiotherapy patients’ dose using cellular radiosensitivity assays with inherent variability. Radiother. Oncol. 50, 67–75 (1999).
Mehanna, H. et al. De-escalation after DE-ESCALATE and RTOG 1016: a Head and Neck Cancer InterGroup framework for future de-escalation studies. J. Clin. Oncol. 38, 2552–2557 (2020).
Bristow, R. G. et al. Combining precision radiotherapy with molecular targeting and immunomodulatory agents: a guideline by the American Society for Radiation Oncology. Lancet Oncol. 19, e240–e251 (2018).
Ma, D. J. et al. Phase II evaluation of aggressive dose de-escalation for adjuvant chemoradiotherapy in human papillomavirus-associated oropharynx squamous cell carcinoma. J. Clin. Oncol. 37, 1909–1918 (2019).
Riaz, N. et al. Precision radiotherapy: reduction in radiation for oropharyngeal cancer in the 30 ROC trial. J. Natl Cancer Inst. 113, 742–751 (2020).
Knegjens, J. L. et al. Tumour volume as prognostic factor in chemoradiation for advanced head and neck cancer. Head. Neck 33, 375–382 (2011).
van Laar, M., van Amsterdam, W. A. C., van Lindert, A. S. R., de Jong, P. A. & Verhoeff, J. J. C. Prognostic factors for overall survival of stage III non-small cell lung cancer patients on computed tomography: a systematic review and meta-analysis. Radiother. Oncol. 151, 152–175 (2020).
Kurth, I., Peitzsch, C., Baumann, M. & Dubrovska, A. The role of cancer stem cells in tumor radioresistance. Cancer Stem Cell https://doi.org/10.1016/j.radonc.2022.02.009 (2014).
Linge, A. et al. Independent validation of tumour volume, cancer stem cell markers and hypoxia-associated gene expressions for HNSCC after primary radiochemotherapy. Clin. Transl. Radiat. Oncol. 16, 40–47 (2019).
Linge, A. et al. Low cancer stem cell marker expression and low hypoxia identify good prognosis subgroups in HPV(-) HNSCC after postoperative radiochemotherapy: a multicenter study of the DKTK-ROG. Clin. Cancer Res. 22, 2639–2649 (2016).
Löck, S. et al. Biomarker signatures for primary radiochemotherapy of locally advanced HNSCC – Hypothesis generation on a multicentre cohort of the DKTK-ROG. Radiother. Oncol. 169, 8–14 (2022).
Wiedenmann, N. E. et al. Serial [18F]-fluoromisonidazole PET during radiochemotherapy for locally advanced head and neck cancer and its correlation with outcome. Radiother. Oncol. 117, 113–117 (2015).
Eschrich, S. A. et al. Validation of a radiosensitivity molecular signature in breast cancer. Clin. Cancer Res. 18, 5134–5143 (2012).
Torres-Roca, J. F. et al. A molecular signature of radiosensitivity (RSI) is an RT-specific biomarker in prostate cancer. Int. J. Radiat. Oncol. Biol. Phys. 90, S157 (2014).
Creelan, B., Eschrich, S. A., Fulp, W. J. & Torres-Roca, J. F. A gene expression platform to predict benefit from adjuvant external beam radiation in resected non-small cell lung cancer. Int. J. Radiat. Oncol. Biol. Phys. 90, S76–S77 (2014).
Ahmed, K. A. et al. Differences between colon cancer primaries and metastases using a molecular assay for tumor radiation sensitivity suggest implications for potential oligometastatic SBRT patient selection. Int. J. Radiat. Oncol. Biol. Phys. 92, 837–842 (2015).
Strom, T. et al. Radiosensitivity index predicts for survival with adjuvant radiation in resectable pancreatic cancer. Radiother. Oncol. 117, 159–164 (2015).
Ahmed, K. A. et al. The radiosensitivity index predicts for overall survival in glioblastoma. Oncotarget 6, 34414–34422 (2015).
Torres-Roca, J. F. et al. Integration of a radiosensitivity molecular signature into the assessment of local recurrence risk in breast cancer. Int. J. Radiat. Oncol. Biol. Phys. 93, 631–638 (2015).
Strom, T. et al. Regional radiation therapy impacts outcome for node positive cutaneous melanoma. J. Natl Compr. Canc. Netw. 15, 473–82. (2017).
Mohammadi, H. et al. Using the radiosensitivity index (RSI) to predict pelvic failure in endometrial cancer treated with adjuvant radiation therapy. Int. J. Radiat. Oncol. Biol. Phys. 106, 496–502 (2020).
Sun, C., Zhang, M., Qiao, Q. & Wang, Y. Integrating intrinsic radiosensitivity and immune status for predicting benefits of radiotherapy in head and neck squamous cell carcinoma. Med. Sci. Monit. 27, 1–11 (2021).
Khan, M. T. et al. Developing tumor radiosensitivity signatures using LncRNAs. Radiat. Res. 195, 324–333 (2021).
Yang, G. et al. Genomic identification of sarcoma radiosensitivity and the clinical implications for radiation dose personalization. Transl. Oncol. 14, 101165 (2021).
Ahmed, K. A. et al. Utilizing the genomically adjusted radiation dose (GARD) to personalize adjuvant radiotherapy in triple negative breast cancer management. EBioMedicine 47, 163–169 (2019).
Scott, J. G. et al. Pan-cancer prediction of radiotherapy benefit using genomic-adjusted radiation dose (GARD): a cohort-based pooled analysis. Lancet Oncol. 22, 1221–1229 (2021).
Cui, Y., Li, B., Pollom, E. L., Horst, K. C. & Li, R. Integrating radiosensitivity and immune gene signatures for predicting benefit of radiotherapy in breast cancer. Clin. Cancer Res. 24, 4754–4762 (2018).
Speers, C. et al. A signature that may be predictive of early versus late recurrence after radiation treatment for breast cancer that may inform the biology of early, aggressive recurrences. Int. J. Radiat. Oncol. Biol. Phys. 108, 686–696 (2020).
Erho, N. et al. Discovery and validation of a prostate cancer genomic classifier that predicts early metastasis following radical prostatectomy. PLoS ONE 8, e66855 (2013).
Cooperberg, M. R. et al. Combined value of validated clinical and genomic risk stratification tools for predicting prostate cancer mortality in a high-risk prostatectomy cohort. Eur. Urol. 67, 326–333 (2015).
Kim, S. II et al. Gene signature for prediction of radiosensitivity in human papillomavirus-negative head and neck squamous cell carcinoma. Radiat. Oncol. J. 38, 99–108 (2020).
Acknowledgements
J.M.P. is supported by The Taylor Family Foundation. C.M.L.W. is supported by the Manchester NIHR Biomedical Research Centre.
Author information
Authors and Affiliations
Contributions
All authors researched data for this article. J.M.P. and C.M.L.W. contributed substantially to discussion of the content, wrote the article, and reviewed and/or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
C.M.L.W. is co-founder of ManTRa Diagnostics, which seeks to commercialize hypoxia-associated gene signatures. J.M.P. and A.P. declare no competing interests.
Peer review
Peer review information
Nature Reviews Clinical Oncology thanks M. Baumann, F. Moraes, J. Torres-Roca and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
ClinicalTrials.gov: https://clinicaltrials.gov/
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Price, J.M., Prabhakaran, A. & West, C.M.L. Predicting tumour radiosensitivity to deliver precision radiotherapy. Nat Rev Clin Oncol 20, 83–98 (2023). https://doi.org/10.1038/s41571-022-00709-y
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41571-022-00709-y
This article is cited by
-
Cancer radioresistance is characterized by a differential lipid droplet content along the cell cycle
Cell Division (2024)
-
ANGPTL4 regulates ovarian cancer progression by activating the ERK1/2 pathway
Cancer Cell International (2024)
-
High expression of PPP1CC promotes NHEJ-mediated DNA repair leading to radioresistance and poor prognosis in nasopharyngeal carcinoma
Cell Death & Differentiation (2024)
-
Olaparib enhances sensitization of BRCA-proficient breast cancer cells to x-rays and protons
Breast Cancer Research and Treatment (2024)
-
Improving the efficacy of combined radiotherapy and immunotherapy: focusing on the effects of radiosensitivity
Radiation Oncology (2023)