Abstract
Tracking the progeny of single cells is necessary for building lineage trees that recapitulate processes such as embryonic development and stem cell differentiation. In classical lineage tracing experiments, cells are fluorescently labelled to allow identification by microscopy of a limited number of cell clones. To track a larger number of clones in complex tissues, fluorescent proteins are now replaced by heritable DNA barcodes that are read using next-generation sequencing. In prospective lineage tracing, unique DNA barcodes are introduced into single cells through genetic manipulation (using, for example, Cre-mediated recombination or CRISPR–Cas9-mediated editing) and tracked over time. Alternatively, in retrospective lineage tracing, naturally occurring somatic mutations can be used as endogenous DNA barcodes. Finally, single-cell mRNA-sequencing datasets that capture different cell states within a developmental or differentiation trajectory can be used to recapitulate lineages. In this Review, we discuss methods for prospective or retrospective lineage tracing and demonstrate how trajectory reconstruction algorithms can be applied to single-cell mRNA-sequencing datasets to infer developmental or differentiation tracks. We discuss how these approaches are used to understand cell fate during embryogenesis, cell differentiation and tissue regeneration.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sulston, J. E., Schierenberg, E., White, J. G. & Thomson, J. N. The embryonic cell lineage of the nematode caenorhabditis elegans. Dev. Biol. 100, 64–119 (1983).
Kretzschmar, K. & Watt, F. M. Lineage tracing. Cell 148, 33–45 (2012).
Shapiro, E., Biezuner, T. & Linnarsson, S. Single-cell sequencing-based technologies will revolutionize whole-organism science. Nat. Rev. Genet. 14, 618–630 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Navin, N. et al. Inferring tumor progression from genomic heterogeneity. Genome Res. 20, 68–80 (2010).
Sun, J. et al. Clonal dynamics of native haematopoiesis. Nature 514, 322–327 (2014). This work introduces the Sleeping Beauty transposase system for prospective genetic lineage tracing in the mouse and reveals a limited contribution of HSCs to steady-state blood production.
Till, J. E. & Mc, C. E. A direct measurement of the radiation sensitivity of normal mouse bone marrow cells. Radiat. Res. 14, 213–222 (1961).
Becker, A. J., Mc, C. E. & Till, J. E. Cytological demonstration of the clonal nature of spleen colonies derived from transplanted mouse marrow cells. Nature 197, 452–454 (1963).
Jacobsen, S. E. W. & Nerlov, C. Haematopoiesis in the era of advanced single-cell technologies. Nat. Cell Biol. 21, 2–8 (2019).
Laurenti, E. & Gottgens, B. From haematopoietic stem cells to complex differentiation landscapes. Nature 553, 418–426 (2018).
Rodriguez-Fraticelli, A. E. et al. Clonal analysis of lineage fate in native haematopoiesis. Nature 553, 212–216 (2018).
Metzger, D., Clifford, J., Chiba, H. & Chambon, P. Conditional site-specific recombination in mammalian cells using a ligand-dependent chimeric Cre recombinase. Proc. Natl Acad. Sci. USA 92, 6991–6995 (1995).
Littlewood, T. D., Hancock, D. C., Danielian, P. S., Parker, M. G. & Evan, G. I. A modified oestrogen receptor ligand-binding domain as an improved switch for the regulation of heterologous proteins. Nucleic Acids Res. 23, 1686–1690 (1995).
Feil, R. et al. Ligand-activated site-specific recombination in mice. Proc. Natl Acad. Sci. USA 93, 10887–10890 (1996).
Feil, R., Wagner, J., Metzger, D. & Chambon, P. Regulation of Cre recombinase activity by mutated estrogen receptor ligand-binding domains. Biochem. Biophys. Res. Commun. 237, 752–757 (1997).
Rinkevich, Y., Lindau, P., Ueno, H., Longaker, M. T. & Weissman, I. L. Germ-layer and lineage-restricted stem/progenitors regenerate the mouse digit tip. Nature 476, 409–413 (2011).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010).
Pei, W. et al. Polylox barcoding reveals haematopoietic stem cell fates realized in vivo. Nature 548, 456–460 (2017).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012). This work introduces the Cre–loxP-based Polylox system for inducible recombination of DNA barcodes in mice, which is used to study haematopoiesis.
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl Acad. Sci. USA 109, E2579–E2586 (2012).
McKenna, A. et al. Whole-organism lineage tracing by combinatorial and cumulative genome editing. Science 353, aaf7907 (2016).
Junker, J. P. et al. Massively parallel clonal analysis using CRISPR/Cas9 induced genetic scars. bioRXiv Preprint at https://doi.org/10.1101/056499 (2017).
Alemany, A., Florescu, M., Baron, C. S., Peterson-Maduro, J. & van Oudenaarden, A. Whole-organism clone tracing using single-cell sequencing. Nature 556, 108–112 (2018). This study presents scScarTrace, a method for prospective genetic lineage tracing in zebrafish using CRISPR–Cas9 to study the clonality of blood, brain, eyes and caudal fin.
Raj, B. et al. Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain. Nat. Biotechnol. 36, 442–450 (2018). This article describes the developmenr of scGESTALT, a method for prospective genetic lineage tracing in zebrafish using CRISPR–Cas9, and the characterization of lineage relationships between cell types in the adult brain.
Spanjaard, B. et al. Simultaneous lineage tracing and cell-type identification using CRISPR-Cas9-induced genetic scars. Nat. Biotechnol. 36, 469–473 (2018). This study introduces LINNAEUS, a method for prospective genetic lineage tracing in zebrafish using CRISPR–Cas9 and uses it to study multiple organs of zebrafish larvae.
Woo, K. & Fraser, S. E. Order and coherence in the fate map of the zebrafish nervous system. Development 121, 2595–2609 (1995).
Solek, C. M., Feng, S., Perin, S., Weinschutz Mendes, H. & Ekker, M. Lineage tracing of dlx1a/2a and dlx5a/6a expressing cells in the developing zebrafish brain. Dev. Biol. 427, 131–147 (2017).
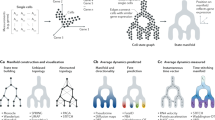
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017). This study presents Monocle2, an scRNA-seq-based trajectory reconstruction algorithm that connects single cells on the basis of transcriptional similarity to infer a developmental or differentiation pseudotime.
Xu, J. et al. Temporal-spatial resolution fate mapping reveals distinct origins for embryonic and adult microglia in zebrafish. Dev. Cell 34, 632–641 (2015).
Henninger, J. et al. Clonal fate mapping quantifies the number of haematopoietic stem cells that arise during development. Nat. Cell Biol. 19, 17–27 (2017).
Kissa, K. & Herbomel, P. Blood stem cells emerge from aortic endothelium by a novel type of cell transition. Nature 464, 112–115 (2010).
Bertrand, J. Y. et al. Haematopoietic stem cells derive directly from aortic endothelium during development. Nature 464, 108–111 (2010).
Tu, S. & Johnson, S. L. Fate restriction in the growing and regenerating zebrafish fin. Dev. Cell 20, 725–732 (2011).
Lin, X. et al. An ectoderm-derived myeloid-like cell population functions as antigen transporters for Langerhans cells in zebrafish epidermis. Dev. Cell 49, 605–617 (2019).
Perli, S. D., Cui, C. H. & Lu, T. K. Continuous genetic recording with self-targeting CRISPR–Cas in human cells. Science 353, aag0511 (2016).
Kalhor, R., Mali, P. & Church, G. M. Rapidly evolving homing CRISPR barcodes. Nat. Methods 14, 195–200 (2017).
Kalhor, R. et al. Developmental barcoding of whole mouse via homing CRISPR. Science 361, aat9804 (2018). This study reports the generation of a mouse line (MARC1) for prospective genetic lineage tracing using CRISPR–Cas9, which is used to build lineage trees of early embryogenesis.
Chan, M. M. et al. Molecular recording of mammalian embryogenesis. Nature 570, 77–82 (2019). The authors develop a method for prospective genetic lineage tracing using CRISPR–Cas9 in a mouse model to study lineage relationships during early embryogenesis.
Biasco, L. et al. In vivo tracking of human haematopoiesis reveals patterns of clonal dynamics during early and steady-state reconstitution phases. Cell Stem Cell 19, 107–119 (2016).
Woodworth, M. B., Girskis, K. M. & Walsh, C. A. Building a lineage from single cells: genetic techniques for cell lineage tracking. Nat. Rev. Genet. 18, 230–244 (2017).
Ludwig, L. S. et al. Lineage tracing in humans enabled by mitochondrial mutations and single-cell genomics. Cell 176, 1325–1339 (2019).
Zarrei, M., MacDonald, J. R., Merico, D. & Scherer, S. W. A copy number variation map of the human genome. Nat. Rev. Genet. 16, 172–183 (2015).
Abyzov, A. et al. Somatic copy number mosaicism in human skin revealed by induced pluripotent stem cells. Nature 492, 438–442 (2012).
Knouse, K. A., Wu, J. & Amon, A. Assessment of megabase-scale somatic copy number variation using single-cell sequencing. Genome Res. 26, 376–384 (2016).
McConnell, M. J. et al. Mosaic copy number variation in human neurons. Science 342, 632–637 (2013).
Cai, X. et al. Single-cell, genome-wide sequencing identifies clonal somatic copy-number variation in the human brain. Cell Rep. 8, 1280–1289 (2014).
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011).
Wang, Y. et al. Clonal evolution in breast cancer revealed by single nucleus genome sequencing. Nature 512, 155–160 (2014).
Casasent, A. K. et al. Multiclonal invasion in breast tumors identified by topographic single cell sequencing. Cell 172, 205–217 e212 (2018). This work demonstrates the capture of CNV by whole-genome amplification in single cells while preserving spatial information using laser-capture microdissection.
Katsonis, P. et al. Single nucleotide variations: biological impact and theoretical interpretation. Protein Sci. 23, 1650–1666 (2014).
Zong, C., Lu, S., Chapman, A. R. & Xie, X. S. Genome-wide detection of single-nucleotide and copy-number variations of a single human cell. Science 338, 1622–1626 (2012).
Lodato, M. A. et al. Somatic mutation in single human neurons tracks developmental and transcriptional history. Science 350, 94–98 (2015).
Leung, M. L. et al. Single-cell DNA sequencing reveals a late-dissemination model in metastatic colorectal cancer. Genome Res. 27, 1287–1299 (2017).
Xu, X. et al. Single-cell exome sequencing reveals single-nucleotide mutation characteristics of a kidney tumour. Cell 148, 886–895 (2012).
Ostertag, E. M. & Kazazian, H. H. Jr. Biology of mammalian L1 retrotransposons. Annu. Rev. Genet. 35, 501–538 (2001).
Hwang, B. et al. Lineage tracing using a Cas9-deaminase barcoding system targeting endogenous L1 elements. Nat. Commun. 10, 1234 (2019).
Evrony, G. D. et al. Single-neuron sequencing analysis of L1 retrotransposition and somatic mutation in the human brain. Cell 151, 483–496 (2012).
Evrony, G. D. et al. Cell lineage analysis in human brain using endogenous retroelements. Neuron 85, 49–59 (2015).
Ellegren, H. Microsatellites: simple sequences with complex evolution. Nat. Rev. Genet. 5, 435–445 (2004).
Reizel, Y. et al. Colon stem cell and crypt dynamics exposed by cell lineage reconstruction. PLOS Genet. 7, e1002192 (2011).
Reizel, Y. et al. Cell lineage analysis of the mammalian female germline. PLOS Genet. 8, e1002477 (2012).
Stewart, J. B. & Chinnery, P. F. The dynamics of mitochondrial DNA heteroplasmy: implications for human health and disease. Nat. Rev. Genet. 16, 530–542 (2015). In this article, the authors demonstrate that mitochondrial mutations can be detected by scRNA-seq and scATAC-seq and can be used to reconstruct clonal relationships of haematopoietic and leukaemic cells.
Nam, A. S. et al. Somatic mutations and cell identity linked by genotyping of transcriptomes. Nature 571, 355–360 (2019).
Kester, L. & van Oudenaarden, A. Single-cell transcriptomics meets lineage tracing. Cell Stem Cell 23, 166–179 (2018).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Park, J. et al. Single-cell transcriptomics of the mouse kidney reveals potential cellular targets of kidney disease. Science 360, 758–763 (2018).
Baron, C. S. et al. Single-cell transcriptomics reveal the dynamic of haematopoietic stem cell production in the aorta. Nat. Commun. 9, 2517 (2018).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Savas, P. et al. Single-cell profiling of breast cancer T cells reveals a tissue-resident memory subset associated with improved prognosis. Nat. Med. 24, 986–993 (2018).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018). The authors present RNA velocity, a trajectory reconstruction algorithm that uses the abundance of spliced and unspliced mRNA detected by scRNA-seq to predict the future state of individual cells.
Tang, F. et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods 6, 377–382 (2009).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Pijuan-Sala, B. et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566, 490–495 (2019).
Frieda, K. L. et al. Synthetic recording and in situ readout of lineage information in single cells. Nature 541, 107–111 (2017).
Lubeck, E., Coskun, A. F., Zhiyentayev, T., Ahmad, M. & Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat. Methods 11, 360–361 (2014).
Stahl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Shah, S. et al. Dynamics and spatial genomics of the nascent transcriptome by intron seqFISH. Cell 174, 363–376 (2018).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Vickovic, S. et al. High-density spatial transcriptomics arrays for in situ tissue profiling. bioRXiv Preprint at https://doi.org/10.1101/563338 (2018).
Zhang, L. et al. Lineage tracking reveals dynamic relationships of T cells in colorectal cancer. Nature 564, 268–272 (2018).
Mooijman, D., Dey, S. S., Boisset, J. C., Crosetto, N. & van Oudenaarden, A. Single-cell 5hmC sequencing reveals chromosome-wide cell-to-cell variability and enables lineage reconstruction. Nat. Biotechnol. 34, 852–856 (2016).
Houseman, E. A. et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinformatics 13, 86 (2012).
Accomando, W. P., Wiencke, J. K., Houseman, E. A., Nelson, H. H. & Kelsey, K. T. Quantitative reconstruction of leukocyte subsets using DNA methylation. Genome Biol. 15, R50 (2014).
Salas, L. A. et al. Tracing human stem cell lineage during development using DNA methylation. Genome Res. 28, 1285–1295 (2018).
Acknowledgements
The authors thank A. Alemany and J. Yeung for valuable feedback on the manuscript.
Author information
Authors and Affiliations
Contributions
C.S.B. researched data and wrote the manuscript. A.v.O. reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing financial interests.
Additional information
Peer review information
Nature Reviews Molecular Cell Biology thanks Bin Zhou and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Long-term haematopoietic stem cells
-
(LT-HSCs). Blood stem cells able to self-renew and differentiate into all types of mature blood cells.
- Primed cells
-
Differentiating cells with a determined cell fate.
- Lateral plate mesoderm
-
A subset of mesodermal tissue found in developing embryos that will form the body walls and circulatory system.
- Primitive wave of haematopoiesis
-
A process by which the embryo generates transient haematopoietic cells, which are necessary for embryonic development.
- Definitive wave of haematopoiesis
-
A process by which definitive adult haematopoietic stem cells are generated during embryogenesis.
- Dome stage
-
A developmental stage of zebrafish embryos that is reached at 4.3 h of development.
- Nuc-seq
-
RNA-sequencing technology for nuclear RNA capture from frozen tissue samples.
- Minimum spanning tree
-
The shortest way to connect all edges of a graph.
- Pseudotime
-
A quantitative measure of biological progression through a process such as cell differentiation.
Rights and permissions
About this article
Cite this article
Baron, C.S., van Oudenaarden, A. Unravelling cellular relationships during development and regeneration using genetic lineage tracing. Nat Rev Mol Cell Biol 20, 753–765 (2019). https://doi.org/10.1038/s41580-019-0186-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-019-0186-3
This article is cited by
-
Single-cell lineage tracing with endogenous markers
Biophysical Reviews (2024)
-
PhyloVelo enhances transcriptomic velocity field mapping using monotonically expressed genes
Nature Biotechnology (2023)
-
Reconstructing growth and dynamic trajectories from single-cell transcriptomics data
Nature Machine Intelligence (2023)
-
Retrospective analysis of enhancer activity and transcriptome history
Nature Biotechnology (2023)
-
In vivo fluorescent labeling and tracking of extracellular matrix
Nature Protocols (2023)