Abstract
A short, 14-amino-acid segment called SP1, located in the Gag structural protein1, has a critical role during the formation of the HIV-1 virus particle. During virus assembly, the SP1 peptide and seven preceding residues fold into a six-helix bundle, which holds together the Gag hexamer and facilitates the formation of a curved immature hexagonal lattice underneath the viral membrane2,3. Upon completion of assembly and budding, proteolytic cleavage of Gag leads to virus maturation, in which the immature lattice is broken down; the liberated CA domain of Gag then re-assembles into the mature conical capsid that encloses the viral genome and associated enzymes. Folding and proteolysis of the six-helix bundle are crucial rate-limiting steps of both Gag assembly and disassembly, and the six-helix bundle is an established target of HIV-1 inhibitors4,5. Here, using a combination of structural and functional analyses, we show that inositol hexakisphosphate (InsP6, also known as IP6) facilitates the formation of the six-helix bundle and assembly of the immature HIV-1 Gag lattice. IP6 makes ionic contacts with two rings of lysine residues at the centre of the Gag hexamer. Proteolytic cleavage then unmasks an alternative binding site, where IP6 interaction promotes the assembly of the mature capsid lattice. These studies identify IP6 as a naturally occurring small molecule that promotes both assembly and maturation of HIV-1.
Similar content being viewed by others
Main
IP6 is a highly negatively charged compound that is present in all mammalian cells at concentrations of 10–40 µM6. Inositol phosphates stimulate in vitro assembly of HIV-1 Gag into immature virus-like particles (VLPs), with previous data suggesting that IP6 interacts with both the MA and NC domains of Gag7,8,9. To understand how IP6 affects HIV-1 assembly, we used an HIV-1 Gag construct spanning the CA to NC domains and having one extra amino acid residue, Ser, preceding the normal N-terminal Pro at the start of the CA domain (s-CANC; Fig. 1a), because this should disfavour formation of the N-terminal β-hairpin that promotes mature assembly10. Longer N-terminal extensions of CANC constructs have been shown to assemble inefficiently into immature VLPs at pH 8, but into mature VLPs at pH 61,10. However, we found that s-CANC still formed mature-like particles at both pH values (Fig. 1b). Notably, the presence of IP6 induced a marked switch to the formation of spherical, immature VLPs (Fig. 1b, d). At pH 8, even a substoichiometric 1:50 molar ratio of IP6 to protein resulted in an approximately 100-fold increase in immature VLPs (Fig. 1b, c). At pH 6, the effect of IP6 was less strong, requiring at least a 1:10 ratio to induce immature assembly (Fig. 1b). We conclude that IP6 imposes an in vitro immature assembly phenotype, even under conditions that favour the mature lattice (pH 6).
a, Map of the HIV-1 Gag protein, indicating the MA, CA, NC and p6 domains, and spacer peptides SP1 and SP2. Gag-derived constructs used in this study are shown underneath. Blue bar, major homology region; purple bar, SP1 helix; NTD and CTD, N-terminal and C-terminal domains of CA; R18, K290 and K359, locations of mutations; N372, C-terminal residue of the s-CASP1 and CACTDSP1 constructs. b, Negative-stain electron microscopy images of mature and immature VLPs formed by s-CANC (50 μM) at pH 6 and pH 8 in the absence or presence of the indicated molar ratios of IP6 (0–10 μM). Scale bars, 100 nm. c, Number of VLPs per 55 µm2 without (–) and with (+) 10 µM IP6 at pH 6 and pH 8; n shown below and mean above box plots. The experiment was repeated three times with similar results. d, Diameters of immature and mature VLPs; n shown below and mean above box plots. e, Representative images of s-CANC VLPs assembled at pH 8 in the absence and presence of IP3, IP4, IP5 and IP6. Scale bars, 100 nm. The experiment was repeated twice with similar results. f, Number of VLPs per 55 µm2 without and with 10 µM IP3–IP6; n = 5, mean shown above box plots. g, Parallel transfections of 293FT wild-type (WT) and IPPK knockout (KO) cells were performed with a VSV-G-pseudotyped HIV-1 provirus containing GFP, and infectivity was measured on WT 293T cells. Graphs show mean ± s.d. of four independent experiments; dots show individual data points. Right panels show sequences of total PCR products of the guide RNA target sites from WT and KO cells; guide RNA sequence is underlined in red. c, d, f, Centre lines show medians; box limits indicate 25th and 75th percentiles as determined by R software24; whiskers extend to minimum and maximum values.
Other inositol derivatives also promoted s-CANC assembly, but to a lesser extent, in the order IP3 < IP4 < IP5 < IP6 (Fig. 1e, f), with efficacy correlating with the number of phosphate groups. Other negatively charged compounds did not promote or only marginally promoted assembly (Extended Data Fig. 1). Overall, these results indicate that charge neutralization is a fundamental aspect of IP6-mediated HIV-1 Gag assembly, and that the details of coordination geometry and/or local stereochemistry are also important.
To address the biological importance of IP6 in HIV-1 replication, we generated a knockout cell line in which the gene encoding inositol pentakisphosphate 2-kinase (IPPK), the enzyme responsible for the final step in IP6 synthesis, was ablated (Fig. 1g). Infectious HIV-1 particle production from these knockout cells was reduced by between 10- and 20-fold (Fig. 1g). We interpret this result as implying that IP6 has a critical role in assembly of immature and/or mature HIV-1.
As the s-CANC construct lacks the MA domain, the effect of IP6 cannot depend on this domain, as previously suggested7,8,9. The NC domain also cannot be essential, because IP6 still promoted assembly in the absence of nucleic acid (Extended Data Fig. 2a, d). Furthermore, IP6 also promoted the formation of abundant immature VLPs from the smaller protein s-CASP1, which lacks the NC domain altogether (Fig. 2a and Extended Data Fig. 2b, d). However, deletion of the SP1 domain abrogated the effect of IP6, as IP6 failed to induce assembly of s-CA into immature VLPs (Extended Data Fig. 2c).
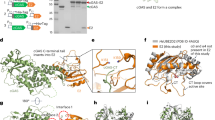
a, IP6-induced assembly of s-CASP1 into immature VLPs. The experiment was repeated four times with similar results. b, IP6-induced assembly of CACTDSP1 into flat micro-crystals. The experiment was repeated six times with similar results. c, Two-dimensional cryo-EM projection map of a micro-crystal. Images of multiple crystals were collected during two rounds of data collection from separate assembly reactions and all crystals had similar unit cells. Two individual crystals had single layer regions and could be further processed. These crystals generated similar maps. d, e, Top view (d) and side view (e) of the CACTDSP1 hexamer X-ray crystal structure showing the protein in grey ribbons and unbiased mFo–DFc difference density in blue mesh, contoured at 2σ. f, Top and side views of IP6 in its myo configuration, docked into the difference density as a rigid body in one of six rotationally equivalent orientations. All six binding modes are shown in Extended Data Fig. 4a. g, Side view of the two rings of Lys290 (green) and Lys359 (cyan) with bound IP6 in the middle. Densities were omitted for clarity, and are shown in Extended Data Fig. 4b.
Both the CA domain of Gag and the mature CA protein are composed of two separately folded sub-domains, CANTD and CACTD. To further define the site of action of IP6, we removed the N-terminal CANTD sub-domain to create CACTDSP1, which makes up the minimal Gag hexagonal lattice3,11. In the presence but not the absence of IP6, and at physiological pH and ionic strength, CACTDSP1 formed flat hexagonal crystals, as shown by negative-stain electron microscopy (Fig. 2b). These crystals had the characteristic immature lattice spacing (Fig. 2c). That s-CASP1 formed a spherical lattice while CACTDSP1 formed a flat lattice suggests that CANTD provides the contacts necessary for enforcing lattice curvature.
We next determined the X-ray crystal structure of CACTDSP1 crystallized in the presence of IP6 (Fig. 2d, e and Extended Data Table 1). This revealed a single, six-fold symmetric density in the middle of the hexameric ring (blue mesh in Fig. 2d–f), indicating that one IP6 molecule binds one CACTDSP1 hexamer. Notably, this density coincides precisely with an unknown density feature observed in cryo-electron microscopy (cryo-EM) maps of the HIV-1 Gag hexamer derived from authentic immature virions2 (Extended Data Fig. 3). This further supports the idea that IP6 is a cofactor of Gag assembly in cells and is a structural component of the HIV-1 particle.
IP6 is an asymmetric molecule with multiple stereoisomers, the most abundant of which is the myo form, with a chair configuration of one axial and five equatorial phosphate groups12; this is the most commonly observed stereoisomer in structures of various IP6-binding proteins13,14,15. In our CACTDSP1 structure, the IP6 density is also consistent with the myo form, with the axial phosphate pointing towards the six-helix bundle (6HB) (Fig. 2f). The bound ligand can adopt six energetically equivalent orientations, and the six-fold symmetric density is therefore the sum of these equivalent positions (Extended Data Fig. 4a). More importantly, the bound IP6 is surrounded by two rings of lysine sidechains—Lys290 from the major homology region loop and Lys359 from the 6HB (Fig. 2g). In our previous crystal structure of the CACTDSP1 hexamer in the absence of IP6, sidechain densities for these lysines were not visible, implying that these residues were highly flexible3. In the current structure, these sidechains are better ordered, and in direct ionic contact through their primary ε-amines with the IP6 phosphate groups (Fig. 2g and Extended Data Fig. 4b).
Consistent with the structure, we found that s-CANC mutant proteins in which Lys290 or Lys359 were replaced with alanine (K290A and K359A) were 100-fold less responsive to added IP6 (Extended Data Fig. 5a–d). These results further indicate that both lysine rings are required for productive IP6 binding. K290R and K359R mutants had less pronounced defects but still did not respond to IP6 as well as wild-type s-CANC, consistent with the high degree of lysine conservation in these positions (99.94% for K290 and 99.84% for K359; http://www.hiv.lanl.gov). Furthermore, the K290A and K359A mutations abolished infectivity (Extended Data Fig. 5e). Thus, optimal HIV-1 assembly in cells appears to require lysines at both positions. The results of previous studies that examined the effects of the above mutations on virus budding from cells and on virus infectivity are consistent with our findings16,17,18,19.
The above data suggest that IP6 acts by stabilizing the 6HB and promoting the formation of the immature Gag hexamer. To test this notion, we examined the dynamic behaviour of the CACTDSP1 hexamer by using all-atom molecular dynamics simulations. In the absence of IP6, the six-fold symmetry of the CACTDSP1 hexamer collapsed after 200 ns and did not recover during the 2 μs of simulation (Extended Data Fig. 6 and Supplementary Video 1). By contrast, six-fold symmetry in the presence of IP6 was maintained, particularly at the top of the 6HB, proximal to the IP6-binding site. Other inositol derivatives and mellitic acid (hexacarboxybenzene) also stabilized the 6HB in our simulations, consistent with their ability to also support immature s-CANC assembly in vitro (Extended Data Fig. 6b, c).
We also examined the effect of IP6 on mature capsid assembly, which is mediated by the CA protein that is generated upon Gag proteolysis. We found that IP6 promoted assembly of HIV-1 CA into mature-like structures1,10,20 (Fig. 3a and Extended Data Fig. 7b, d). Compared to immature s-CANC assembly, however, higher amounts of IP6 were required (Extended Data Fig. 7b, d). Mellitic acid (Extended Data Fig. 7c, e) and IP5, but not IP4 or IP3 (Extended Data Fig. 7f, g), stimulated mature CA assembly, although less potently than IP6.
a, Representative negative stain images of mature CA assemblies at pH 6 and 100 mM NaCl with increasing IP6 concentrations (0–1,250 μM). The experiment was repeated five times with similar results. b, Representative image of failed CA R18A assembly even in the presence of 1,250 μM IP6. The experiment was repeated three times with similar results. c, d, Top view (c) and side view (d) of a CA hexamer crystal structure showing the protein in yellow ribbons and unbiased mFo–DFc difference density in blue mesh, contoured at 2.2σ. e, Side views of myo-IP6 docked into the difference density in two possible binding modes. f, Illustration of a single IP6 molecule bound within a chamber enclosed by the N-terminal β-hairpins and the Arg18 ring (magenta). In a second crystal form, IP6 densities were observed both above and below the Arg18 ring (Extended Data Fig. 8).
The mature HIV-1 CA hexamer also contains a positively charged ring, made up of Arg18 sidechains (Arg150 in Gag numbering)21,22. This ring was previously shown to mediate transport of nucleoside triphosphates, which facilitates reverse transcription of the encapsulated genome23. We therefore tested whether IP6 would promote assembly of the HIV-1 CA R18A mutant, and found that it did not (Fig. 3b). HIV-1 virions containing this mutation were also non-infectious17,23 (Extended Data Fig. 5e). We next crystallized the mature CA hexamer in the presence of IP6 (Extended Data Table 1). Although IP6 can bind both above and below the ring (Extended Data Fig. 8), densities were most pronounced in the upper binding site, inside a chamber surrounded by the N-terminal β-hairpins of CA (Fig. 3c–f). Thus, IP6 also binds and promotes assembly of the mature HIV-1 CA lattice.
Our results lead to the following model (Fig. 4). IP6 facilitates the formation of the six-helix CA–SP1 bundle by binding to two rings of primary amines at Lys290 and Lys359, thereby neutralizing otherwise repulsive charges at the centre of the HIV-1 Gag hexamer (Fig. 4c). Although other negatively charged molecules can also bind this pocket, our data suggest that IP6 is the most potent in promoting assembly, probably because it has the most optimal binding geometry. Some 300–400 molecules of IP6—one per hexamer—are incorporated into the virus particle as a structural component of the immature Gag shell (Fig. 4c). During virus maturation, proteolysis of Gag disrupts the 6HB, thus releasing IP6 and at the same time unmasking the Arg18 binding site in mature CA. IP6 then binds to this newly exposed site in CA (Fig. 4d), promoting the formation of CA hexamers and in turn the mature CA lattice. This involvement of a small molecule in two distinct steps in virus assembly, by binding to highly conserved sites, suggests strategies for possible therapeutic intervention in HIV-1 replication.
a, Diagram of HIV-1 Gag, with the indicated positions of R150 (triangle; R18 in mature CA), K290 (circle; K158 in mature CA), K359 (square; K227 in mature CA). Dotted lines indicate protease cleavage sites. b, Diagram of Gag organization in immature virions (left). Following cleavage of Gag by protease (that is, maturation), CA re-organizes to form a mature core around viral RNA (right). c, d, Surface representations of the CASP1 and CA hexamers in the immature (c) and mature virus (d), with IP6 shown in its binding sites. The marked rearrangement of CA upon maturation is evident, as is the change in IP6 binding site between immature and mature viruses. CANTD, blue; CACTD, orange; 6HB, purple; IP6, red.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized and investigators were not blinded to allocation during experiments and outcome assessment.
Protein purification
DNA coding for HIV-1 Gag proteins were cloned into a His6–SUMO vector25. The proteins were expressed in Escherichia coli and purified using standard Ni2+ affinity chromatography followed by cleavage of the SUMO moiety by ULP1 protease. In brief, bacterial pellets were resuspended in buffer and lysed by sonication and cellular debris removed by centrifugation. The supernatant was filtered through a 0.2-μm filter, applied to a Ni2+ affinity resin, and eluted with imidazole. The eluted protein was dialysed overnight in the presence of ULP1 protease, and subjected to Ni2+ chromatography a second time to remove the SUMO tag and ULP1 protease.
All proteins containing the NC domain were purified with additional steps for more stringent removal of nucleic acid. Following bacterial lysis and centrifugation, nucleic acid was precipitated by addition of 0.03% (v/v) polyethyleneimine followed by centrifugation. Ammonium sulfate to 20% saturation was added to the resulting supernatant, and the precipitate was resuspended in buffer (20 mM Tris-HCl, pH 8, 100 mM NaCl, 2 mM TCEP (tris(2-carboxyethyl)phosphine), 5 μM ZnCl2). The protein was then purified by anion exchange and Ni2+ chromatography as above. All purification steps were performed at 4 °C or on ice. All of the final purified proteins, at concentrations of 2–5 mg/ml and having A260/A280 ratios of <0.6, were flash-frozen in liquid nitrogen and stored at –80 °C.
In vitro assembly
Assembly of s-CANC VLPs was performed by dialysing 50 μM protein against buffer (50 mM MES, pH 6 or 50 mM Tris-HCl, pH 8, 100 mM NaCl, 5 μM ZnCl2, 2 mM TCEP) with a single-stranded 50-mer oligonucleotide (GT25) at a 1:5 molar ratio of oligonucleotide to protein for 4 h at 4 °C. All reactions were adjusted to a final volume of 100 μl with buffer following dialysis. Working stocks of 10 mM inositol phosphates were made fresh (IP6, TCI cat# P0409; IP3–IP5, Cayman Chemical cat #s IP3-60960, IP4-60980, and IP5-10009851) with the pH adjusted to 6.0 with NaOH, and added both to the assembly reaction and dialysis buffer. Both s-CASP1 and CACTDSP1 assembly reactions were performed as described for s-CANC but with 500 μM protein and 500 μM IP6. Mature CA assembly was performed by dilution into buffer (50 mM MES, pH 6, 100 mM NaCl) to 250 μM final concentration in the presence of increasing amounts of IP6. Note that under these low-salt conditions, HIV-1 CA does not spontaneously assemble efficiently. The mature reactions were diluted 1:10 before spotting on EM grids. All VLP assemblies were visualized by EM negative staining with uranyl acetate. Quantification was performed by counting particles on at least five images from at least two different assembly reactions. Box plot; centre lines show the medians; box limits indicate the 25th and 75th percentiles as determined by R software24; whiskers extend to minimum and maximum values.
CRISPR knockout
The lentiCRISPR v2 vector was a gift from F. Zhang (Addgene plasmid # 52961)26. The VSV-G expression vector27 was obtained through the NIH AIDS Research and Reference Reagent Program. HEK293FT cells were purchased from Invitrogen. Cell lines were tested for, and showed no mycoplasma contamination. The plasmid v906 is an HIV-1 NL4-3 derived provirus lacking Vpr, Vif, Env, and containing CMV GFP in place of Nef. The construct has several silent restriction sites added to the CA domain of Gag for cloning purposes. The IPPK-targeted guide RNA (5′-AACAGCGCTGCGTCGTGCTG-3′) was cloned into lentiCRISPR v2, which was then used to transduce 293FT cells, followed by selection with puromycin at 1 μg/ml. Clonal isolates of the stably transduced cells were obtained by limiting dilution. To confirm the knockout, genomic DNA was isolated from clonal isolates using the DNeasy blood and tissue kit (Qiagen) following the manufacturer’s protocol. The guide RNA target sequence was amplified from genomic DNA using primers 5′-GAAATGTGTGCCACTGTGTTTA-3′ and 5′-ATGATGGACACACCACTTTCT-3′. The PCR product was directly sequenced.
Infectivity assays
Equivalent numbers of 293FT WT or IPPK KO cells were plated in 35-mm dishes and transfected with 900 ng of v906 and 100 ng of VSV-G. Medium was collected two days post-transfection and frozen at −80 °C to lyse cells in the supernatant. Thawed supernatants were centrifuged at 1,500g for 5 min to remove cellular debris. Infections were performed in fresh 293FT cells. Cells were collected two days later, fixed with 4% paraformaldehyde, and analysed for GFP expression using an Accuri C6 flow cytometer.
Two-dimensional crystallography
CACTDSP1 2D crystals were produced by incubating 0.8 mM protein with 0.8 mM IP6 at room temperature for 30 min. Samples were placed on a carbon-coated grid, washed with 0.1 M KCl, blotted to near dryness and flash frozen by plunging in liquid ethane. Low-dose images were collected on a Tecnai F20 equipped with 4k × 4k Ultrascan CCD camera (Gatan) under low electron-dose conditions (~20 e−/A2). Images were converted to MRC format and manual indexing, unbending, and corrections for CTF were performed with 2dx28.
X-ray crystallography
Purified CACTDSP1 protein (stock = 4.5 mg/ml) was mixed with equal volume of IP6 (stock = 1.4 mM) and incubated briefly at room temperature. Crystals were formed by the vapour diffusion method in sitting drops containing a 1:1 ratio of the protein/IP6 mix and precipitant (0.2 M NaCl, 20% PEG 3,350, 0.1 M Bis-Tris, pH 5.35). Hexagonal plate crystals grew after 2 days of incubation at 17 °C. Crystals were cryoprotected in 25% ethylene glycol and flash-frozen in liquid nitrogen. Diffraction data were collected at the Advanced Photon Source beamline 22-ID and were indexed and scaled with HKL200029. The structure was solved by molecular replacement using as search model one monomer of the previously reported CACTDSP1 hexamer structure (PDB 5I4T)3, with Lys290 and Lys359 sidechains truncated at Cα. Refinement and model building was performed using the PHENIX suite of programs30 and Coot31. Refinement of the protein was first completed before modelling the IP6 and Lys290/Lys359 sidechain densities. The IP6 density was unambiguously identified from mFo−DFc difference maps, and the interpretation that the density was due to bound IP6 was further supported by comparison with difference maps from our previously reported CACTDSP1 structure in the absence of IP63. Given the resolution of the data and crystallographic averaging of the ligand density, we assumed that the bound IP6 was in the myo conformation and refined the ligand as a rigid body with 1/6 occupancy. Only weak residual difference densities were observed after this treatment, suggesting that the modelled IP6 conformation was a reasonable interpretation of the data.
Disulfide-stabilized CA A14C/E45C/W184A/M185A was prepared as previously described32,33. IP6-containing samples were prepared for crystallization as described for CACTDSP1, except that the protein stock concentration in this case was 10 mg/ml. P6 crystals were obtained in precipitant containing 2% Tacsimate, 14% PEG 8,000, 0.1 M Tris, pH 8.4, whereas P212121 crystals were obtained in 8% PEG 8,000, 0.1 M Tris, pH 8.2. Data were collected at Advanced Photon Source beamline 22-BM (P6 form) or 22-ID (P212121 form) and processed with HKL200029. The crystals were isomorphous with previously deposited structures solved in the absence of IP6 (PDB 3H47 and 3H4E)32, and so initial refinement was through rigid body placement of the deposited coordinates (with Arg150 sidechains and waters removed). Refinement of protein-only models were first completed before modelling the IP6 and Arg150 sidechain densities. As with the immature hexamer, IP6 densities were unambiguously identified by mFo−DFc difference maps and by difference density comparisons of CA hexamers crystallized with and without IP6. The IP6 densities in the mature hexamers were modelled as follows. For the P6 crystal form, a single well-defined IP6 density was found inside the β-hairpin chamber (Fig. 3c–e). As in the case of the immature hexamer, the ligand density was also six-fold symmetric due to crystallographic averaging, but in this case indicated at least two binding modes, one with the axial phosphate pointing away from the Arg18 ring and a second pointing towards the ring. Two IP6 molecules were therefore docked into the density, again in the myo form and refined as rigid bodies with 1/12 occupancy (Fig. 3e). Again, only weak residual difference densities were observed after this treatment, suggesting that the modelled IP6 conformations were reasonable interpretations of the data. For the P212121 form, IP6 densities were observed on both sides of the Arg18 ring (there were 2 hexamers in the asymmetric unit and so we observed 4 density features) (Extended Data Fig. 8). These were modelled by docking myo-IP6 in one or two configurations that appeared most consistent with the local density distribution, and then refined as rigid bodies with appropriate occupancy. In this case, significant residual difference densities were observed at the ligand positions after refinement, indicating that additional binding modes were possible. However, we did not attempt to model these multiple overlapping binding modes. The P6 form crystallized in the presence of Tacsimate (Hampton Scientific), which is a mixture of organic carboxylic acids. The excess of negatively charged precipitant therefore appears to have inhibited binding of IP6 below the Arg18 ring; this can be reasonably interpreted to mean that IP6 has greater affinity for the site enclosed by the β-hairpins. The P212121 form crystallized in the absence of Tacsimate, allowing IP6 binding on both sides of the Arg18 ring.
Statistics for all three crystal structures are reported in Extended Data Table 1. Structure visualizations and images were made by using PyMol (Schrödinger Scientific).
Molecular dynamics simulations
The structure of the IP6-bound CACTDSP1 hexamer was used to derive bound and unbound CACTDSP1 models. The IPs and mellitic acid molecules were placed in the central pore of the hexamer between K290 and K359 rings in the corresponding models by aligning the carbons present in the central cyclic ring. All models were then solvated with TIP3P water34 and the salt concentration was set to 150 mM NaCl. Sixteen chloride molecules and twenty-three sodium ions were placed near the hexamer using the CIONIZE plugin in VMD35 to minimize the electrostatic potential. The resulting CACTDSP1 models each contained a total of 30,000 atoms.
After model building, the systems were initially subjected to minimization in two stages, both using the conjugated gradient algorithm36 with linear searching37. Each stage consisted of 10,000 steps of energy minimization. During the first stage, only water molecules and ions were free to move, while the protein and IP molecules, if any, were fixed. In the second stage, the backbone atoms of the protein were restrained with a force constant of 10.0 Kcal mol−1 Å−2 Convergence of the minimizations were confirmed once the variances of gradients were not greater than 1 Kcal mol−1 Å−1. During thermalization the systems were heated from 50 K to 310 K in 20 K increments over 1 ns. Subsequently, the systems were equilibrated, while the backbone atoms of CACTDSP1 were restrained. The positional restraints were gradually released at a rate of 1.0 Kcal mol−1 Å−2 per 400 ps from 10.0 to 0.0 Kcal mol−1 Å−2. NAMD 2.1238 was employed during minimization/thermalization and equilibration steps.
Simulations of IP6-bound and unbound CACTDSP1 were then performed on the special purpose computer Anton239 in the Pittsburgh supercomputing centre for 2 μs. The CHARMM 36m40 force-field was used for all simulations. Parameters for IP6 were derived by analogy following the CGENFF protocol41. During the simulation, the temperature (310 K) and pressure (1 atm) were maintained by using the Multigrator integrator42 and the simulation time-step was set to 2.5 fs/step, with short-range forces evaluated at every time step, and long-range electrostatics evaluated at every second time step. Short-range non-bonded interactions were cut off at 17 Å; long range electrostatics were calculated using the k-Gaussian Split Ewald method43.
Simulations of CACTDSP1 bound to IP3, IP4, and IP5 were performed for 2 μs on TACC Stampede 2 using NAMD 2.1238. The molecular simulations were conducted under isothermal (310 K) and isobaric (1 atm) conditions, regulated by the Langevin thermostat44 and the Nosé-Hoover Langevin piston45,46, respectively. All bonds to hydrogen atoms were constrained with the SHAKE algorithm47. A time step of 2 fs was used for all simulations. Long-range electrostatics were calculated using the Particle-Mesh-Ewald method, as implemented in NAMD38, with a cutoff of 1.2 nm. Full electrostatic interactions were calculated every two time steps while nonbonded interactions were performed every time step.
Analysis of MD simulations
Root-mean-square deviations (RMSDs) and root mean square fluctuations (RMSFs) of the Cα of the CACTDSP1 hexamers were computed using the measure command in VMD35. Before RMSD and RMSF calculations, the structure of the hexamer was aligned to a common reference. RMSFs of each monomer in a central hexamer were calculated to obtain RMSF standard deviations of an entire hexamer.
Reporting summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Coordinates and structure factors have been deposited at the RCSB Protein Data Bank (PDB) database, under accession numbers 6BHR, 6BHT, and 6BHS. All other data are available from the authors on request; see author contributions for specific datasets.
Change history
29 August 2018
In this Letter, the Protein Data Bank (PDB) accessions were incorrectly listed as ‘6BH5, 6BHT and 6BHS’ instead of ‘6BHR, 6BHT and 6BHS’; this has been corrected online.
References
Gross, I. et al. A conformational switch controlling HIV-1 morphogenesis. EMBO J. 19, 103–113 (2000).
Schur, F. K. et al. An atomic model of HIV-1 capsid-SP1 reveals structures regulating assembly and maturation. Science 353, 506–508 (2016).
Wagner, J. M. et al. Crystal structure of an HIV assembly and maturation switch. eLife 5, e17063 (2016).
Keller, P. W., Adamson, C. S., Heymann, J. B., Freed, E. O. & Steven, A. C. HIV-1 maturation inhibitor bevirimat stabilizes the immature Gag lattice. J. Virol. 85, 1420–1428 (2011).
Wang, M. et al. Quenching protein dynamics interferes with HIV capsid maturation. Nat. Commun. 8, 1779 (2017).
Letcher, A. J., Schell, M. J. & Irvine, R. F. Do mammals make all their own inositol hexakisphosphate? Biochem. J. 416, 263–270 (2008).
Campbell, S. et al. Modulation of HIV-like particle assembly in vitro by inositol phosphates. Proc. Natl Acad. Sci. USA 98, 10875–10879 (2001).
Datta, S. A. K. et al. Interactions between HIV-1 Gag molecules in solution: an inositol phosphate-mediated switch. J. Mol. Biol. 365, 799–811 (2007).
Munro, J. B. et al. A conformational transition observed in single HIV-1 Gag molecules during in vitro assembly of virus-like particles. J. Virol. 88, 3577–3585 (2014).
von Schwedler, U. K. et al. Proteolytic refolding of the HIV-1 capsid protein amino-terminus facilitates viral core assembly. EMBO J. 17, 1555–1568 (1998).
Accola, M. A., Strack, B. & Göttlinger, H. G. Efficient particle production by minimal Gag constructs which retain the carboxy-terminal domain of human immunodeficiency virus type 1 capsid-p2 and a late assembly domain. J. Virol. 74, 5395–5402 (2000).
Loewus, F. A. & Murthy, P. P. N. Myo-inositol metabolism in plants. Plant Sci. 150, 1–19 (2000).
van Galen, J. et al. Interaction of GAPR-1 with lipid bilayers is regulated by alternative homodimerization. Biochim. Biophys. Acta 1818, 2175–2183 (2012).
Ouyang, Z., Zheng, G., Tomchick, D. R., Luo, X. & Yu, H. Structural basis and IP6 requirement for Pds5-dependent cohesin dynamics. Mol. Cell 62, 248–259 (2016).
Macbeth, M. R. et al. Inositol hexakisphosphate is bound in the ADAR2 core and required for RNA editing. Science 309, 1534–1539 (2005).
Chang, Y. F., Wang, S. M., Huang, K. J. & Wang, C. T. Mutations in capsid major homology region affect assembly and membrane affinity of HIV-1 Gag. J. Mol. Biol. 370, 585–597 (2007).
von Schwedler, U. K., Stray, K. M., Garrus, J. E. & Sundquist, W. I. Functional surfaces of the human immunodeficiency virus type 1 capsid protein. J. Virol. 77, 5439–5450 (2003).
Melamed, D. et al. The conserved carboxy terminus of the capsid domain of human immunodeficiency virus type 1 gag protein is important for virion assembly and release. J. Virol. 78, 9675–9688 (2004).
Rihn, S. J. et al. Extreme genetic fragility of the HIV-1 capsid. PLoS Pathog. 9, e1003461 (2013).
Ganser-Pornillos, B. K., von Schwedler, U. K., Stray, K. M., Aiken, C. & Sundquist, W. I. Assembly properties of the human immunodeficiency virus type 1 CA protein. J. Virol. 78, 2545–2552 (2004).
Ganser-Pornillos, B. K., Cheng, A. & Yeager, M. Structure of full-length HIV-1 CA: a model for the mature capsid lattice. Cell 131, 70–79 (2007).
Pornillos, O., Ganser-Pornillos, B. K. & Yeager, M. Atomic-level modelling of the HIV capsid. Nature 469, 424–427 (2011).
Jacques, D. A. et al. HIV-1 uses dynamic capsid pores to import nucleotides and fuel encapsidated DNA synthesis. Nature 536, 349–353 (2016).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2013).
Malakhov, M. P. et al. SUMO fusions and SUMO-specific protease for efficient expression and purification of proteins. J. Struct. Funct. Genomics 5, 75–86 (2004).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Chang, L. J., Urlacher, V., Iwakuma, T., Cui, Y. & Zucali, J. Efficacy and safety analyses of a recombinant human immunodeficiency virus type 1 derived vector system. Gene Ther. 6, 715–728 (1999).
Gipson, B., Zeng, X., Zhang, Z. Y. & Stahlberg, H. 2dx-user-friendly image processing for 2D crystals. J. Struct. Biol. 157, 64–72 (2007).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Pornillos, O. et al. X-ray structures of the hexameric building block of the HIV capsid. Cell 137, 1282–1292 (2009).
Pornillos, O., Ganser-Pornillos, B. K., Banumathi, S., Hua, Y. & Yeager, M. Disulfide bond stabilization of the hexameric capsomer of human immunodeficiency virus. J. Mol. Biol. 401, 985–995 (2010).
Jorgensen, W. L. & Jenson, C. Temperature dependence of TIP3P, SPC, and TIP4P water from NPT Monte Carlo simulations: seeking temperatures of maximum density. J. Comput. Chem. 19, 1179–1186 (1998).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38, 27–28 (1996).
Fletcher, R. & Reeves, C. M. Function minimization by conjugate gradients. Comput. J. 7, 149–154 (1964).
Sun, W. & Yuan, Y.-X. Optimization Theory and Methods: Nonlinear Programming (Springer US, 2006)
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Shaw, D. E. et al. In Proc. Intl Conf. High Performance Computing, Networking, Storage and Analysis 41–53 (IEEE Press, New Orleans, 2014).
Best, R. B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone φ, ψ and side-chain χ(1) and χ(2) dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Vanommeslaeghe, K. et al. CHARMM general force field: A force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Lippert, R. A. et al. Accurate and efficient integration for molecular dynamics simulations at constant temperature and pressure. J. Chem. Phys. 139, 164106 (2013).
Shan, Y., Klepeis, J. L., Eastwood, M. P., Dror, R. O. & Shaw, D. E. Gaussian split Ewald: A fast Ewald mesh method for molecular simulation. J. Chem. Phys. 122, 54101 (2005).
Berendsen, H. J. C., Postma, J. P. M., Vangunsteren, W. F., Dinola, A. & Haak, J. R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Martyna, G. J., Tobias, D. J. & Klein, M. L. Constant pressure molecular dynamics algorithms. J. Chem. Phys. 101, 4177–4189 (1994).
Feller, S. E., Zhang, Y. H., Pastor, R. W. & Brooks, B. R. Constant pressure molecular dynamics simulation—the Langevin Piston Method. J. Chem. Phys. 103, 4613–4621 (1995).
Ryckaert, J.-P., Ciccotti, G. & Berendsen, H. J. Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J. Comput. Phys. 23, 327–341 (1977).
Acknowledgements
We thank J. Briggs for discussions and reading of the manuscript. This work was supported by the National Institutes of Health (NIH) grants R01-GM107013 (V.M.V.), R01-GM105684 (G. W. Feigenson), P30-GM110758 and P50-GM082251 (J.R.P.), R01-AI129678 (O.P. and B.K.G.-P.), U54-GM103297 (O.P.), and R01-GM110776 (M.C.J.). F.K.M.S. was supported by Deutsche Forschungsgemeinschaft grant BR 3635/2-1 awarded to J. A. G. Briggs. J.M.W. was supported by NIH postdoctoral fellowship grant F32-GM115007. Anton computer time was provided by the Pittsburgh Supercomputing Center (PSC) through NIH grant R01-GM116961. The Anton machine at PSC was generously made available by D. E. Shaw Research. This work used the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by National Science Foundation (NSF) grant number OCI-1053575. Specifically, it used the Bridges system, which is supported at PSC by NSF award number ACI-1445606. Some of The EM work was conducted at the Molecular Electron Microscopy Core facility at the University of Virginia.
Reviewer information
Nature thanks E. Freed and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
R.A.D. performed protein purification and in vitro assembly. F.K.M.S. did comparative analyses of cryo-EM and crystal structure data. K.K.Z, J.M.W., B.K.G.-P. and O.P. carried out crystallization trials and structure determination. B.K.G.-P. performed 2D cryo-EM. J.R.P. and C.X. performed all-atom MD simulations. T.D.L., C.L.R. and M.C.J., performed cell biology and virology. The manuscript was written primarily by R.A.D., J.R.P., B.K.G.-P., O.P. and V.M.V. The project was originally conceived by R.A.D., with input from all authors throughout experimentation and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Effect of acidic molecules on immature s-CANC assembly.
a, Representative negative-stain electron microscopy images. Scale bars, 200 nm. The experiment was repeated twice with similar results. b, Number of immature VLPs per 55 µm2. n = 5, mean shown above box plots; centre lines show medians; box limits indicate 25th and 75th percentiles as determined by R software; whiskers extend to minimum and maximum values.
Extended Data Fig. 2 s-CANC and s-CASP1 VLPs.
a–c, Representative negative-stain electron microscopy images of s-CANC (a), s-CASP1 (b) and s-CA (c) proteins assembled in the absence of GT25 and in the presence of the indicated IP6 concentrations. Scale bars, 200 nm. d, Diameters of immature VLPs; mean diameter above plot; n below plot. Centre lines show medians; box limits indicate 25th and 75th percentiles as determined by R software; whiskers extend to minimum and maximum values.
Extended Data Fig. 3 Comparison of the HIV-1 Gag cryo-EM structure with the CACTDSP1–IP6 crystal structure.
a, The crystal structure of CACTDSP1 bound to IP6 (cyan) was superimposed on a previously described model of the CA-SP1 segment build into cryo-EM densities of immature HIV-1 particles (PDB 5L93, orange2). Note the close correspondence in K359 rotamers, which were modelled independently in the two structures. For visualization purposes, only one of the six possible IP6 conformations is displayed. b, RMSD calculations of the crystal structure and PDB 5L93. For full-length (residues 149–237) and CA-SP1 (residues 223–237), the RMSDs were calculated only for the atoms that were modelled in both maps. If a sidechain was not modelled, the entire residue was omitted from the calculation. The overall agreement of the models is very high, indicating that the crystal structure corresponds well with conformations found in the virus. c, The CACTDSP1 bound to IP6 (orange and red, respectively) was fitted into two previously published cryo-EM densities2 from VLPs collected from cells (EMD-2706 and EMD-4017). Both maps are shown at 8.8 Å, which is the resolution of the lower resolved map, EMD-2706. In the zoomed insets, only the density corresponding to IP6 is shown. Matching of models and maps and RMSD calculations were performed in Chimera.
Extended Data Fig. 4 Interpretation of the IP6 density in the immature CACTDSP1 hexamer structure.
a, Top and side views of the unbiased mFo–DFc difference density (blue mesh, 2σ) ascribed to the bound IP6. Shown are six IP6 molecules docked in six rotationally equivalent positions, consistent with the six-fold rotational symmetric density. b, Top view of the docked IP6 molecules within the CACTDSP1 hexamer. Unbiased mFo–DFc difference densities (blue mesh) are also shown for both the bound IP6 and sidechains of Lys290 (green) and Lys359 (cyan). Density for Lys359 is more pronounced, which we interpret to mean that this residue adopts a more restricted range of rotamers for binding IP6.
Extended Data Fig. 5 Quantification of wild-type and mutant HIV s-CANC assembly at pH 6 and pH 8.
a, c, Number of immature (purple) and mature (orange) VLPs per 55 μm2 without (−) and with (+) 10 μM IP6 at pH 6 and pH 8. Mean above and n below box plots. Centre lines show medians; box limits indicate 25th and 75th percentiles as determined by R software; whiskers extend to minimum and maximum values. b, d, Representative negative stain electron microscopy images of wild-type and mutant s-CANC assembly in the absence (−) and presence (+) of 10 μM IP6 at pH 6 and pH 8. Scale bar, 400 nm. Repeated three times with similar results. e, Infectivity relative to wild-type virus of IP6 binding residues mutated to alanine and CA residue numbering in parenthesis. Error bars represent s.d., individual data points represented as dots; from four independent experiments.
Extended Data Fig. 6 IP6 modulates the stability of the 6HB.
a, Structural changes observed after 2 μs of molecular dynamics simulations of CACTDSP1 with and without bound IP6. b, RMSDs of the ligand-bound and unbound forms of the CACTDSP1 hexamer. c, RMSFs of the central hexamer during the simulation. The RMSF was averaged over the six central monomers; dashed line shows the s.d. for each residue.
Extended Data Fig. 7 Quantification of mature HIV-1 CA assembly and VLP diameter at pH 6.
a, Example of CA assembly in the absence of IP6 or mellitic acid. b, c, Representative negative-stain electron microscopy images of assemblies induced by IP6 (b) and mellitic acid (c). Scale bars, 200 nm. Tubes (T), cones (C), and other (O) morphologies are marked by coloured arrowheads. a–c, Repeated four times with similar results. d, Number of assembled CA tubes (blue), cones (orange) and other (green) per 55 μm2 at increasing IP6 concentrations. Mean shown above plots, n = 5. e, Number of assembled tubes (blue), cones (orange) and other (green) per 55 μm2 at increasing mellitic acid concentrations. Mean shown above and n below box plots. f, Representative images of mature VLPs assembled with IP5 and IP6 at 50 mM NaCl. Scale bars, 100 nm. Repeated three times with similar results. g, Number of CA VLPs per 10 µm2 without and with IP3, IP4, IP5, and IP6. Mean shown above, n = 5. d, e, g, Centre lines show medians; box limits indicate 25th and 75th percentiles as determined by R software; whiskers extend to minimum and maximum values.
Extended Data Fig. 8 Crystal structure of IP6 bound to the mature CA hexamer.
a, b, Top view (a) and side view (b) of a second CA hexamer crystal structure (P212121 space group) showing the protein in yellow ribbons and unbiased mFo–DFc difference density in blue mesh, contoured at 2.5σ. c, Close-up view showing IP6 densities both above and below the ring of Arg18 residues (magenta).
Supplementary information
Supplementary Figure 1
FACs gating strategy. a, Events were plotted along forward scatter (FSC) and side scatter (SSC) axises using FlowJo. Events with the right morphology were gated as "Live" cells and the position of the gate was copied onto all samples. b, Events from the "Live" gate were isolated and plotted along GFP and RFP intensity axes. The "Non-Fluorescent" gate was created based on a HEK293FT fluorescence negative sample (plot not shown) and the gate was copied onto all isolated "Live" samples. The "GFP-Positive" gate was created based on a HEK293FT GFP positive sample (plot not shown) and the gate was copied onto all isolated "Live" samples. Representative comparison of two cell types transduced with HIV Env deficient virus with a GFP reporter (HEK293FT = WT and IPPK KO = HEK293FT with inositol-pentakisphosphate 2-Kinase knocked out).
Video 1: 2-μs trajectories of IP6-unbound (left) and bound (right) CACTDSP1 models.
The protein hexamer is shown in cartoon representation and IP6 molecule is in stick representation.
Rights and permissions
About this article
Cite this article
Dick, R.A., Zadrozny, K.K., Xu, C. et al. Inositol phosphates are assembly co-factors for HIV-1. Nature 560, 509–512 (2018). https://doi.org/10.1038/s41586-018-0396-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0396-4
This article is cited by
-
HIV-1 capsids enter the FG phase of nuclear pores like a transport receptor
Nature (2024)
-
HIV-1 is dependent on its immature lattice to recruit IP6 for mature capsid assembly
Nature Structural & Molecular Biology (2023)
-
Enrich and switch: IP6 and maturation of HIV-1 capsid
Nature Structural & Molecular Biology (2023)
-
Molecular architecture and conservation of an immature human endogenous retrovirus
Nature Communications (2023)
-
Structural basis of HIV-1 maturation inhibitor binding and activity
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.