Abstract
Protein ubiquitination is a multifaceted post-translational modification that controls almost every process in eukaryotic cells. Recently, the Legionella effector SdeA was reported to mediate a unique phosphoribosyl-linked ubiquitination through successive modifications of the Arg42 of ubiquitin (Ub) by its mono-ADP-ribosyltransferase (mART) and phosphodiesterase (PDE) domains. However, the mechanisms of SdeA-mediated Ub modification and phosphoribosyl-linked ubiquitination remain unknown. Here we report the structures of SdeA in its ligand-free, Ub-bound and Ub–NADH-bound states. The structures reveal that the mART and PDE domains of SdeA form a catalytic domain over its C-terminal region. Upon Ub binding, the canonical ADP-ribosyltransferase toxin turn-turn (ARTT) and phosphate-nicotinamide (PN) loops in the mART domain of SdeA undergo marked conformational changes. The Ub Arg72 might act as a ‘probe’ that interacts with the mART domain first, and then movements may occur in the side chains of Arg72 and Arg42 during the ADP-ribosylation of Ub. Our study reveals the mechanism of SdeA-mediated Ub modification and provides a framework for further investigations into the phosphoribosyl-linked ubiquitination process.
Similar content being viewed by others
Main
Ubiquitination is one of the most prevalent protein modifications in eukaryotic cells, regulating a wide array of essential cellular processes1,2. Ubiquitination is carried out by a three-enzyme cascade (E1, E2 and E3), and results in the transfer of ubiquitin (Ub) to a lysine residue of the substrate3. Prokaryotes do not contain the Ub-proteasome system, but a variety of bacterial pathogens adopt intricate mechanisms to influence the host Ub system to support their own survivals4. Legionella pneumophila, the causative agent of a potentially fatal pneumonia known as Legionnaires’ disease5,6, can survive and replicate within host cells by creating a vacuole7,8,9. The biogenesis of the Legionella-containing vacuole is based on the approximately 300 Legionella substrates (effectors) that are translocated into host cells10,11,12,13. Notably, recent studies have identified that the SidE family effectors from L. pneumophila could catalyse Ub transfer to several endoplasmic reticulum-associated human Rab GTPases14 and the endoplasmic reticulum protein reticulon 4 (RTN4)15, using a unique approach that differs from the canonical ubiquitination pathway16. By targeting RTN4, the SidE family proteins could control the dynamics of tubular endoplasmic reticulum and promote structural transformations of the tubules15.
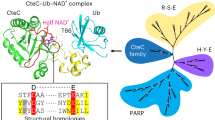
The SidE family members are large proteins (approximately 1,500 residues)6 (Extended Data Fig. 1), such as SdeA, which contains a deubiquitinase (DUB) domain (SdeA DUB)17, a PDE domain (SdeA PDE), an mART domain (SdeA mART) and a C-terminal domain (SdeA CTD) (Fig. 1a). During the ubiquitination process, the R42 residue of Ub is first ADP-ribosylated with NAD+ by SdeA mART, and then the phosphodiester bond of the ADP-ribosylated Ub (ADPR-Ub) is cleaved by SdeA PDE to make phosphoribosyl Ub (PR-Ub)15,16. PR-Ub can either remain by itself or be linked via a phosphodiester bond to the hydroxyl group of serine residues in either the substrates or SdeA itself in a reaction catalysed by SdeA PDE. However, structural investigation of the mechanism behind these modifications is required.
Overall structure of SdeA
To understand the mechanism underlying SdeA-mediated Ub modifications, we first solved the crystal structure of a truncated SdeA(amino acids 231–1190, hereafter called SdeA(231–1190)) (Extended Data Table 1) at 3.39 Å resolution. This region of the protein includes SdeA PDE, SdeA mART and a part of SdeA CTD (SdeA pCTD). SdeA(231–1190) exhibited ubiquitination activities that were similar to full-length SdeA (Extended Data Fig. 2a, b). The crystal belonged to the C2221 space group and two SdeA molecules were found in the asymmetric unit. However, results from the PISA server18, gel filtration chromatography and analytical ultracentrifuge analysis indicated that SdeA mainly exists as a monomer in solution (Extended Data Fig. 2d, e). Notably, the two SdeA molecules in the asymmetric unit exhibited obvious conformational variations, with a core root mean square deviation (r.m.s.d.) of 1.82 Å among 793 Cα atoms. Superimposition of the two molecules revealed a better alignment of their PDE and mART domains, but prominent differences between their pCTDs, indicating that SdeA pCTD might be flexible in solution (Extended Data Fig. 2f). In the SdeA(231–1190) structure, SdeA mART and SdeA PDE interact primarily through hydrophobic interactions and form a catalytic core (Fig. 1b, c and Extended Data Fig. 3a–d), which sits on top of SdeA pCTD. In contrast to the more conserved PDE and mART domains, SdeA pCTD represented an overall novel fold completely composed of α-helices (Fig. 1b), which could be divided into two subdomains on two sides below SdeA mART (Extended Data Fig. 2g, h).
Structure of the mART domain of SdeA
SdeA mART contains a typical Rossmann fold19 (Fig. 2a), and exhibits the basic characteristics that are conserved among all known bacterial mART toxins20,21. SdeA mART folds in a two-lobe structure with an N-terminal α-helical lobe (residues 594–758) and a C-terminal β-sandwich lobe (Fig. 2a). Notably, structural alignment revealed several unique features in the SdeA mART structure (Extended Data Fig. 4a–d). First, the canonical ARTT and PN loops in SdeA mART are both in different conformations from those of the other mART proteins. Second, the N-terminal α-helical lobe of SdeA mART also shows an overall fold different from those of the others. Third, two consecutive protruding helices are connected by a loop (residues 789–797, here named the plug), which inserts into and interacts with SdeA PDE. This plug loop is, to our knowledge, unique among known mART structures (Fig. 2a and Extended Data Figs. 3a–d, 4a–d). Notably, the much higher ADP-ribosylation activity of SdeA(193–935)H277A when compared to SdeA(597–935) (the single SdeA mART) indicates that unlike SdeA PDE, which is fully active as a single domain (that is, it is able to process ADPR-Ub to PR-Ub and catalyse ubiquitination (Extended Data Fig. 3e)), SdeA mART needs to be stabilized by SdeA PDE to be active. Moreover, deletion of the plug loop of SdeA mART markedly reduced the activity of SdeA(193–998) (∆NC SdeA)16 (Fig. 2f and Extended Data Figs. 3f, 4e–h).
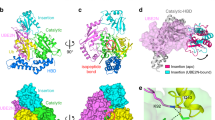
a, Structure of SdeA mART. b, Overall structure of the SdeA(231–1190)–Ub complex. SdeA is coloured as in Fig. 1a; three Ub molecules are coloured magenta. c, Structural superimposition of apo-SdeA mART (grey) and the SdeA mART–Ub complex (coloured as in b) shows conformational changes. d, Enlarged view of the outlined region in c, major SdeA mART residues which move upon Ub binding are shown in stick representation. e, Expanded view showing a superimposition of regions of the ARTT and PN loops before and after Ub binding. The conformational change is highlighted with a blue arrow. f, Relative amounts of the unmodified Ub, PR-Ub and ADPR-Ub were studied by top-down liquid chromatography–mass spectrometry analysis after they were isolated from the reaction mixtures. WT, wild type.
SdeA mART has a unique Ub-binding mode
To our knowledge, SdeA mART is the first mART domain ever reported to catalyse ADP-ribosylation of Ub. To understand the molecular mechanism by which SdeA mART recognizes and mediates ADP-ribosylation of Ub, we solved the structure of the SdeA(231–1190)–Ub complex by mixing the two proteins directly in molar ratios of 1:4, 1:6 and 1:8 (SdeA:Ub) before crystallization. Notably, the SdeA(231–1190) structure was solved with three bound Ubs, one at SdeA mART, and the other two at SdeA pCTD (Fig. 2b and Extended Data Fig. 5a–d). The binding of Ub caused a prominent conformational change (r.m.s.d. = 2.24 Å among 830 Cα atoms) of SdeA, particularly in the N-terminal region of SdeA pCTD (Extended Data Fig. 5b), again demonstrating the flexibility of SdeA pCTD. However, SdeA pCTD was dispensable for the in vitro activity of SdeA, as SdeA(193–935) was fully capable of Ub modification and RAB33B ubiquitination (Extended Data Figs. 4i, 5a). Therefore, we focused on the Ub bound to SdeA mART in the subsequent studies.
To our knowledge, the Ub–SdeA mART binding represents a novel binding mode that differs from all known Ub-protein interactions1,22. Burying a surface area of 607.8 Å2, the overall structure of SdeA mART in the complex is similar to that of apo-SdeA mART, with an r.m.s.d. of 1.04 Å among 293 Cα atoms (Fig. 2c). Nonetheless, the ARTT and PN loops undergo marked conformational changes upon Ub binding (Fig. 2d, e). In response to Ub binding, the Cα atom of SdeAE860 in the ARTT loop moves towards Ub by 6.0 Å to interact with UbR72 via its side chain, which is simultaneously stabilized by SdeAQ713 (Figs. 2d, 3a). In addition to UbR72, another Ub residue that is essential for its recognition is UbR74, which inserts into a negatively charged groove on SdeA mART by forming electrostatic interactions with the side chains of SdeAD691 and SdeAD707 from the α-helical lobe (Fig. 3a and Extended Data Fig. 6a, b). Meanwhile, the Cα atom of SdeAW832 in the PN loop moves 3.8 Å away from its original position upon Ub binding (Fig. 2d). Notably, SdeAW832 acts as a gate for Ub binding, as it occupies the site in the loop that links β3 and α2 of Ub in the apo-SdeA structure (Fig. 2d). Together, the ARTT and PN loops and the α-helical lobe of SdeA mART contribute to Ub recognition.
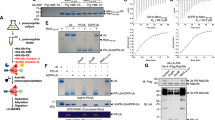
a, Detailed interactions between Ub (magenta) and SdeA mART (green). Polar interactions are represented by red dashed lines. b, Wild-type ∆NC SdeA or the mutant proteins were incubated with Ub, RAB33B and NAD+ for 30 min at 37 °C. The samples were then subjected to SDS–PAGE, with Coomassie staining (top) and phospho-specific staining with Pro-Q Diamond (bottom). c, Galactose-inducible pYES2 plasmids containing wild-type ∆NC SdeA or the mutants were transformed into yeast W303 strain. Five microlitres of cells in three tenfold serial dilutions were spotted on both glucose- and galactose-containing plates lacking uracil for two days before image acquisition. d, Wild-type Ub and the indicated mutants were incubated with ∆NC SdeA, RAB33B and NAD+ for 30 min at 37 °C. The samples were then treated as in b. Uncropped blots and gel images for b and d are shown in Supplementary Fig. 1. e, Superimposition of the structures of SdeA–Ub and SdeA–Ub–NADH. The whole SdeA–Ub complex is shown in grey. SdeA and Ub in the SdeA–Ub–NADH complex are shown as in a. f, MST assays of the binding of NAD+ to wild-type SdeA(231–1190) and the indicated mutants. Means are indicated by horizontal lines and individual values from three independent experiments are shown with their markers. Binding curves and Kd values are also shown. NA, for the N723A, R729A and R766A mutants, Kd could not be determined.
Of the SdeA residues that interact with Ub, SdeAE860 has been shown to have an essential role in ADP-ribosylation of Ub14,16. Moreover, SdeA mutant proteins (D691A/D707A, T831D and W832A) all showed decreased activities in Ub phosphoribosylation and substrate ubiquitination, and these mutations also inhibited the yeast toxicity of SdeA (Fig. 3a–c and Extended Data Fig. 7a, b). Consistently, the UbR74A mutant showed no modification upon treatment with SdeA, as has previously been observed in UbR42A and UbR72A mutants16 (Fig. 3d). We also found that these three Ub mutants were defective in the SdeA mART-catalysed reaction, as they could not be ADP-ribosylated by SdeA(193–935)H277A (Fig. 2f and Extended Data Fig. 4f).
The interaction pair of SdeAE830 and UbK6, and the I44 patch of Ub seemed to have a minor role in SdeA mART-Ub binding, as mutations of these residues did not influence Ub modification or substrate ubiquitination (Fig. 3b–d). SdeA mART-mediated ADP-ribosylation of Ub is specific, as Ub-like proteins SUMO1 and NEDD8 could not be ADP-ribosylated by SdeA(193–935)H277A, which is consistent with their structural differences (Extended Data Fig. 6c, d). Structural alignments also suggested that SdeA mART is able to modify both the proximal and distal moieties of K63-, K48-, K11- and M1-linked diubiquitins (Extended Data Fig. 5e–h), a finding that is consistent with a previous study23.
To further investigate the Ub modification mechanism of SdeA, we solved the structure of SdeA(231–1190)–Ub–NADH complex using a soaking approach (Extended Data Table 1). NADH is reported to be an inhibitor of the mART activity of the enzyme component of the iota-toxin from Clostridium perfringens24 and is predicted to occupy the binding site of NAD+. Similarly, NADH also had an inhibitory effect on the SdeA-catalysed ubiquitination of RAB33B (Extended Data Fig. 2c). In the structure of the complex, NADH in the cavity forms a ring-like conformation (Fig. 3e and Extended Data Fig. 7e). The overall structure of SdeA mART–Ub remains unchanged upon NADH binding (r.m.s.d. of 0.36 Å among 307 Cα atoms). Marked conformational changes only occur in the side chains of SdeAR766 and SdeAW832, which interact with the phosphate group of NADH through hydrogen bonding and with the nicotinamide ring through π–π interactions, respectively (Fig. 3e). Mutations of the NADH-binding residues of SdeA all impaired the Ub modification, substrate ubiquitination and yeast toxicity of SdeA (Fig. 3b, e and Extended Data Fig. 7c, d, f). The decreased binding to NAD+ by the mutant proteins was confirmed using the microscale thermophoresis (MST) assay (Fig. 3f). Notably, although the SdeA variants with mutations in the Ub- or NADH-binding sites were not able to catalyse ubiquitination with normal Ubs, they could utilize and process ADPR-Ub to complete the ubiquitination reaction (Extended Data Fig. 7g–i), indicating that none of the mutations interfere with the activity of SdeA PDE.
Suggested mechanism of SdeA mART ADP-ribosylation
An SN1 nucleophilic reaction mechanism of the ADP-ribosylation of arginine residues by mART proteins has been proposed by several studies25,26. However, a notable point of the SdeA–Ub–NADH complex structure is the 11.7 Å distance between the nucleophile (NH1/2 atom of UbR42) and the electrophile (C1D of the N-ribose, the ribose linked to the nicotinamide group of NAD+), which is too far for an SN1 attack (Fig. 4a). Nevertheless, UbR72, which forms polar interactions with the side chain of SdeAE860, the backbone carbonyl of SdeAQ713 and the O3D atom of the NADH ribose group, is much closer to the site of the electrophile with a distance of 4.6 Å (Fig. 4a). Because we could not obtain the structure of the SdeA–Ub–NAD+ complex, we performed molecular dynamics simulations with NADH replaced by NAD+ in the complex, during which the positions of the side chains of UbR42 and UbR72, and NAD+ remained almost unchanged (Fig. 4b and Extended Data Fig. 7j–m).
a, The conformations of UbR42 and UbR72 in the SdeA mART–Ub–NADH complex are shown. b, Molecular dynamics simulation results of the SdeA mART–Ub–NAD+ complex (left) and the SdeA mART–Ub–intermediate system (right). c, In vitro glutathione S-transferase (GST) pull-down assays to detect the IcmS binding region of SdeA. d, Superposition of the gel filtration chromatograms of SdeA(1191–1350), the IcmS–IcmW–DotLc complex and a mixture of both. Coomassie blue staining of the peak fractions after SDS–PAGE are shown on the right. c–d, Uncropped gel images are shown in Supplementary Fig. 1.
In the proposed SN1 mechanism, the highly folded and strained conformation of the nicotinamide mononucleotide region of NAD+ induces an equilibrium shift towards formation of an oxocarbenium cation intermediate (NAD+ with its nicotinamide group cleaved, hereafter referred to as the intermediate)25,26. Therefore, we also performed molecular dynamics simulations with NADH replaced by the intermediate in the SdeA mART–Ub–NADH structure (Extended Data Fig. 7j–m). Notably, after the simulation, UbR72 moved towards the ARTT loop and away from the intermediate. Instead, UbR42 entered the active site, occupied the original position of UbR72, and formed electrostatic interactions with SdeAE860 (Fig. 4b). After the system reached equilibrium, the average distance between the nucleophile (UbR42) and the electrophile was 4.46 Å (Fig. 4b and Extended Data Fig. 7m). Consistent with the conformational change observed in molecular dynamics simulation, the conformation of UbR42 is highly variable among the available structures of Ub (Extended Data Fig. 8a).
Together with the structural and biochemical results, we propose that during the catalytic process, UbR72 might function as a ‘probe’, together with UbR74, by anchoring Ub on SdeA mART. After cleavage of the nicotinamide group from NAD+, the strain in the highly folded structure of the intermediate would be alleviated, which might destabilize the binding of UbR72, causing it to leave. This in turn could facilitate the approach of UbR42 to the active site. However, the exact catalytic cycle still needs further investigation. Moreover, SdeA mART-produced ADPR-Ub will be processed to PR-Ub, and linked to the target protein by SdeA PDE, the mechanism of which can be gained from the structure of SdeA PDE with its substrates27,28.
SdeA CTD interacts with IcmS–IcmW
Because the N-terminal SdeA DUB and approximately 300 residues of the C-terminal were not included in our crystallized construct, we next studied the solution structure of four constructs of SdeA using the small-angle X-ray scattering (SAXS) method to investigate the spatial position of SdeA PDE–SdeA mART within SdeA (Extended Data Table 2 and Extended Data Fig. 8c–h). Both the scattering profile comparison and the reconstructed molecular envelope indicated that the crystal structure of SdeA(231–1190) is similar to its structure in solution (Extended Data Fig. 8d, e). Superimposing the envelope of SdeA(231–1190) onto the envelope of SdeA(1–1190) further revealed the extra electron density within the envelope of SdeA(1–1190) for SdeA DUB (Extended Data Fig. 8f), which forms a triangle-shaped catalytic core with SdeA mART and SdeA PDE. The envelope of the C-terminal SdeA(1092–1496) indicated the helical-bundle shape of this region (Extended Data Fig. 8g). Further superimposition of the SAXS envelopes of SdeA(1–1499), SdeA(1–1190), SdeA(1092–1496) and the crystal structure of SdeA(231–1190) reconfirmed the positions of SdeA DUB and SdeA CTD (Extended Data Fig. 8h). Notably, in vitro binding assays revealed that SdeA CTD is involved in binding to the adaptor protein complex IcmS–IcmW6,29,30 and the minimal binding region is SdeA(1191–1350), which could form a tight complex with IcmS–IcmW–DotLc (residues 656–798 of DotL)31 (Fig. 4c, d and Extended Data Fig. 8b). This suggests that SdeA CTD might function in the translocation of SdeA into host cells.
Discussion
An unresolved problem is how the ADPR-Ub is delivered from SdeA mART to SdeA PDE. One possibility is that two or more SdeAs might be close to each other in vivo and ADPR-Ubs produced by one SdeA mART could be used by the PDE domain of an adjacent one. Our study provides mechanistic insight into the structure and function of SdeA and serves as a foundation for the further studies of phosphoribosyl-linked ubiquitination.
Methods
Protein expression and purification
The full length and various segments of L. pneumophila SdeA were amplified by PCR and cloned into pGEX6p-1 or pET22b vectors to produce GST-tagged fusion proteins with a PreScission Protease cleavage site between GST and the target proteins, or His-tagged protein. The SdeA mutants were generated by two-step PCR and were subcloned, overexpressed and purified in the same way as the wild-type protein. The SdeA clone with a deletion of residues 789–797 was made by bridging PCR, and a GSG sequence was added between the two SdeA fragments. The proteins were expressed in Escherichia coli strain BL21 and induced using 0.2 mM isopropyl-β-d-thiogalactopyranoside (IPTG) when the cell density reached an OD600 nm of 0.8. For GST-tagged proteins, after growth at 16 °C for 12 h, the cells were collected, re-suspended in lysis buffer (1 × PBS, 2 mM dithiothreitol (DTT) and 1 mM phenylmethanesulfonyl fluoride) and lysed by sonication. The cell lysate was centrifuged at 20,000g for 45 min at 4 °C to remove cell debris. The supernatant was applied onto a self-packaged GST-affinity column (2 ml glutathione Sepharose 4B; GE Healthcare) and contaminant proteins were removed with wash buffer (lysis buffer plus 200 mM NaCl). The fusion protein was then digested with PreScission protease at 4 °C overnight. The protein with an additional five-amino-acid tag (GPLGS) at the N terminus was eluted with lysis buffer. The eluant was concentrated and further purified using a Superdex-200 (GE Healthcare) column equilibrated with a buffer containing 10 mM Tris–HCl pH 8.0, 200 mM NaCl, and 5 mM DTT. The purified protein was analysed by SDS–PAGE. The fractions containing the target protein were pooled and concentrated to 20 mg ml−1. Selenomethionine (Se–Met)-labelled SdeA was expressed in E. coli B834 (DE3) cells grown in M9 minimal medium supplemented with 60 mg l−1 Se–Met (Sigma-Aldrich) and specific amino acids: Ile, Leu and Val at 50 mg l−1; Lys, Phe and Thr at 100 mg l−1. The Se–Met protein was purified as described above. The SdeA(231–1190) segment was also cloned into pET22b vector, to make a construct with a C-terminal His tag, which was also used in purification and crystallization.
The fragment of human RAB33B cDNA (residues 1–229) was cloned into the MCS1 of pRSFDuet vector to produce His-tagged fusion protein. The fusion protein was induced in E. coli Rosetta (DE3) by 0.2 mM IPTG when the cell density reached an OD600 nm of 0.8. Recombinant His-tagged protein was purified using Ni-affinity column chromatography, ion exchange chromatography and was further subjected to gel filtration chromatography (Superdex-200 column) in buffer containing 10 mM Tris-HCl pH 8.0, 200 mM NaCl, 5 mM DTT.
All Ub mutants used in the ubiquitination assay were cloned into pGEX6p-1 vectors to produce GST-tagged fusion proteins. The proteins were purified according to the protocols for GST-tagged proteins, concentrated to 20 mg ml−1 and stored at −80 °C until use. In addition, the GST-tagged wild-type Ub was used in the self-ubiquitination experiments of SdeA(231–1190).
For the IcmS protein and its complexes, IcmS was cloned into pGEX6p-1 vector and purified as stated above. His-tagged IcmS and IcmW were cloned into the MCS1 and MCS2 sites of pRSFDuet vector, respectively. Then the IcmS–IcmW complex was purified as His-tagged proteins according to the protocol described above. For coexpression of IcmS–IcmW–DotLc complex, DotLc, which was cloned into pET22b vector, was co-purified with the bacteria containing the above pRSFDuet vector (His–IcmS and IcmW), according to the same protocol as the IcmS–IcmW complex. For coexpression of the IcmS–IcmW–DotLc–LvgA complex, LvgA was cloned into a modified pET15b vector to produce N-terminal His–MBP tagged LvgA with a PreScission protease digestion site between them. The E. coli BL21 (DE3) strain transformed with this vector and the above mentioned pRSFDuet vector expressed the three proteins. DotLc in the above mentioned vector was expressed from the E. coli BL21 (DE3) strain. Equal volumes of E. coli cultures were co-sonicated and cleared lysate was subjected to Ni-affinity column chromatography. After elution from the column, the protein was treated with PreScission protease, and the complex was purified through ion exchange chromatography and gel filtration chromatography.
Crystallization, data collection and structure determination
The SdeA(231–1190) was concentrated to 20 mg ml−1 in 10 mM Tris-HCl pH 8.0, 200 mM NaCl and 5 mM DTT. Crystals were grown using the hanging-drop vapour diffusion method. Crystals of SdeA were grown at 18 °C by mixing an equal volume of the protein (20 mg ml−1) with reservoir solution containing 50% Tacsimate pH 7.0, 0.1 M Tris pH 8.8, 6% sorbitol. The crystals appeared overnight and grew to full size in about 4–5 days. The crystals were cryoprotected in reservoir solution containing 10% glycerol before transfer to liquid nitrogen. Se–Met-labelled protein was crystallized in the same buffer, and the crystals diffracted better than the native crystals. After hundreds of crystal diffraction tests at Micro/Max-007HF from Rigaku and beamlines BL17U1 and BL19U1 of the Shanghai Synchrotron Radiation Facility (SSRF)32, the crystal of the Se–Met-labelled protein suitable for structure determination was finally obtained. Purified C-terminal His-tagged SdeA(231–1190) was mixed with purified Ub in molar ratios of 1:4, 1:6 and 1:8, in which the final concentration of SdeA(231–1190) was 24 mg ml−1. The crystals of the SdeA–Ub complex were also grown at 18 °C by mixing an equal volume of the protein mixture with a reservoir solution containing 0.1 M sodium malonate pH 8.0, 0.1 M Tris pH 8.0, and 30% w/v polyethylene glycol 1,000. The crystals appeared after 2 days and grew to full size in about 4–5 days. These crystals were also used in the NADH-soaking experiment.
All the data were collected at SSRF beamline BL17U1 and BL19U1, integrated and scaled using the HKL2000 package33. Further processing was carried out using programs from the CCP4 suite34. SHELXD35 was used to locate the positions of selenium sites in SdeA. The identified anomalous scattering sites were input into PHASER36 for single-wavelength anomalous dispersion (SAD) phasing. The real-space constraints were applied to the electron density map in DM37. As the resolution of the data was low, the final model rebuilding of SdeA was performed manually using Coot38 and the apo SdeA structure was refined with PHENIX39 against the SAD data using non-crystallographic symmetry and stereochemistry information as restraints. The structures of the SdeA(231–1190)–Ub and SdeA(231–1190)–Ub–NADH complexes were solved by molecular replacement with the structure of apo- SdeA(231–1190) and Ub (PDB: 1UBQ) as templates. Final Ramachandran statistics: 92.6% favoured, 6.0% allowed and 1.4% outliers for apo- SdeA(231–1190) structure; 93.8% favoured, 4.9% allowed and 1.3% outliers for SdeA(231–1190)–Ub structure; 93.2% favoured, 5.5% allowed and 1.3% outliers for SdeA(231–1190)–Ub–NADH structure. Structural illustrations were generated using PyMOL (v.1.8.0.0, https://pymol.org/). Data collection and structure refinement statistics are summarized in Extended Data Table 1.
MST assay
The NAD+ affinity of the purified wild-type SdeA(231–1190) and its mutants was measured using the Monolith NT.115 (Nanotemper Technologies). All the proteins used were desalted to MST buffer (10 mM HEPES pH 7.5, 150 mM NaCl) before the experiment. The SdeA proteins were fluorescently labelled according to the manufacturer’s procedure and the protein concentration was adjusted to 10 μM. Then fluorescent dye NT-647-NHS was added, mixed and incubated for 30 min at 25 °C in the dark. For each assay, the labelled protein (about 0.1 μM) was mixed with the same volume of unlabelled NAD+ of 16 different serial concentrations at room temperature. The samples were then loaded into premium capillaries (NanoTemper Technologies) and measured at 25 °C by using 20% LED power and medium MST power. Each assay was repeated three times. Data analyses were performed using MO.Affinity Analysis v.2.2.4 software. With a confidence of 68%, the Kd value is within the given range.
Structural analysis by SAXS
SAXS measurements were carried out at room temperature at the beamline 12 ID-B of the Advanced Photon Source, Argonne National Laboratory and the beamline BL19U2 of the National Center for Protein Science Shanghai and Shanghai Synchrotron Radiation Facility. The scattered X-ray photons were recorded with a PILATUS 1M detector (Dectris) at 12 ID-B and a PILATUS 100k detector (Dectris) at BL19U2. The setups were adjusted to achieve scattering q values of 0.005 < q < 0.89 Å−1 (12ID-B) or 0.009 < q < 0.415 Å−1 (BL19U2), in which q = (4π/λ)sinθ, and 2θ is the scattering angle. Thirty 2D images were recorded for each buffer or sample solution using a flow cell, with an exposure time of 0.5–2 s to minimize radiation damage and obtain good signal-to-noise ratio. No radiation damage was observed as confirmed by the absence of systematic signal changes in sequentially collected X-ray scattering images. The 2D images were reduced to 1D scattering profiles using Matlab (12ID-B) or BioXTAS Raw (BL19U2). Scattering profiles of the proteins were calculated by subtracting the background buffer contribution from the sample-buffer profile using the program PRIMUS40 according to standard procedures41. Concentration series measurements (fourfold and twofold dilution and stock solution) for the same sample were carried out to remove the scattering contribution owing to inter-particle interactions and to extrapolate the data to infinite dilution. The forward scattering intensity I(0) and the radius of gyration (Rg) were calculated from the data of infinite dilution at low q values in the range of qRg < 1.3, using the Guinier approximation: lnI(q) ≈ ln(I(0))−Rg2q2/3. These parameters were also estimated from the scattering profile with a broader q range of 0.006–0.30 Å−1 using the indirect Fourier transform method implemented in the program GNOM42, along with the pair distance distribution function (PDDF), P(r) and the maximum dimension of the protein, Dmax. The parameter Dmax (the upper end of the distance r), was chosen so that the resulting PDDF has a short, near-zero-value tail to avoid underestimation of the molecular dimension and consequent distortion in low-resolution structural reconstruction. The Porod volume of solutes (Vporod), the volume-of-correlation (Vc), were calculated using the programs PRIMUS and Scatter, respectively. The molecular masses of solutes were calculated on a relative scale using the Rg/Vc power law as previously described43, as well as from AUTOPOROD44, independently of protein concentration and with minimal user bias. The theoretical scattering intensity of the atomic structure model was calculated and fitted to the experimental scattering intensity using CRYSOL45. Low-resolution ab initio shape reconstructions were performed with the program DAMMIN, which generates models represented by an ensemble of densely packed beads46, using scattering data within the q range of 0.006–0.30 Å−1. Thirty-two independent runs for both programs were performed, and the resulting models were subjected to averaging by DAMAVER47 and were superimposed by SUPCOMB48 on the basis of the normalized spatial discrepancy criteria and were filtered using DAMFILT to generate the final model.
Analytical ultracentrifugation
Sedimentation velocity experiments were performed at 20 °C with a Beckman XL-I analytical ultracentrifuge (Beckman Coulter) equipped with a four-cell An-60 Ti rotor. Samples were clarified by centrifugation at 12,000 r.p.m. for 10 min in a tabletop centrifuge before the experiment. Reaction buffer containing 10 mM Tris, pH 8.0, 200 mM NaCl and 5 mM DTT was used as the reference solution, and ∼400 μl of SdeA(231–1190) peak fractions (OD280 nm = 0.7) was loaded in two-channel centerpieces fitted with sapphire windows in a four-hole rotor. Absorbance scans were taken at 280 nm versus radial location during centrifugation at 30,000 r.p.m. The differential sedimentation coefficients, c(s), frictional coefficients and molecular mass were calculated using SEDFIT software.
Molecular dynamics simulation
Molecular dynamics simulations were based on the crystal structure of the SdeA–Ub–NADH complex described in this manuscript. The simulations were carried out under two conditions: (1) the NAD+-bound complex structure (the SdeA mART–Ub–NADH structure with the NADH displaced by NAD+) and (2) the intermediate-bound complex (the SdeA mART–Ub–NADH structure with the nicotinamide group of the ligand removed). Hydrogen atoms were added with the optimal hydrogen-bonding networks and side-chain protonation states determined at pH 7.0 by PROPKA49,50. The protein-chain termini of SdeA were capped with neutral acetyl and methylamide groups. Each system was solvated in a cubic water box with a 10 Å buffer and neutralizing counter ions were added. To mimic experimental assay conditions, a 0.15 M NaCl salt bath was introduced. We used the OPLS-AA 2005 force field51 parameter set for the protein and ligand, and TIP3P model for water. The parameters for ligands (NAD+ and intermediate) were generated from the LigParGen52 web server, which applied BOSS software53 to assign the bonded and van der Waals parameters by analogy to the existing atom types. The charges were calculated and assigned by a semi-empirical AM154 calculation using CM1A charge model55,56. Simulations were performed with the Desmond software package57. Prepared systems were first minimized using 5,000 steps of a steepest descent algorithm, then equilibrated as follows: the system was heated from 0 to 310 K in the isothermal–isobaric (NpT) ensemble over 100 ps with harmonic restraints of 10.0 kcal∙mol−1∙Å−2 on heavy atoms of protein and ligand, and initial velocities sampled from the Boltzmann distribution. Further equilibration was performed at 310 K with harmonic restraints on the protein and ligand starting at 10.0 kcal∙mol−1∙Å−2 and reduced by 1.0 kcal∙mol−1∙Å−2 in a stepwise fashion every 2 ns, for a total of 20 ns of additional restrained equilibration. Production runs were then made for 200 ns duration in the NpT ensemble. The M-SHAKE algorithm58 was applied to constrain all bonds involving hydrogen atoms with a time step of 2 fs. The short-range electrostatic and Lennard–Jones interactions were cut off at 9 Å. Long-range electrostatic interactions were computed by the particle mesh Ewald method59.
Yeast toxicity assay
Yeast strain W303 was used for all the experiments. Yeast was grown at 30 °C in YPD for transformation or appropriate selective medium lacking uracil and containing either 2% glucose or galactose as a carbon source. For expression in yeasts, genes were cloned into pYES2 vector containing the galactose-inducible promoter. The integrity of all constructs was verified by sequencing analysis. For each construct, about 1 μg of plasmid DNA was used to transform yeast cells using the standard lithium acetate method. For yeast toxicity experiments, W303 strain cells carrying the defined plasmids were grown overnight in synthetic media lacking uracil and containing 2% glucose. The cells were collected, washed once with sterile water and resuspended in sterile water to an OD600 nm of 1.0, 0.1 or 0.01. Then 5-μl aliquots of this suspension were spotted onto solid synthetic defined medium lacking uracil and containing either 2% glucose or galactose for protein expression. Plates were grown at 30 °C and images were acquired after 2 days of growth.
Preparation of the ADPR-Ub
For producing ADPR-Ub, 0.1 μM ∆NC SdeAH277A was used in the reaction mixture after tests of enzyme concentrations for the best conversion. In brief, 0.1 μM ∆NC SdeAH277A was incubated with 0.4 mM NAD+ and 35 μM Ub at 37 °C for 3 h, after which the reaction mixture was concentrated and loaded onto the Superdex-75 column (GE Helthcare). The peak fractions of Ub were pooled and subjected to mass spectrometric analysis to verify that all the Ubs were in the ADPR-Ub form. ADPR-Ub could be prepared to 100% purity by this method. They were then stored at −80 °C until use.
In vitro ubiquitin-modification and RAB33B-ubiquitination assays
For auto-ubiquitination of SdeA(231–1190) experiments, 6 μM of purified GST-tagged ubiquitin was incubated with 0.9 μM SdeA(231–1190) and RAB33B at 37 °C for 1 h in the presence or absence of 0.1 mM NAD+ in a buffer containing 50 mM Tris pH 7.5, 1 mM DTT. After the reaction, the samples were analysed using SDS–PAGE and Coomassie staining. For the RAB33B-ubiquitination experiments, to test whether SdeA(231–1190) was functioning normally, 0.6 μM SdeA, 0.9 μM wild-type SdeA(231–1190), SdeA(231–1190)E860A/E862A or SdeA(231–1190)H277A were incubated with 1 mM NAD+ and 35 μM Ub in the presence or absence of 9.5 μM His–RAB33B at 37 °C for 5 min. The samples were then analysed using Tricine gel, Coomassie staining and immunoblotting with anti-His and anti-Ub antibodies. For the ubiquitination experiments of wild-type ∆NC SdeA and the mutants, 1 μM ∆NC SdeA (wild type and mutants) was incubated with 0.1 mM NAD+, 35 μM Ub and 7.4 μM His–RAB33B at 37 °C for 30 min. For the NADH-inhibition experiment, 1, 2 or 5 mM NADH was added to the reaction mixture, in which the other components were the same concentrations as above. The samples were then analysed using SDS–PAGE. For phospho-specific staining of PR-Ub, Pro-Q Diamond stain was used according to the manufacturer’s instructions. For immunoblotting analysis, primary antibodies were used: anti-His (1:5,000, Transgene, HT501) and anti-Ub (1:500, Santa Cruz Biotechnology, sc-8017). For the time kinetics of the ubiquitination reactions, 1.09 μM ∆NC SdeA (wild type and mutants) or combinations of its fragments were incubated with 1 mM NAD+, 35 μM Ub, 9.5 μM His–RAB33B at 37 °C for indicated times. The samples were stained with Coomassie and Pro-Q diamond phosphoprotein stain.
Top-down LC–MS analysis of modified Ub and Ub-like proteins
Wild-type Ub was purchased from Boston Biochem (U-100H), and the untagged Ub mutants were purified from E. coli cells. The ubiquitination reactions were performed in a 100-μl system, in which 35 μM Ub, Ub mutants or Ub-like proteins was incubated with specific SdeA fragments or mutants for 2 h in a buffer containing 50 mM NaCl, 50 mM Tris pH 7.5 and 2 mM DTT. The reaction mixtures were then run through 30-kDa molecular mass cut-off filters to obtain the modified Ub or other Ub-like proteins below the filter. The proteins were then subjected to LC–MS analysis. A linear ion-trap mass spectrometer (LTQ Velos Pro, Thermo Scientific) was used for total molecular mass analyses. Liquid chromatography separation was carried out on an EASY-nLC 1200 system (Thermo Scientific). The capillary column (75 μm × 150 mm) with a laser-pulled electrospray tip was home-packed with 4-μm, 100 Å Magic C4AQ silica-based particles (Michrom BioResources). The mobile phase consisted of solvent A (97% H2O, 3% ACN, and 0.1% FA) and solvent B (20% H2O, 80% ACN and 0.1% FA). The following gradient was used: solvent B was started at 20% for 3 min and then raised to 50% in 20 min; subsequently, solvent B was rapidly increased to 70% in 2 min and maintained for 20 min before 100% solvent A was used for column equilibration. Eluted peptides from the capillary column were electrosprayed directly onto the mass spectrometer for mass-spectrometry analyses. One full mass-spectrometry scan (m/z 600–1,500) was acquired.
In vitro GST pull-down assay
To detect whether SdeA could bind IcmS alone or its complexes, GST-fused full-length SdeA protein was preloaded on glutathione resins and then incubated with IcmS, IcmS–IcmW, IcmS–IcmW–DotLc or IcmS–IcmW–DotLc–LvgA at 18 °C for 1 h. The samples bound on glutathione resins were washed three times with the washing buffer (50 mM Tris, pH 7.5, 50 mM NaCl) and then analysed by SDS–PAGE and Coomassie blue staining. To detect the region of SdeA responsible for IcmS binding, GST-fused full-length IcmS protein was preloaded on glutathione resins and then incubated with different fragments of SdeA protein at 18 °C for 1 h. The samples were treated as stated above.
Gel-filtration binding assay
The SdeA(1191–1350) and IcmS–IcmW–DotLc complex purified as described above were subjected to gel-filtration analysis (Superdex 200, GE Healthcare). They were mixed at a molar ratio of about 1:1 and incubated at 18 °C for 4 h before gel-filtration analysis in buffer containing 10 mM Tris pH 8.0, 100 mM NaCl. Samples from relevant fractions were applied to SDS–PAGE and visualized by Coomassie blue staining.
Statistics and reproducibility
No statistical methods were used to predetermine sample size. All of the in vitro assays presented in this work were repeated at least three times with similar results. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Coordinates and structure factors for the complexes have been deposited in the Protein Data Bank (PDB) under accessions: 5YIM, (SdeA(231–1190); 5YIK, SdeA(231–1190)–Ub; and 5YIJ, SdeA(231–1190)–Ub–NADH. Uncropped versions of all gels are displayed in Supplementary Fig. 1. All other data are available from the corresponding author upon reasonable request.
References
Komander, D. & Rape, M. The ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012).
Yau, R. & Rape, M. The increasing complexity of the ubiquitin code. Nat. Cell Biol. 18, 579–586 (2016).
Hershko, A., Ciechanover, A. & Varshavsky, A. The ubiquitin system. Nat. Med. 6, 1073–1081 (2000).
Hubber, A., Kubori, T. & Nagai, H. Modulation of the ubiquitination machinery by Legionella. Curr. Top. Microbiol. Immunol. 376, 227–247 (2013).
Jeong, K. C., Sexton, J. A. & Vogel, J. P. Spatiotemporal regulation of a Legionella pneumophila T4SS substrate by the metaeffector SidJ. PLoS Pathog. 11, e1004695 (2015).
Bardill, J. P., Miller, J. L. & Vogel, J. P. IcmS-dependent translocation of SdeA into macrophages by the Legionella pneumophila type IV secretion system. Mol. Microbiol. 56, 90–103 (2005).
Horwitz, M. A. Formation of a novel phagosome by the Legionnaires’ disease bacterium (Legionella pneumophila) in human monocytes. J. Exp. Med. 158, 1319–1331 (1983).
Swanson, M. S. & Isberg, R. R. Association of Legionella pneumophila with the macrophage endoplasmic reticulum. Infect. Immun. 63, 3609–3620 (1995).
Kagan, J. C. & Roy, C. R. Legionella phagosomes intercept vesicular traffic from endoplasmic reticulum exit sites. Nat. Cell Biol. 4, 945–954 (2002).
Luo, Z. Q. & Isberg, R. R. Multiple substrates of the Legionella pneumophila Dot/Icm system identified by interbacterial protein transfer. Proc. Natl Acad. Sci. USA 101, 841–846 (2004).
Zhu, W. et al. Comprehensive identification of protein substrates of the Dot/Icm type IV transporter of Legionella pneumophila. PLoS ONE 6, e17638 (2011).
Lifshitz, Z. et al. Computational modeling and experimental validation of the Legionella and Coxiella virulence-related type-IVB secretion signal. Proc. Natl Acad. Sci. USA 110, E707–E715 (2013).
Qiu, J. & Luo, Z. Q. Legionella and Coxiella effectors: strength in diversity and activity. Nat. Rev. Microbiol. 15, 591–605 (2017).
Qiu, J. et al. Ubiquitination independent of E1 and E2 enzymes by bacterial effectors. Nature 533, 120–124 (2016).
Kotewicz, K. M. et al. A single Legionella effector catalyzes a multistep ubiquitination pathway to rearrange tubular endoplasmic reticulum for replication. Cell Host Microbe 21, 169–181 (2017).
Bhogaraju, S. et al. Phosphoribosylation of ubiquitin promotes serine ubiquitination and impairs conventional ubiquitination. Cell 167, 1636–1649.e13 (2016).
Sheedlo, M. J. et al. Structural basis of substrate recognition by a bacterial deubiquitinase important for dynamics of phagosome ubiquitination. Proc. Natl Acad. Sci. USA 112, 15090–15095 (2015).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Rao, S. T. & Rossmann, M. G. Comparison of super-secondary structures in proteins. J. Mol. Biol. 76, 241–256 (1973).
Han, S., Arvai, A. S., Clancy, S. B. & Tainer, J. A. Crystal structure and novel recognition motif of Rho ADP-ribosylating C3 exoenzyme from Clostridium botulinum: structural insights for recognition specificity and catalysis. J. Mol. Biol. 305, 95–107 (2001).
Jeong, B. R. et al. Structure function analysis of an ADP-ribosyltransferase type III effector and its RNA-binding target in plant immunity. J. Biol. Chem. 286, 43272–43281 (2011).
Dikic, I., Wakatsuki, S. & Walters, K. J. Ubiquitin-binding domains—from structures to functions. Nat. Rev. Mol. Cell Biol. 10, 659–671 (2009).
Puvar, K. et al. Ubiquitin chains modified by the bacterial ligase SdeA are protected from deubiquitinase hydrolysis. Biochemistry 56, 4762–4766 (2017).
Sakurai, J., Nagahama, M., Oda, M., Tsuge, H. & Kobayashi, K. Clostridium perfringens iota-toxin: structure and function. Toxins (Basel) 1, 208–228 (2009).
Tsurumura, T. et al. Arginine ADP-ribosylation mechanism based on structural snapshots of iota-toxin and actin complex. Proc. Natl Acad. Sci. USA 110, 4267–4272 (2013).
Tsuge, H. et al. Structural basis of actin recognition and arginine ADP-ribosylation by Clostridium perfringens ι-toxin. Proc. Natl Acad. Sci. USA 105, 7399–7404 (2008).
Akturk, A. et al. Mechanism of phosphoribosyl-ubiquitination mediated by a single Legionella effector. Nature https://doi.org/10.1038/s41586-018-0147-6 (2018).
Kalayil, S. et al. Insights into catalysis and function of phosphoribosyl-linked serine ubiquitination. Nature https://doi.org/10.1038/s41586-018-0145-8 (2018).
Ninio, S., Zuckman-Cholon, D. M., Cambronne, E. D. & Roy, C. R. The Legionella IcmS–IcmW protein complex is important for Dot/Icm-mediated protein translocation. Mol. Microbiol. 55, 912–926 (2005).
Cambronne, E. D. & Roy, C. R. The Legionella pneumophila IcmSW complex interacts with multiple Dot/Icm effectors to facilitate type IV translocation. PLoS Pathog. 3, e188 (2007).
Kwak, M. J. et al. Architecture of the type IV coupling protein complex of Legionella pneumophila. Nat. Microbiol. 2, 17114 (2017).
Wang, Q. S. et al. The macromolecular crystallography beamline of SSRF. Nucl. Sci. Tech. 26, 010102 (2015).
Otwinowski, Z., Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Schneider, T. R. & Sheldrick, G. M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Cowtan, K. D. & Zhang, K. Y. Density modification for macromolecular phase improvement. Prog. Biophys. Mol. Biol. 72, 245–270 (1999).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002).
Konarev, P. V., Volkov, V. V., Sokolova, A. V., Koch, M. H. J. & Svergun, D. I. PRIMUS - a Windows-PC based system for small-angle scattering data analysis. J. Appl. Crystallogr. 36, 1277–1282 (2003).
Wang, J. et al. A method for helical RNA global structure determination in solution using small-angle X-ray scattering and NMR measurements. J. Mol. Biol. 393, 717–734 (2009).
Svergun, D. I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 25, 495–503 (1992).
Rambo, R. P. & Tainer, J. A. Accurate assessment of mass, models and resolution by small-angle scattering. Nature 496, 477–481 (2013).
Petoukhov, M. V., Konarev, P. V., Kikhney, A. G. & Svergun, D. I. ATSAS 2.1—towards automated and websupported small-angle scattering data analysis. J. Appl. Crystallogr. 40, s223–s228 (2007).
Svergun, D. I., Barberato, C. & Koch, M. H. J. CRYSOL—a program to evaluate X-ray solution scattering of biological macromolecules from atomic coordinates. J. Appl. Crystallogr. 28, 768–773 (1995).
Svergun, D. I. Restoring low resolution structure of biological macromolecules from solution scattering using simulated annealing. Biophys. J. 76, 2879–2886 (1999).
Volkov, V. V. & Svergun, D. I. Uniqueness of ab initio shape determination in small-angle scattering. J. Appl. Crystallogr. 36, 860–864 (2003).
Kozin, M. B. & Svergun, D. I. Automated matching of high- and low-resolution structural models. J. Appl. Crystallogr. 34, 33–41 (2001).
Bas, D. C., Rogers, D. M. & Jensen, J. H. Very fast prediction and rationalization of pK a values for protein–ligand complexes. Proteins 73, 765–783 (2008).
Li, H., Robertson, A. D. & Jensen, J. H. Very fast empirical prediction and rationalization of protein pK a values. Proteins 61, 704–721 (2005).
Banks, J. L. et al. Integrated modeling program, applied chemical theory (IMPACT). J. Comput. Chem. 26, 1752–1780 (2005).
Dodda, L. S., Cabeza de Vaca, I., Tirado-Rives, J. & Jorgensen, W. L. LigParGen web server: an automatic OPLS-AA parameter generator for organic ligands. Nucleic Acids Res. 45, W331–W336 (2017).
Jorgensen, W. L. & Tirado-Rives, J. Molecular modeling of organic and biomolecular systems using BOSS and MCPRO. J. Comput. Chem. 26, 1689–1700 (2005).
Storer, J. W., Giesen, D. J., Cramer, C. J. & Truhlar, D. G. Class IV charge models: a new semiempirical approach in quantum chemistry. J. Comput. Aided Mol. Des. 9, 87–110 (1995).
Udier-Blagović, M., Morales De Tirado, P., Pearlman, S. A. & Jorgensen, W. L. Accuracy of free energies of hydration using CM1 and CM3 atomic charges. J. Comput. Chem. 25, 1322–1332 (2004).
Dodda, L. S., Vilseck, J. Z., Tirado-Rives, J. & Jorgensen, W. L. 1.14*CM1A-LBCC: Localized Bond-Charge Corrected CM1A Charges for Condensed-Phase Simulations. J. Phys. Chem. B 121, 3864–3870 (2017).
Bowers, K. J. et al. Scalable algorithms for molecular dynamics simulations on commodity clusters. In Proc. of the 2006 ACM/IEEE Conference on Supercomputing 43–43 (2006).
Kräutler, V., Van Gunsteren, W. F. & Hünenberger, P. H. A fast SHAKE algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations. J. Comput. Chem. 22, 501–508 (2001).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089 (1993).
Petoukhov, M. V. et al. New developments in the ATSAS program package for small-angle scattering data analysis. J. Appl. Crystallogr. 45, 342–350 (2012).
Acknowledgements
We thank B. Zhou for the gift of the yeast W303 strain and the pYES2 vector; Y. Zhang for help in the yeast experiments; J. Ren, X. Zhang and R. Qiao for discussions about the mechanisms of SdeA; S. Zhang for the electron microscopy tests of the SdeA sample; C. Yan for help with the data collection process; the Tsinghua University Branch of China National Center for Protein Sciences Beijing and Y. Xue for providing facility support for NMR analysis of the protein samples; the Protein Chemistry Facility at the Center for Biomedical Analysis of Tsinghua University and W. Zhang for sample analysis; the staff at beamline BL17U1 and BL19U1 of the Shanghai Synchrotron Radiation Facility for their assistance with data collection and X. Zuo at the Advanced Photon Source (APS), Argonne National Laboratory (ANL) and the staff of BL19U2 beamline at the National Center for Protein Science Shanghai and Shanghai Synchrotron Radiation Facility for assistance during data collection. Use of the scattering beamline 12-ID-B resource at APS, ANL is allocated under the GUP-52757 to X.F. This work was supported by the National Key Research and Development Program of China (2017YFA0506500), the National Natural Science Foundation of China (31670766, 21532004, 21475005, and 21622501) and the Fundamental Research Funds for the Central Universities (buctylkxj03).
Reviewer information
Nature thanks K. Gehring and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Y.F. designed and supervised the project. Y.D., Y.M., Y.H. and W.W. purified the proteins, grew and optimized the crystals, and tested the diffractions of hundreds of crystals. Y.M., Y.X., Y.D., Z.L., Y.H., H.W. and N.X. performed the in vitro activity analysis, MST analysis, yeast toxicity assay and GST pull-down assays. Y.Zho. conducted the molecular dynamics simulations, and C.-Q.X. and J.L. conducted calculations to analyse the results of the molecular dynamics simulations. Y.Zha. and X.F. performed the SAXS analysis of different constructs of SdeA. M.W., H.D. and X.L. performed the mass spectrometric analysis. M.P., T.T., and L.L. contributed to experiment design and helped to supervise the project. J.W., Y.F., M.Y. and S.F. collected and analysed the crystallographic data and solved the crystal structure. Y.F. analysed the data and wrote the paper with the help of all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Sequence alignment of SidE family members, spanning the regions corresponding to the SdeA fragment used in crystallization.
Residues with 100% homology, over 75% homology and over 50% homology are shaded in dark blue, pink and light blue, respectively. Secondary structural elements of SdeA are shown above the sequences. The residue ranges of the PDE, mART, pCTD and the α-helical lobe of the mART domain are marked with brackets or lines.
Extended Data Fig. 2 SdeA(231–1190) is an active monomer in solution.
a, SdeA, wild-type SdeA(231–1190), SdeA(231–1190)E860A/E862A or SdeA(231–1190)H277A were incubated with NAD+ and Ub in the presence or absence of His–RAB33B. Ubiquitinated His–RAB33B were analysed using tricine gels, Coomassie staining and immunoblotting with anti-His and anti-Ub antibodies. b, SdeA(231–1190) and RAB33B(15–202) were incubated with GST–Ub and NAD+, and self-ubiquitinated SdeA was detected by Coomassie staining, immunoblotting with anti-Ub antibodies, and Pro-Q diamond phosphoprotein staining. c, SdeA(231–1190), NAD+ and RAB33B were incubated with 0, 1, 2 or 5 mM NADH. The ubiquitination reactions were analysed using tricine gel and Coomassie staining. d, Analytical ultracentrifugation results showed that SdeA(231–1190) is a monomer. Analytical ultracentrifugation analysis yielded a sedimentation coefficient of 5.13 S, and a molecular mass of approximately 106 kDa. The buffer is 10 mM Tris pH 8.0, 200 mM NaCl and 5 mM DTT. e, Gel filtration profile of the SdeA(231–1190) protein and the molecular markers on Superdex-75 column (GE Healthcare) are shown. The sizes of the molecular markers are marked on top of the peaks. The samples of SdeA(231–1190) collected from the Superdex-75 column were run on SDS–PAGE gels and detected by Coomassie staining. a–e, Similar results were obtained in three independent experiments. a–c, e, Uncropped blots and gel images are shown in Supplementary Fig. 1. f, Two views of the superimposition of the structures of the two molecules in the asymmetric unit, coloured in different colours. g, Structure of the CTD region in the crystallized protein can be divided into two parts (left and right). The α helices are numbered according to their orders in the residue region from 908 to 1190. h, Topological diagram of the CTD region shown in g. The N and C termini of the pCTD domain are labelled.
Extended Data Fig. 3 Interactions between SdeA mART and SdeA PDE are essential for the activity of SdeA mART.
a, Overview of the interactions between SdeA mART and SdeA PDE. SdeA is coloured as in Fig. 1b. The major interaction region between the two domains is outlined. b, An expanded view of the region outlined in a. Interaction residues are shown in stick representation and the red dashed lines represent polar interactions. The plug loop in SdeA mART is indicated. c, The interaction between SdeA mART and SdeA PDE. SdeA mART and SdeA PDE are shown in cartoon and surface electrostatic models, respectively. d, A view of the interaction from c rotated by 180 degrees. In this view, SdeA mART and SdeA PDE are shown as surface electrostatic and cartoon models, respectively. e, Testing the ability of SdeA PDE to process ADPR-Ub into PR-Ub. SdeA(231–588), wild-type ∆NC SdeA or the H277A mutant were incubated with ADPR-Ub and RAB33B for 30 min. The samples were stained with Coomassie and Pro-Q diamond phosphoprotein stain. f, Testing the importance of domain interaction for the activity of SdeA mART. Various SdeA segments, and mixtures of SdeA(231–588) and SdeA(597–935) or SdeA(193–935)H277A, were incubated with RAB33B, NAD+ and Ub for 30 min. The samples were analysed using Coomassie staining, immunoblotting with anti-Ub antibodies and Pro-Q diamond phosphoprotein staining. e, f, Similar results were obtained in three independent experiments. Uncropped blots and gel images are shown in Supplementary Fig. 1.
Extended Data Fig. 4 SdeA mART exhibits novel conformations of ARTT and PN loops.
a, Superimposition of SdeA mART (green) and HopU1 (pink) from Pseudomonas syringae (PDB: 3U0J). The ARTT loop is indicated. The r.m.s.d. value is indicated beside the PDB code (panels b and c are arranged in the same way). b, Superimposition of SdeA mART (green) and ADP-ribosyltransferase Vis (blue) (PDB: 4XZK). c, Superimposition of SdeA mART (green) and XopAI from Xanthomonas axonopodis pv. citri (cyan) (PDB: 4ELN). d, Superimposition of SdeA mART structure (green) and the three other structures from a–c. e, f, Mass spectra of the samples in Fig. 2f. The sample name and their molecular masses are indicated in the figures. g, h, Different fragments and different combinations of SdeA proteins were incubated with Ub, RAB33B and NAD+ at 37 °C for the indicated amounts of time. The samples were analysed using Coomassie staining and Pro-Q diamond phosphoprotein staining. i, Testing the ubiquitination ability of the catalytic core. 0.09 or 0.9 μM ∆NC SdeA or SdeA(193–935) was incubated with or without Ub and NAD+ for the indicated amounts of time. The samples were analysed using Coomassie staining, immunoblotting with anti-Ub antibodies and Pro-Q diamond phosphoprotein staining. e–i, Similar results were obtained in three independent experiments. g–i, Uncropped blots and gel images are shown in Supplementary Fig. 1.
Extended Data Fig. 5 SdeA(231–1190) binds three Ub molecules.
a, Overall structure of the SdeA(231–1190)–Ub complex. SdeA is coloured as in Fig. 1b. The three Ub molecules are coloured in magenta and labelled as Ub1–3 according to the order of their binding region in SdeA(231–1190). Q935 and S998, which are two common C termini of the clones used in this study, are shown as spheres. b, Ub binding causes prominent structural changes of SdeA. The SdeA–Ub complex structure is shown as in a, and the apo-SdeA structure is coloured in pink. The N-terminal region of SdeA pCTD which undergoes pronounced conformational changes is outlined with a circle. c, d, Expanded views of the two Ub binding sites in SdeA pCTD. The proteins are coloured as in a. Red dashed lines indicate polar interactions. e–h, Structural alignments of the Ub molecule (magenta) in the SdeA mART–Ub complex with the proximal (yellow) and distal (orange) Ubs of the K11- (e), K48- (f), K63- (g) and M1-linked (h) diubiquitins. The two R42 residues in each of the four diUbs are shown in stick representation.
Extended Data Fig. 6 Specific recognition of Ub by SdeA mART.
a, The interaction between SdeA mART and Ub. SdeA mART is shown as a surface electrostatic potential model and Ub is in magenta cartoon representation. The R42, R72 and R74 residues of Ub are shown in stick representation. b, UbR72 and UbR74 are bound in the negatively charged groove of SdeA mART. The front part of SdeA mART is cut away to reveal the inner surface. c, Superimposition of SUMO1 (PDB: 1WM3), NEDD8 (PDB: 1NDD) and Ub in the SdeA mART-Ub complex. The conserved Arg residues in Ub, SUMO1 and NEDD8 are shown in stick representation, out of which UbR42, UbR72, and UbR74 are marked. The polar interactions with UbR72 and UbR74 are shown as red dashed lines. d, The purified SUMO and NEDD8 proteins were incubated with SdeA(193–935)H277A and NAD+ under the conditions stated in the ‘Top-down LC–MS analysis of modified Ub and Ub-like proteins’ section of the Methods. Mass spectra of the samples are also shown. The sample names and their molecular masses are indicated in the figures. Similar results were obtained in three independent experiments.
Extended Data Fig. 7 Molecular dynamics simulations indicate the movements of the side chains of UbR42 and UbR72.
a–d, Wild-type ∆NC SdeA and indicated mutants were incubated with Ub, RAB33B and NAD+ at 37 °C for the indicated amounts of time. The samples were analysed using Coomassie staining and Pro-Q diamond phosphoprotein staining. Similar results were obtained in three independent experiments. Uncropped blots and gel images are shown in Supplementary Fig. 1. e, The structure of the SdeA mART–Ub–NADH complex. SdeA mART is shown as an electrostatic surface potential model. White, blue and red indicate neutral, positive and negative surfaces, respectively. Shown in green mesh is the 2Fo−Fc electron density map contoured at 1σ around the NADH molecule. f, Galactose-inducible pYES2 plasmids containing wild-type ∆NC SdeA or the mutants were transformed into yeast W303 strain. Five microlitres of cells in three tenfold serial dilutions were spotted on both glucose- and galactose-containing plates lacking uracil for two days before image acquisition. g, Purified ADPR-Ub proteins were treated with or without wild-type ∆NC SdeA. The samples were analysed using Coomassie staining and Pro-Q diamond phosphoprotein staining. h, Purified ADPR-Ub protein was subjected to top-down LC–MS analysis. The results indicated 100% ADPR-Ub. i, Wild-type ∆NC SdeA or other mutants were incubated with RAB33B and the prepared ADPR-Ub verified in g and h. The samples were analysed using SDS–PAGE, with Coomassie staining and Pro-Q diamond phosphoprotein staining. f–i, Similar results were obtained in three independent experiments. g, i, Uncropped blots and gel images are shown in Supplementary Fig. 1. j, k, The time series for the r.m.s.d. of the non-hydrogen atoms of the protein–ligand complex (j) and the ligand (k) in the SdeA mART–Ub–intermediate and SdeA mART–Ub–NAD+ systems during molecular dynamics simulations. These two plots indicate that both systems have reached equilibrium during the 200-ns simulations. l, m, The time series for the shortest distance between the NH1/2 atom of UbR72 and C1D of the ligand (l) and the distance between the NH1/2 atom of UbR42 and C1D of the ligand (m) in the two systems during molecular dynamics simulations.
Extended Data Fig. 8 The overall shape of SdeA and the function of SdeA CTD.
a, Superimposition of various Ub structures (PDB codes: 5M93 (orange), 1UBQ (pink), 5CRA (chain C, cyan), 3ZLZ (chain B, yellow) and 4BOZ (chain B, grey)) onto the SdeA mART–Ub–NADH structure with R42 residues of all the Ub molecules shown in stick representation. SdeA mART and Ub from the SdeA mART–Ub–NADH complex are shown in green and magenta, respectively. b, In vitro GST pull-down assays to detect the interactions of SdeA with IcmS or its complexes. GST-fused SdeA protein was incubated with IcmS, the IcmS–IcmW complex, the IcmS–IcmW–DotLc (residues 656–783 of DotL) ternary complex or the IcmS–IcmW–DotLc–LvgA quaternary complex. The protein samples bound to glutathione resins were washed three times and analysed by SDS–PAGE and Coomassie blue staining. IcmS/W represents IcmS + IcmW. The band marked with an asterisk represents the degraded GST tag. Similar results were obtained in three independent experiments. Uncropped blots and gel images are shown in Supplementary Fig. 1. c, Experimental PDDFs (pair distance distribution function) for SdeA(231–1190), SdeA(1–1499), SdeA(1–1190) and SdeA(1092–1496). d, Overlay of the experimental scattering profiles (exp) from the four samples in the SAXS analysis with the back-calculated scattering profile of the crystal structure of SdeA(231–1190) (cal). e, Fitting the crystal structure of SdeA(231–1190) into the SAXS envelope of SdeA(231–1190). Two perpendicular views are shown. f, Superimposition of the SAXS envelopes of SdeA(231–1190) (coloured as in e) and SdeA(1–1190) (light magenta) with the crystal structure of SdeA(231–1190) fitted. g, SAXS envelopes of SdeA(1092–1496). h, Superimposition of the SAXS envelopes of SdeA(1–1190) (light magenta), SdeA(1092–1496) (cyan) and SdeA(1–1499) (wheat) with the crystal structure of SdeA(231–1190) fitted.
Supplementary information
Supplementary Figure 1
This file contains the uncropped scans with size marker indications.
Rights and permissions
About this article
Cite this article
Dong, Y., Mu, Y., Xie, Y. et al. Structural basis of ubiquitin modification by the Legionella effector SdeA. Nature 557, 674–678 (2018). https://doi.org/10.1038/s41586-018-0146-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0146-7
This article is cited by
-
Legionella metaeffector MavL reverses ubiquitin ADP-ribosylation via a conserved arginine-specific macrodomain
Nature Communications (2024)
-
Molecular basis of threonine ADP-ribosylation of ubiquitin by bacterial ARTs
Nature Chemical Biology (2024)
-
Serine-ubiquitination regulates Golgi morphology and the secretory pathway upon Legionella infection
Cell Death & Differentiation (2021)
-
Structural basis for protein glutamylation by the Legionella pseudokinase SidJ
Nature Communications (2021)
-
Structural insights into the mechanism and inhibition of transglutaminase-induced ubiquitination by the Legionella effector MavC
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.