Abstract
Medulloblastoma is a malignant childhood cerebellar tumour type that comprises distinct molecular subgroups. Whereas genomic characteristics of these subgroups are well defined, the extent to which cellular diversity underlies their divergent biology and clinical behaviour remains largely unexplored. Here we used single-cell transcriptomics to investigate intra- and intertumoral heterogeneity in 25 medulloblastomas spanning all molecular subgroups. WNT, SHH and Group 3 tumours comprised subgroup-specific undifferentiated and differentiated neuronal-like malignant populations, whereas Group 4 tumours consisted exclusively of differentiated neuronal-like neoplastic cells. SHH tumours closely resembled granule neurons of varying differentiation states that correlated with patient age. Group 3 and Group 4 tumours exhibited a developmental trajectory from primitive progenitor-like to more mature neuronal-like cells, the relative proportions of which distinguished these subgroups. Cross-species transcriptomics defined distinct glutamatergic populations as putative cells-of-origin for SHH and Group 4 subtypes. Collectively, these data provide insights into the cellular and developmental states underlying subtype-specific medulloblastoma biology.
Similar content being viewed by others
Introduction
Medulloblastoma (MB) comprises a series of molecularly and clinically diverse malignant childhood cerebellar tumours1. While advances in treatment have improved survival, many patients suffer from neurological sequelae or still succumb to their disease. Genomic studies of bulk patient cohorts have defined four consensus molecular subgroups (WNT, SHH, Group 3 and Group 4)2, each characterized by discrete genomic landscapes, patient demographics and clinical phenotypes3,4,5,6,7. The association between genotypes, transcriptional profiles, and patient age at diagnosis suggests that distinct MB subgroups arise from the transformation of different cell types in precise spatiotemporal patterns. Such genotype-to-cell-type associations have been partially investigated for WNT and SHH MBs, which are thought to originate from cells in the extracerebellar lower rhombic lip8 and from cerebellar granule neuron progenitors (GNPs)9,10, respectively. By contrast, cellular origins of Group 3 and Group 4 MB remain unconfirmed. Overlapping transcriptional and epigenetic signatures observed in bulk profiling studies have consistently hampered definitive classification of Group 3 and Group 4 tumours and suggest that they may share common developmental origins3,11. Thus, a better understanding of MB cellular composition and substructure according to subgroup is a critical goal, especially for the poorly characterized Group 3 and Group 4.
Single-cell RNA sequencing (scRNA-seq) has emerged as a powerful method to comprehensively characterize cellular states in healthy and diseased tissues12. Whereas in central nervous system malignancies, scRNA-seq has been applied to decipher adult and paediatric gliomas13,14,15,16, such approaches have yet to be deployed across MB subgroups. Here we applied full-length scRNA-seq across consensus MB subgroups to infer cellular trajectories, deconvolute bulk MB expression cohorts and nominate developmental origins. We find that WNT, SHH and Group 3 tumours exhibit subgroup-specific cellular trajectories that consist of malignant undifferentiated and differentiated neuronal-like populations, whereas Group 4 tumours recapitulate more differentiated populations of known lineage. Collectively, these data provide insights into the molecular and cellular architecture of MB across all subgroups, with the potential to inform future studies aimed at improving patient outcomes.
MB and cerebellar transcriptomes
We prospectively obtained fresh surgical resections from 25 patients with MB (23 diagnostic samples and 2 recurrences) and 11 patient-derived xenograft (PDX) models (Fig. 1a, b, Extended Data Fig. 1a, b, Supplementary Table 1a). Each tumour sample was classified on a molecular level using DNA methylation arrays17 (Fig. 1b, Extended Data Fig. 1b). The majority of tumours were also characterized by whole-genome (n = 5) or whole-exome (n = 12) sequencing (Fig. 1b, Supplementary Table 1b). To perform full-length scRNA-seq, cells were dissociated, sorted for viability and profiled using the Smart-seq2 protocol18 (see Methods). Analysis of known subgroup-specific signature genes19 demonstrated expected expression patterns (Extended Data Fig. 1b, c). Pairwise correlation of aggregated scRNA-seq and DNA methylation array data further substantiated subgroup classifications and PDX model fidelity (Extended Data Fig. 1d). Scoring single cells using published transcriptional signatures revealed that WNT and SHH tumours consist exclusively of cells scoring highly for their respective signatures. Conversely, cells derived from Group 3 and Group 4 tumours exhibited some degree of transcriptional overlap (Extended Data Fig. 1e). In total, 8,734 single cells passed quality control, with a median of 4,561 genes detected per cell (Supplementary Table 1a).
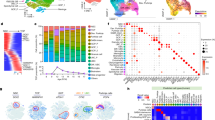
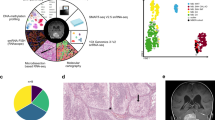
a, Summary of human MB and developing mouse cerebellar scRNA-seq datasets. E10, embryonic day 10; P10, postnatal day 10. b, Clinical and molecular details of the MB single-cell cohort. Asterisks indicate recurrent MB samples (n = 2/25). Samples subjected to whole-genome (G) or whole-exome (E) sequencing are indicated. c, t-distributed stochastic neighbour embedding (t-SNE) representation of the cerebellar scRNA-seq dataset. Colours represent assigned cerebellar cell types. d, Radar plot showing the CCA coefficients between each mouse cerebellar cell type (n = 18) and bulk human MB expression data3. Asterisks indicate significant correlations (FDR corrected permutation test, P < 0.05). Malig., malignant; Glu, glutamatergic; GABA, GABAergic; seq., sequencing.
To classify single cells into malignant and non-malignant subsets, we used two complementary strategies. First, we inferred genome-wide copy-number variations (CNVs) from the scRNA-seq data as previously described13 (see Methods). This analysis identified large-scale genomic gains and losses in most (21 out of 25) patient samples, including hallmark alterations such as monosomy 6 (WNT) and isochromosome 17q (Groups 3 and 4; Extended Data Fig. 2a–e). Few cells (n = 36) from these patients lacked discernable CNVs (see Methods). Second, we clustered single cells across all samples according to their transcriptional profiles. A minority of single cells in our cohort clustered with reference immune cells (n = 6) or oligodendrocytes (n = 22) (Extended Data Fig. 3a, b). All cells that lacked CNVs and/or clustered with normal reference populations were deemed as non-malignant and excluded from further analysis (n = 43). Across individual tumours, 96–100% of cells were classified as malignant, consistent with previous estimates of high MB tumour cell fractions based on genome sequencing20. We further validated these assignments by quantifying genetic mutations identified by bulk tumour DNA sequencing in our scRNA-seq data (1,937 mutant and 1,952 wild-type transcripts detected; see Methods, Extended Data Fig. 3c–f).
To relate MB single-cell profiles to normal developmental hierarchies, we leveraged recently generated scRNA-seq data for mouse cerebellar development spanning 13 embryonic and early postnatal time points21 (total of 78,156 single cells; Fig. 1a, c, Extended Data Fig. 4a–e, Supplementary Table 1c). Canonical correlation analysis (CCA; see Methods) facilitated cross-species comparisons between our mouse cerebellar single-cell, human MB single-cell and bulk3 expression datasets. SHH MB was highly correlated with GNP populations (cosine distance = 0.54), consistent with literature9,10, supporting GNPs as the cell-of-origin for this subgroup (Fig. 1d, Extended Data Fig. 4f). Notably, Group 4 MB was highly correlated with unipolar brush cells (UBC; cosine distance = 0.50) and glutamatergic cerebellar nuclei (GluCN; cosine distance = 0.49). By contrast, we did not detect high-confidence correlations between any cerebellar populations and either WNT or Group 3 subgroups.
Malignant trajectories within WNT MB
Children with WNT MB account for about 10% of patients with MB and have an excellent prognosis22. Somatic CTNNB1 mutations or germline APC mutations, both of which drive constitutive WNT signalling, are found in nearly all WNT MBs3,23. Five WNT tumours were included in our dataset. Pairwise correlation analysis revealed multiple distinct transcriptional states that were consistently identified within these tumours (Fig. 2a). Inferring CNVs from our scRNA-seq data identified four cases with monosomy 6, a stereotypic genomic feature of this subgroup (Fig. 2b, Extended Data Fig. 2a). The fifth case (BCH807) exhibited chromosome 19 gain and was negative for nuclear β-catenin by immunohistochemistry (data not shown), both of which are atypical characteristics for this subgroup despite high-confidence molecular classification as WNT MB (Fig. 1b). SJ99 exhibited marked heterogeneity at both a transcriptional and genetic level, with evidence for two distinct subclones. Subclone SJ99-A exhibited monosomy 6 and chromosome 17p loss, whereas subclone SJ99-B exhibited broad gains and losses affecting nearly every chromosome. Investigation of genetically supported single-nucleotide variants (SNVs) confirmed expression of mutant transcripts in 57.2% of cells (including key WNT MB driver genes CTNNB1, DDX3X and TP53; Fig. 2c).
a, Pairwise correlations between the expression profiles of all WNT MB cells (n = 1,780). Cells are ordered by hierarchical clustering within each sample or genetic subclone. NM, non-malignant. b, Single-cell derived CNVs by chromosome (columns). c, Mutant and wild-type transcripts detected in single cells. d, Relative expression of 90 genes representing WNT MB metaprograms (rows) across cells from MUV44 and SJ99. Cells positive for the cell cycle program (WNT-A) are indicated. e, Scatter plot of the WNT-C and WNT-D metaprogram expression scores for all WNT MB cells.
Non-negative matrix factorization (NMF) was applied to define underlying transcriptional programs specific to each tumour (Extended Data Fig. 5a, b, Supplementary Table 2a, see Methods). This analysis revealed highly similar programs in all five WNT MBs, which we grouped accordingly into four metaprograms (WNT-A, WNT-B, WNT-C and WNT-D). To interpret the characteristics of each metaprogram, we evaluated their underlying gene signatures. WNT-A contained numerous markers of cell cycle activity (such as TOP2A, CDK1 and RRM2; P < 0.001, Fisher’s exact test; Supplementary Table 2b). WNT-C was characterized by markers of neurogenesis or neuronal differentiation (such as STMN2, KIF5C and SYT11; P < 0.001; Fig. 2d). WNT-B consisted of ribosomal and metabolic genes (NME2, HK2 and PGM5), and WNT-D contained select WNT-pathway genes (LRP4 and APCDD1) and immediate early response genes (JUNB and EGR1; Fig. 2d). Cells scoring highest for both WNT-B and WNT-D expressed elevated levels of additional canonical WNT pathway genes (DKK2, AXIN2 and WIF1) and MYC (Extended Data Fig. 5c, Supplementary Table 2c). We interpret these metaprograms as reflecting cell cycle activity (WNT-A), neuronal-like differentiation (WNT-C) and two WNT-driven states (WNT-B and WNT-D), with WNT-B characterized by elevated protein biosynthesis and metabolism (Fig. 2d). RNA in situ hybridization performed on the same tumours validated the expression of specific metaprogram marker genes in subpopulations of cells (Extended Data Fig. 5d). Moreover, scoring each cell in our cohort for these four metaprograms defined a putative developmental trajectory for WNT MB, with cell cycle activity restricted to cells that were both high in WNT-B and low in WNT-C and WNT-D (Fisher’s exact test, P < 0.001), suggesting that this subpopulation fuels WNT MB growth (Fig. 2e). Notably, each metaprogram was identified in at least four samples (Extended Data Fig. 5a), suggesting that the programs reflect shared features of WNT MB.
Developmental trajectories within SHH MB
As the dominant subgroup in both infants (≤3 years old) and adults (≥18 years old)22, SHH MB accounts for about one third of all patients with MB. Outcomes are heterogeneous and associated with underlying genetics, demographics and clinical features24. Our dataset included three patients with SHH MB, ranging in age from 3 to 13 years (Fig. 1b). Pairwise correlation and unsupervised NMF analysis revealed three transcriptional programs (SHH-A, -B and -C) shared among these tumours (Fig. 3a, Extended Data Fig. 6a, b, Supplementary Table 2a, b). SHH-A contained markers of cell cycle activity (for example, TOP2A, CDK1 and RRM2; P < 0.001, Fisher’s exact test). SHH-B was enriched for ribosomal genes and translational initiation and elongation factors (EIF3E and EEF1A1; P < 0.001), and markers of canonical SHH signalling (PTCH1 and BOC; P < 0.001; Fig. 3b). SHH-C was defined by markers of neuronal differentiation (STMN2, MAP1B, TUBB2B and SEMA6A; P < 0.001; Fig. 3b). We interpret these programs as reflecting cell cycle activity (SHH-A), undifferentiated progenitors (SHH-B) and more differentiated neuronal-like programs (SHH-C). Scoring each SHH MB cell for these programs defined a putative developmental trajectory, with proliferating cells restricted to undifferentiated progenitors (Fig. 3c). These respective programs were partially recapitulated in SHH subgroup PDX models (Extended Data Fig. 6c, d).
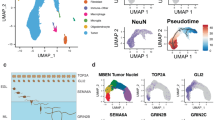
a, Pairwise correlations between the expression profiles of all SHH MB cells (n = 1,135). NM, non-malignant. b, Relative expression of 60 genes representing SHH MB metaprograms across cells from MUV41 and SJ577. Cells positive for the cell cycle program (SHH-A) are indicated. c, Scatter plot of the normalized SHH-B and SHH-C metaprogram expression scores for all SHH MB cells. d, CCA coefficients between SHH MB single-cells and mouse glutamatergic cell types. Cells are ordered as in b. e, Patient age (top), expression levels of ATOH1 and NEUROD1 (middle), and CCA coefficients between tumours and mouse glutamatergic cell types (bottom) for bulk SHH MBs3 (n = 100). f, Expression of Neurod1 (left) and Atoh1 (right) in the mouse cerebellum. In situ hybridization data were obtained from the Allen Developing Mouse Brain Atlas (© 2008 Allen Institute for Brain Science. Allen Developing Mouse Brain Atlas http://developingmouse.brain-map.org). P4, postnatal day 4.
To investigate the developmental significance of these findings, we used CCA to compare SHH MB metaprograms to mouse cerebellar populations. SHH-B correlated with undifferentiated UBC–GNP and GNP populations, whereas SHH-C correlated with UBC–GN intermediate and differentiated granule neuron populations (Fig. 3d, Extended Data Fig. 7a–d). To validate these observations in a larger cohort, we implemented a focused analysis of UBC, GNP and granule neuron populations, assessing correlations between these cell types and bulk SHH MB expression profiles (Fig. 3e, Extended Data Fig. 7e). This analysis broadly split SHH MBs into two age-associated categories: infant tumours correlated with intermediate and mature granule neurons (marked by high expression of NEUROD1), whereas adult tumours correlated with GNPs and mixed UBC and GN progenitors (marked by high expression of ATOH1; Fig. 3e, f, Extended Data Fig. 7f–j). Together, our data suggest that infant and adult SHH MBs are enriched for temporally distinct GNP (or UBC) populations and/or have distinct differentiation capacities, further supporting their divergent biology25,26,27.
Malignant programs within Group 3/4 MB
Group 3 and Group 4 tumours account for about 60% of MB diagnoses and remain the least understood with respect to disease biology and developmental origins7. Group 3 tumours are frequently metastatic at diagnosis and are typified by genomic amplification or overexpression of MYC, which is associated with unfavourable outcomes11,28. Group 4 tumours are metastatic at diagnosis in approximately one third of patients and harbour recurrent chromatin modifier alterations28,29. Recent bulk-profiling studies have demonstrated marked molecular and clinical heterogeneity in Group 3 and Group 4, with a subset of tumours exhibiting overlapping molecular signatures that confound robust classification3,30,31.
On the basis of this prior knowledge, we performed a combined analysis of the scRNA-seq data for all 17 Group 3 and Group 4 tumours. Pairwise correlation analysis of single cells largely discriminated between subgroups, with a subset of ‘intermediate’ tumours exhibiting transcriptional ambiguity (MUV34, BCH825 and SJ625; Fig. 4a). NMF analysis of the combined series identified three distinct transcriptional programs (Group 3/4-A, -B and -C) (Extended Data Fig. 8a–c, Supplementary Table 2a, b). Group 3/4-A contained markers of cell cycle activity (for example, TOP2A, CDK1 and RRM2; P < 0.001, Fisher’s exact test). Group 3/4-B was primarily characterized by ribosomal and translational initiation/elongation genes (EIF3E and EEF1A1; P < 0.001; Fig. 4b) as well as by MYC and MYC target genes (for example, HLX). Group 3/4-C contained well-recognized neuronal lineage markers (STMN2, SOX4, ZIC1 and SYT11; P < 0.01; Fig. 4b). We interpret that these programs reflect cell cycle activity (Group 3/4-A), undifferentiated progenitor-like programs with high MYC activity (Group 3/4-B) and differentiated neuronal-like programs (Group 3/4-C; Fig. 4b).
a, Pairwise correlations between the expression profiles of all Group 3/4 MB cells (n = 4,873). NM, non-malignant. DNA methylation-based subgroup prediction scores are indicated (top). b, Relative expression of 60 genes representing Group 3/4 MB metaprograms from SJ617, MUV34 and BCH1031. Cells positive for the cell cycle program (Group 3/4-A) are indicated. c, d, Scatter plots of the Group 3/4-B and Group 3/4-C metaprogram expression scores for all Group 3 (c) and Group 4 (d) MB cells.
Scoring each Group 3/4 MB cell for these programs revealed distinct patterns: prototypic Group 3 tumours were dominated (>88% of cells) by the undifferentiated progenitor-like program (Group 3/4-B), whereas the differentiated neuronal-like program (Group 3/4-C) was observed in almost all cells (>95%) from prototypic Group 4 tumours, consistent with their neuronal differentiation phenotype11,28 (Fig. 4c, d, Supplementary Table 2d). Group 3 tumours with MYC amplifications (SJ17 and MUV29; Extended Data Fig. 2c) lacked neuronal differentiation altogether (<2% of cells), suggesting that oncogenic MYC expression may potentiate an undifferentiated progenitor-like state. Notably, Group 3/4 intermediate tumours (MUV34, BCH825 and SJ625) comprised a mixture of both malignant cell states, containing 12–20% of cells characterized by the undifferentiated program, with the remainder of cells characterized by the differentiated program. These transcriptional programs were also evident in nine Group 3/4 PDX models (Extended Data Fig. 8d, e). Our results indicate that Group 3/4 MBs contain cells along a common continuum of neuronal differentiation.
The observation that Group 3 and Group 4 MBs both contained cells scoring high for the neuronal-like differentiation program (Group 3/4-C) prompted us to examine whether varying proportions of cells with this shared program could underlie the molecular overlap seen in bulk tumour profiles. Quantifying the Group 3/4-B and C programs in bulk MB gene expression data3 (n = 248 Group 3/4 MBs) recapitulated observations made in our single-cell cohort (Fig. 5a). Sorting these profiles by their relative scores for these programs confirmed that prototypic Group 3 MBs were largely characterized by the undifferentiated progenitor-like program (Group 3/4-B), whereas prototypic Group 4 MBs were dominated by the differentiated neuronal-like program (Group 3/4-C). A considerable fraction of tumours (19.8%) exhibited evidence of both programs (Fig. 5a, Extended Data Fig. 9a). These intermediate tumours were characterized by elevated DNA methylation-based prediction scores (≥0.2) for both subgroups (odds ratio = 8.9, P < 0.001, Fisher’s exact test). We validated these results by performing immunohistochemistry on a series of 22 Group 3/4 MBs, using MYC and TUJ1 (which is encoded by TUBB3) as biomarkers of the Group 3/4-B and Group 3/4-C programs, respectively (Fig. 5b, Extended Data Fig. 9b). Prototypical Group 3 MBs exhibited high expression of MYC and few TUJ1-positive cells, whereas prototypical Group 4 MBs were devoid of MYC-expressing cells and universally positive for TUJ1. Tumours classified as intermediate Group 3/4 MB by DNA methylation contained varying proportions of both MYC-expressing and TUJ1-expressing cells, consistent with our single-cell results.
a, Relative expression of 60 genes representing Group 3/4 MB metaprograms across bulk Group 3/4 MBs. DNA methylation-based subgroup prediction scores are indicated (bottom). b, DNA methylation-based subgroup prediction scores (top) and relative immunohistochemistry-based expression levels of MYC and TUJ1 (encoded by TUBB3; middle) in 22 Group 3/4 MBs. Four representative immunohistochemistry images are shown (bottom). c, t-SNE representation of bulk Group 3/4 MBs classified according to published DNA methylation subtypes3,32 (n = 740; left). Subtypes are delineated by curved lines. Samples are coloured by differentiation state as defined in a. Right, quantification of undifferentiated, intermediate or differentiated expression state per subtype. Asterisks indicate enrichment of intermediate samples (Fisher’s exact test, P < 0.001). Scale bars, 25 μm. Undiff., undifferentiated; Diff., differentiated; Interm., intermediate.
We next investigated whether recently described3,32 DNA methylation-based subtypes of Group 3/4 MB were related to the metaprograms inferred from scRNA-seq. We found that DNA methylation subtypes I and V, both of which contain a mixture of Group 3 and Group 4 MBs, were significantly enriched for tumours with intermediate expression patterns (P < 0.001, Fisher’s exact test; Fig. 5c, Extended Data Fig. 9c). These results suggest that a continuum of cellular states accounts for the molecular substructure seen in Group 3/4 that complicates accurate consensus classification.
Lineage-specific correlates of Group 4 MB
We next sought to compare and interrelate the different subgroup-specific metaprograms. To this effect, we applied all observed metaprograms (n = 10) to all 7,745 malignant cells in our dataset. Pairwise correlation of expression scores confirmed high similarity among cell cycle programs (WNT-A, SHH-A, Group 3/4-A; average r = 0.99) (Fig. 6a). The undifferentiated progenitor-like programs (WNT-B, SHH-B and Group 3/4-B) exhibited low correlations (average r = 0.23), in agreement with their distinct underlying biology. By contrast, the neuronal-like differentiation programs (WNT-C, SHH-C and Group 3/4-C) were highly correlated (average r = 0.77; Fig. 6a, b, Extended Data Fig. 9d), consistent with shared capacity for neuronal differentiation across subgroups. We reasoned that the neuronal-like differentiation programs defined in each subgroup consist of general neuronal differentiation markers, potentially masking markers of specific lineages. To elucidate markers that might inform developmental origins, we compared genes specific to neuronal-like cells in the different subgroups (n = 260; relative to undifferentiated cell populations; see Methods). Half of these genes (52%) were shared between at least two subgroups and included general markers of neuronal differentiation (for example, ENO2, SYT11, TUBB3 and MAP2), while the remainder were exclusive to individual subgroups (13–20%; Fig. 6c, Extended Data Fig. 9e, Supplementary Table 3). Glutamatergic lineage-specific transcription factors EOMES and LMX1A ranked among the most-differentially expressed genes specific to the Group 3/4-C program (Fig. 6c, Supplementary Table 3). In mice, these transcription factors have essential roles in defining neuronal cell fates in the embryonic upper rhombic lip (uRL), including UBCs and GluCN, both of which are born out of the uRL during cerebellar morphogenesis33,34. As our earlier CCA analysis identified both UBCs and GluCN as being highly correlated with Group 4 MB expression datasets (Fig. 1d, Extended Data Fig. 4f), we performed a deeper analysis into these correlations. Discriminatory UBC markers were specifically expressed in Group 4 single cells and bulk tumour profiles, implicating a possible developmental link between UBCs and Group 4 MB (Fig. 6d, e, Extended Data Fig. 9f). Similar results were observed for GluCN, although the highest correlations were limited to a subset of Group 4 tumours (Extended Data Fig. 10a–f). Collectively, these associations further implicate UBCs and GluCN of the embryonic cerebellum as candidate cells-of-origin for Group 4 MB.
a, Pairwise correlation of expression scores of metaprograms defined separately in each subgroup and applied across cells from all subgroups. b, Expression scores for neuronal differentiation metaprograms across all cells (n = 7,745). c, Relative expression levels of genes specific to neuron-like cells in different subgroups (n = 126). Cells are ordered as in b. d, Average expression levels of 30 UBC-associated genes (rows) in identified mouse cerebellar cell types (columns). Genes are ordered by their relative expression in UBCs. e, Relative expression of homologous genes in d in all cells from the MB single-cell cohort (columns). Cells are ordered as in b.
Discussion
Despite extensive characterization of MB genomic landscapes, effective subgroup-specific therapies have yet to emerge, suggesting that a deeper understanding of the biological and cellular basis of MB is essential. This is particularly urgent for Group 3 and Group 4 MB, which often bear inferior outcomes. As a first challenge, these subgroups have proven difficult to accurately classify, confounded by transcriptional and epigenetic ambiguity. Our combined single-cell analysis of Group 3/4 MBs confirmed that prototypic Group 3 MBs are dominated by undifferentiated progenitor-like cells, whereas prototypic Group 4 MBs consist almost exclusively of more differentiated neuronal-like cells. Of note, we identified a subset of intermediate tumours characterized by varying proportions of both undifferentiated and more differentiated populations (Extended Data Fig. 10g). These findings offer a novel molecular and cellular explanation for the challenges associated with Group 3 and Group 4 sub-classification and provide a framework for future classifications that incorporate population heterogeneity.
Cellular origins for WNT and SHH MB have been mostly informed from genetically faithful mouse models8,9. Cross-species transcriptional analyses performed here confirmed significant correlations between SHH MB and GNPs of variable differentiation states that were associated with patient age. Moreover, our analyses identified UBCs and GluCN as cellular correlates of Group 4 MB subtypes, building on previous studies that have implicated glutamatergic cellular origins for Group 44. For WNT MB, we failed to identify significant correlation between malignant single-cell programs and cerebellar populations, consistent with an extracerebellar origin for this subgroup8. No significant correlations were detected between Group 3 MB and our cerebellar dataset. This observation may be attributed to transformation and cellular reprogramming driven by specific oncogenes (that is, MYC) or may imply that Group 3 MBs have an extracerebellar origin. It is also plausible that our mouse reference atlas was incomplete and lacked populations pertinent to either WNT or Group 3 MB origins. Technical limitations of comparing single-cell datasets between species should not be underestimated, warranting future studies of the cellular correlates between human cerebellar and MB single cells.
In conclusion, our work provides a cellular atlas of MB across all subgroups and a cross-species comparison of cerebellar development, highlighting putative subgroup-specific origins. Our analyses also define the cellular states underlying each MB subgroup, disentangling determinants of intra- and intertumoral heterogeneity. These findings will enable future studies to assess translational opportunities and to evaluate the impact of therapeutic approaches on the spectrum of cellular states that drive MB.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Tissue sample collection and dissection
Human primary tumours
Patients and their parents at Boston Children’s Hospital, the Medical University of Vienna and St Jude Children’s Research Hospital gave consent preoperatively according to Institutional Review Board guidelines. Fresh tumours were collected at the time of surgery and processed directly. Tumour samples from Boston Children’s Hospital and the Medical University of Vienna were mechanically and enzymatically dissociated using a papain-based brain tumour dissociation kit (Miltenyi Biotec). Tumour samples from St Jude Children’s Research Hospital were pre-cut and dissociated for 30 min at 37 °C in papain solution (10 units/ml, Worthington, LS003126) containing N-acetyl-l-cysteine (160 μg/ml, Sigma-Aldrich, A9165) and DNase I (12 μg/ml, Sigma-Aldrich, DN25), rinsed in Neurobasal medium (Gibco, 21103049) supplemented with B-27 (Gibco, 17504044), N-2 (Gibco, 17502048) and l-glutamine (Gibco, 10378016), and filtered using a 40-µm strainer.
Mouse cerebellum
Mouse cerebellar tissue from Crl:CD1 (ICR) mice at 13 distinct developmental time points was previously isolated21. Embryonic time points include each day between E10 to E18 and postnatal time points include P0, P4, P7, and P10. Two biological replicates were included at each time point and three at E14. Cerebella were dissociated as previously described.
PDXs
PDXs were acquired from R. Wechsler-Reya (Sanford Burnham Prebys Medical Discovery Institute), X.-N. Li (Baylor College of Medicine) and the Brain Tumour Resource Laboratory (https://www.btrl.org). PDXs were injected into the cerebellum of NSG mice. Mice were observed daily and were euthanized when signs of sickness, including lethargy and neurological abnormalities, appeared. Low passage PDXs (<10) were dissected, pre-cut and dissociated for 30 min at 37 °C in papain solution (10 units/ml, Worthington, LS003126) containing N-acetyl-l-cysteine (160 μg/ml, Sigma-Aldrich, A9165) and DNase I (12 μg/ml, Sigma-Aldrich, DN25), rinsed in Neurobasal medium and filtered using a 40-µm strainer. The experiments were conducted in accordance with the National Institute of Health’s Guide for the Care and Use of Laboratory Animals and according to the guidelines established by the St Jude Children’s Research Hospital Institutional Animal Care and Use Committee. Procedures in the protocol were approved by the Animal Care and Use Committee (ACUC) of SJCRH (Animal Assurance Number: A3077-01).
Fluorescence-activated cell sorting
Dissociated tumour cells (from fresh primary tumours and PDXs) were resuspended in cold 1% bovine serum albumin in phosphate buffered saline (PBS-BSA 1%). Cells were first stained with CD45–Vioblue direct antibody conjugate (Miltenyi Biotec, 130-092-880) in PBS-BSA 1% for 20 min at 4 °C, washed and then co-stained with 1 µM calcein AM (Life Technologies, C3100MP) and 0.33 µM TO-PRO-3 iodide (Life Technologies, T3605) in PBS-BSA 1%. Sorting was performed with FACSAria Fusion (Becton Dickinson) using 488 nm (calcein AM, 530/30 filter), 640 nm (TO-PRO-3, 670/30 filter) and 405 nm (Vioblue, 450/50 filter) lasers. Non-stained controls were included with all tumours. CD45-positive cells were counterselected for the St Jude samples only and viable medulloblastoma cells were identified by staining positive with calcein AM but negative for TO-PRO-3. Forward scatter area (FSC-A) versus side scatter width (SSC-W) criteria were used to discriminate doublets and select single cells. Single cells were sorted into 96-well plates containing cold TCL buffer (Qiagen, 1031576) containing 1% β-mercaptoethanol, snap frozen on dry ice, and then stored at −80 °C before whole-transcriptome amplification, library preparation and sequencing.
Generation and processing of DNA methylation data
All single-cell patient and PDX samples were analysed using Illumina Infinium Methylation EPIC BeadChip arrays according to the manufacturer’s instructions. Data were generated from both freshly frozen and formalin-fixed paraffin-embedded (FFPE) tissue samples. Medulloblastoma subgroup predictions were obtained from a web-platform for DNA methylation-based classification of central nervous system tumours (www.molecularneuropathology.org, version 11b435). Resulting assignment of samples to WNT, SHH, Group 3 and Group 4 subgroups were used for all downstream analyses. A similar classification system was used for predicting medulloblastoma subtypes32. CNV analysis from EPIC methylation array data was performed using the conumee Bioconductor package. Identified CNVs were compared to those predicted from the single-cell data (shown in Extended Data Fig. 2).
Generation of whole-exome and whole-genome sequencing data
Human genomic whole-exome sequencing libraries were generated using the SureSelectXT kit specific for the Illumina HiSeq instrument (Agilent Technologies), followed by exome enrichment using the SureSelectXT Human All Exon V5 without UTRs bait set. The resulting exome-enriched libraries of tumour and normal samples were then sequenced using paired-end 100-cycle sequencing on a NovaSeq 6000 (Illumina) according to the manufacturer’s instructions. Whole-genome sequencing libraries were constructed using the TruSeq DNA PCR-free sample preparation kit according to the manufacturer’s instructions. Tumour and normal samples were sequenced on an Illumina HiSeq 2500 instrument as previously described36. Somatic SNVs and INDELs were determined via the Mutect2 algorithm as implemented in GATK v.4.0. Coding and splice-related variants were subsequently annotated using the Medal Ceremony annotation pipeline. Additionally, all reported somatic variants were manually curated in IGV.
Human scRNA-seq data generation and processing
Whole-transcriptome amplification, library construction and sequencing were performed as previously described following the Smart-seq2 modified protocol13. Expression levels were quantified as Ei,j = log2(TPMi,j/10 + 1), where TPMi,j refers to transcript-per-million for gene i in sample j, as calculated by RSEM37. TPM values were divided by 10 as we estimated that the complexity of single-cell libraries was in the order of 100,000 transcripts and would like to avoid counting each transcript ~10 times, as would be the case with TPM, which may inflate the difference between the expression level of a gene in cells in which the gene is detected and those in which it is not detected.
To filter out low-quality cells, we first removed cells for which less than 2,500 genes were detected. For each processed 96-well plate, we then determined the average number of genes detected per cell minus two times its standard deviation. We then additionally filtered out the cells that were below that threshold. For the remaining cells, we calculated the aggregate expression of each gene as Ea(i) = log2(average(TPMi,1...n) + 1), and excluded genes with Ea<4. In each subgroup (WNT, SHH, and Group 3/4), we defined relative expression by centering the expression levels, Eri,j = Ei,j − average[Ei,1...n] for the remaining cells and genes. On average, we detected ~4,500 genes per cell. Gene expression values were uploaded to the Gene Expression Omnibus (accession number GSE119926).
Pearson correlation coefficients between expression profiles of cells that passed quality filtering was calculated using centred gene expression levels (for each subgroup separately, shown in Figs. 2a, 3a, 4a). Cells were ordered by hierarchical clustering using 1 − correlation coefficient as the distance and Ward’s linkage, within each sample or genetic subclone (for samples SJ99 and BCH825).
Identification of CNVs in single-cell data
CNVs were estimated as previously described13 by applying a moving average to the relative expression, with a sliding window of 100 genes within each chromosome after sorting genes by their chromosomal location (shown in Extended Data Fig. 2). Non-malignant tumour cells were determined by unsupervised clustering of all single-cell-derived copy-number profiles for each sample with 190 copy-number profiles derived from two non-malignant cell types (tumour-associated oligodendrocytes and immune cells14). For the majority of tumours (21/25), most of the cells did not cluster with the non-malignant cells but formed their own cluster(s) and showed clear evidence of CNVs. A small fraction of tumour cells clustered with the non-malignant cells (<4%). Given the high percentage of malignant cells in these tumours, we decided to classify all cells from the remaining four tumours (MUV41, SJ577, MUV34, and SJ625) as malignant. For two samples (SJ99 and BCH825) genetic subclones were identified based on their CNV profiles.
Identification of SNVs in single-cell data
To detect mutant transcripts in our full-length scRNA-seq expression data (shown in Fig. 2c and Extended Data Fig. 3c–f), sequencing reads were first aligned to the human genome build hg19 using STAR v.2.5.1b. RefSeq gene annotations were supplied to guide alignment. Variants were then quantified in each single cell at the genomic position at which they were detected in the whole-genome/whole-exome sequencing data using samtools mpileup v.1.3. For some genes multiple variants were detected (for example, four different variants were detected for CTNNB1 in WNT MB) and quantified separately. To detect mutant and wild-type transcripts, we required one or more supporting reads. We then filtered variants that were detected as mutant in less than three cells, or that were considered erroneously called as they were detected at elevated frequency in samples in which they were not detected in the genome sequencing data. A total of 82 variants remained after this filtering step. Mutations in highly expressed transcripts were detected in the majority of cells from the respective sample (for example, OTX2 Q103R mutation in MUV39). Mutations in less highly expressed transcripts were detected less frequently. Only a small number of mutant transcripts were detected in samples in which they were not detected by genome sequencing (for example, only ten mutant transcripts were detected for the respective other CTNNB1 variants in the WNT MB single cells; Fig. 2c), illustrating the high specificity of our approach.
Identification of intratumour NMF programs and cellular hierarchies
Transcriptional programs were determined as previously described14 by applying NMF to the centred expression data38. Negative values were converted to zero. Analysis was performed for each sample and subclone individually (excluding samples for which less than 100 cells were profiled), using only the malignant cells and setting the number of factors to four for WNT and three for SHH and Group 3/4 tumours. For each of the resulting factors, we considered the 30 genes with the highest NMF scores to be characteristic of that factor (provided in Supplementary Table 2a). All single cells within the WNT, SHH, or Group 3/4 subgroups where then scored for these NMF programs (as described below, shown in Extended Data Figs. 5a, 6a, 8a). Hierarchical clustering, with one minus Pearson correlation as the distance metric and Ward’s linkage, of the scores for each program revealed four (WNT subgroup) or three (SHH and Group 3/4 subgroups) main correlated sets of programs. The 30 genes with the highest average NMF score within each correlated program set (excluding ribosomal protein genes) were then used to define a total of ten subgroup-specific metaprograms (provided in Supplementary Table 2b).
To interpret the characteristics of each metaprogram, we manually inspected their underlying gene signatures. Additionally, we tested for enrichment of described gene sets (GO biological processes cell cycle and neuron differentiation, KEGG hedgehog signalling pathway, and manually curated ribosomal proteins and translational initiation–elongation factors) in each metaprogram using Fisher’s exact test.
Generation of single-cell program expression scores
Single-cell expression scores were generated in a similar way as described previously13. Given a set of genes (Gj) reflecting a NMF program or metaprogram, we calculate for each cell i, a score, SCj(i), quantifying the relative expression of Gj in cell i, as the average relative expression (Er) of the genes in Gj, compared to the average relative expression of a control gene set \({G}_{j}^{{\rm{cont}}}\): \({{\rm{SC}}}_{j}(i)=\mathrm{average}[Er({G}_{j},i)]-\mathrm{average}[Er({G}_{j}^{{\rm{cont}}},i)]\). The control gene set contains 100 genes with the most similar aggregate expression level. In this way, the control gene set has a comparable distribution of expression levels to that of the considered gene set, and the control gene set is 100-fold larger, such that its average expression is analogous to averaging over 100 randomly selected gene sets of the same size as the considered gene set.
Single cells were assigned to different cell populations based on the maximum expression score for their respective subgroup-specific metaprograms, excluding the cycling programs. The fraction of cells per tumour sample assigned to each cell population is provided in Supplementary Table 2d. Scores for the cycling programs were binarized into cycling and non-cycling (larger and smaller than 1, respectively). For illustration of the cellular hierarchies in SHH MB, scores were normalized by minimizing the average minimum difference of all cells per sample to −1 or 1 (shown in Fig. 3c). For the pan-subgroup analysis of all malignant medulloblastoma cells we re-centred expression values across the dataset and calculated expression scores for each of the ten metaprograms. The pairwise correlation of expression scores is shown in Fig. 6a.
Determination of cell population-specific genes
For comparison of cell populations in WNT tumours, we calculated the average expression level of all cells per population (log2-transformed, un-centred expression levels were used). For this analysis we excluded cells from BCH807, as it was very different from the other four WNT MBs and represents an atypical case (without cells scoring highest for the WNT-B metaprogram, highly proliferative, negative staining for nuclear β-catenin and lacking monosomy of chromosome 6). We then determined all genes with a difference smaller or larger than 1 between the average log2-transformed expression levels when comparing the undifferentiated proliferative population (highest for metaprogram WNT-B) against the neuron-like population (WNT-C) or undifferentiated post-mitotic population (WNT-D). A total of 640 genes were identified in this way (provided in Supplementary Table 2c, Extended Data Fig. 5c).
For comparison of neuron-like cell populations between medulloblastoma subgroups, we first determined genes that were specific to any of the neuron-like populations. For every subgroup (WNT, SHH and Group 3/4), the average expression level of all neuron-like cells was compared to the average expression level of all undifferentiated cells from each subgroup, determining genes with a difference larger than 1 between the average log2-tranformed expression levels. This way we determined a total of 260 genes that were specific to the neuron-like cell population of at least one subgroup (provided in Supplementary Table 3). Genes that were specific to two or three subgroups were grouped as shared genes (Fig. 6c, Extended Data Fig. 9e).
RNA in situ hybridization
Paraffin-embedded tissue sections from two WNT MB tumours of the single-cell cohort (SJ99 and SJ129) were obtained from St Jude Children’s Research Hospital. Sections were mounted on glass slides and stored at −80 °C. Slides were stained using the RNAscope 2.5 HD Duplex Detection Kit (Advanced Cell Diagnostics (ACD), 322430). Slides were baked for 1 h at 60 °C, deparaffinized and dehydrated with xylene and ethanol. The tissue was pretreated with RNAscope Hydrogen Peroxide (ACD, 322335) for 10 min at room temperature and RNAscope Target Retrieval Reagent (ACD, 322000) for 15 min at 98 °C. RNAscope Protease Plus (ACD, 322331) was then applied to the tissue for 30 min at 40 °C. Hybridization probes were prepared by diluting the C2 probe (red) 1:50 into the C1 probe (green). Advanced Cell Technologies RNAscope Target Probes used included Hs-MKI67 (ACD, 591771 and 591771-C2), Hs-DKK2 (ACD, 531131-C2), Hs-STMN2 (ACD, 525211-C2), Hs-ZFP36 (ACD, 427351) and Hs-EGR1 (ACD, 457671). Probes were added to the tissue and hybridized for 2 h at 40 °C. A series of ten amplification steps were performed using instructions and reagents provided in the RNAscope 2.5 HD Duplex Detection Kit. Tissue was counterstained with Gill’s haematoxylin for 25 s at room temperature followed by mounting with VectaMount mounting media (Vector Laboratories).
Immunohistochemistry
Double labelling immunohistochemistry was performed using a 1:8,000 dilution of anti-tubulin β3 (clone TUJ1, Biolegend) and 1:25 dilution of anti-MYC (clone Y69, Abcam) diluted in Ventana antibody diluent (Roche Tissue Diagnostic, 251-018) and detected using the UltraView Red (Roche Tissue Diagnostics, 760-501) and UltraView DAB (Roche Tissue Diagnostics, 760-500) detection kits, respectively. Each target was evaluated using a semiquantitative system to construct a H-score, obtained by multiplying the intensity of the stain (0: no staining; 1: weak staining; 2: moderate staining, and 3: strong staining) by the percentage (0 to 100) of cells showing that staining intensity (H-score range, 0 to 300).
Mouse scRNA-seq data generation and processing
Single cells from developing mouse cerebellar tissue were processed using the microfluidics-based 10x Chromium protocol, as previously described21. In brief, single cells were prepared using the Chromium v.1 Single Cell 3′ Library and Gel Bead Kit according to the manufacturers’ specifications. Quantification and quality checks for the library were performed using an Agilent Technologies DNA 1000 chip. Libraries were sequenced on an Illumina HiSeq 2500 machine. Raw sequencing data have been uploaded to the European Nucleotide Archive (accession PRJEB23051).
Mouse developing cerebellum cells were filtered and normalized using the scanpy Python package39. Genes expressed in less than 50 cells and cells expressing less than 200 genes were removed. Additionally, cells with less than 524 and greater than 3,206 total counts (±3 median absolute deviations) were removed. Furthermore, those cells with greater than 5% of their total counts mapping to mitochondrial genes were removed. Gene expression values were then divided by the total number of transcripts and multiplied by 10,000. Normalized values were calculated by natural-log transforming these values. We calculated scaled expression (z scores for each gene) for downstream analysis.
Identification of cell types in developing mouse cerebellum
The scanpy package implemented in Python was applied to identify cell types among 82,228 cells expressing a total of 16,475 genes. After two rounds of clustering (using the Louvain method), populations predicted to be of non-cerebellar origins were excluded. Removed populations were enriched for haemoglobin, oligodendrocycte, and/or immune associated genes. The remaining 78,156 cells were visualized by t-SNE (using the first 100 principle components as input) and clustered a third time. We then merged clusters if the Mantel Spearman correlation between gene distance matrices (using Manhattan distance) was greater than 0.9 (Fig. 1c, Extended Data Fig. 4c). Resulting clusters, in conjunction with marker genes, were used to identify major cell types in the developing cerebellum.
Integrated analysis of mouse and human datasets
Gene expression matrices for human and mouse datasets were restricted to the 16,919 high-confidence homologous genes with gene order conservation and whole-genome alignment scores greater than 75%, as defined by Ensembl. We removed genes without expression in at least 200 cells and filtered out those with gene dispersion across cells/samples less than equal to zero in each dataset. We also regressed out individual-specific effects in the single-cell data.
For CCA, the first 30 canonical correlation vectors were calculated to project each expression matrix into the maximally correlated subspace, as similarly described previously40. In brief, CCA is implemented as singular value decomposition, by implicitly restarted Lanczos bidiagonalization algorithm, of a distance matrix between two gene expression matrices.
We adopted a correlation of differential expression approach to measure similarity between biological groups in two different studies. Such a procedure has previously been shown to be effective in implicating cellular origins for WNT and SHH medulloblastoma subgroups41. Gene expression for each cell, or centroids for each cluster when at the cluster level, is subtracted by mean gene expression of all other cluster centroids to determine differential expression. Cosine distance is then used to calculate correlations between differential expression vectors between studies as a metric for similarity (Figs. 1d, 3d, e, Extended Data Figs. 4f, 7j, 10a, f). Significance is assessed by 10,000 permutations, followed by FDR correction, for cluster labels of interest. Genes driving CCA differential correlations between human and mouse datasets were investigated by identifying genes both differentially expressed in the cell type of interest (Mann–Whitney U-test) and correlated with the CCA correlation (Pearson correlation). Significant genes were those predicted to drive CCA differential correlations.
NMF applied to the centred mouse expression data, with negative values assigned to zero and rank set to two, determined an undifferentiated and differentiated program. Both programs were projected onto a centred dataset of interest, scaled to a range of zero and one, then differentiated programs were subtracted from undifferentiated programs to calculate differentiation scores42 (shown in Extended Data Fig. 4b).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The scRNA-seq and array-based DNA methylation data of 36 patient and PDX samples described in this study have been deposited in the Gene Expression Omnibus (GEO) with the accession code GSE119926. The scRNA-seq data of the developing mouse cerebellum have been deposited to the European Nucleotide Archive (ENA) with the accession code PRJEB23051.
References
Gajjar, A. J. & Robinson, G. W. Medulloblastoma-translating discoveries from the bench to the bedside. Nat. Rev. Clin. Oncol. 11, 714–722 (2014).
Taylor, M. D. et al. Molecular subgroups of medulloblastoma: the current consensus. Acta Neuropathol. 123, 465–472 (2012).
Northcott, P. A. et al. The whole-genome landscape of medulloblastoma subtypes. Nature 547, 311–317 (2017).
Lin, C. Y. et al. Active medulloblastoma enhancers reveal subgroup-specific cellular origins. Nature 530, 57–62 (2016).
Hovestadt, V. et al. Decoding the regulatory landscape of medulloblastoma using DNA methylation sequencing. Nature 510, 537–541 (2014).
Northcott, P. A. et al. Subgroup-specific structural variation across 1,000 medulloblastoma genomes. Nature 488, 49–56 (2012).
Northcott, P. A. et al. Medulloblastomics: the end of the beginning. Nat. Rev. Cancer 12, 818–834 (2012).
Gibson, P. et al. Subtypes of medulloblastoma have distinct developmental origins. Nature 468, 1095–1099 (2010).
Yang, Z. J. et al. Medulloblastoma can be initiated by deletion of Patched in lineage-restricted progenitors or stem cells. Cancer Cell 14, 135–145 (2008).
Oliver, T. G. et al. Loss of patched and disruption of granule cell development in a pre-neoplastic stage of medulloblastoma. Development 132, 2425–2439 (2005).
Cho, Y. J. et al. Integrative genomic analysis of medulloblastoma identifies a molecular subgroup that drives poor clinical outcome. J. Clin. Oncol. 29, 1424–1430 (2011).
Tanay, A. & Regev, A. Scaling single-cell genomics from phenomenology to mechanism. Nature 541, 331–338 (2017).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Filbin, M. G. et al. Developmental and oncogenic programs in H3K27M gliomas dissected by single-cell RNA-seq. Science 360, 331–335 (2018).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Venteicher, A. S. et al. Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq. Science 355, eaai8478 (2017).
Hovestadt, V. et al. Robust molecular subgrouping and copy-number profiling of medulloblastoma from small amounts of archival tumour material using high-density DNA methylation arrays. Acta Neuropathol. 125, 913–916 (2013).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protocols 9, 171–181 (2014).
Northcott, P. A. et al. Rapid, reliable, and reproducible molecular sub-grouping of clinical medulloblastoma samples. Acta Neuropathol. 123, 615–626 (2012).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Carter, R. A. et al. A single-cell transcriptional atlas of the developing murine cerebellum. Curr. Biol. 28, 2910–2920 (2018).
Kool, M. et al. Molecular subgroups of medulloblastoma: an international meta-analysis of transcriptome, genetic aberrations, and clinical data of WNT, SHH, Group 3, and Group 4 medulloblastomas. Acta Neuropathol. 123, 473–484 (2012).
Waszak, S. M. et al. Spectrum and prevalence of genetic predisposition in medulloblastoma: a retrospective genetic study and prospective validation in a clinical trial cohort. Lancet Oncol. 19, 785–798 (2018).
Shih, D. J. et al. Cytogenetic prognostication within medulloblastoma subgroups. J. Clin. Oncol. 32, 886–896 (2014).
Northcott, P. A. et al. Pediatric and adult sonic hedgehog medulloblastomas are clinically and molecularly distinct. Acta Neuropathol. 122, 231–240 (2011).
Kool, M. et al. Genome sequencing of SHH medulloblastoma predicts genotype-related response to smoothened inhibition. Cancer Cell 25, 393–405 (2014).
Merk, D. J. et al. Opposing effects of CREBBP mutations govern the phenotype of Rubinstein–Taybi syndrome and adult SHH medulloblastoma. Dev. Cell 44, 709–724 (2018).
Northcott, P. A. et al. Medulloblastoma comprises four distinct molecular variants. J. Clin. Oncol. 29, 1408–1414 (2011).
Jones, D. T. et al. Dissecting the genomic complexity underlying medulloblastoma. Nature 488, 100–105 (2012).
Cavalli, F. M. G. et al. Intertumoral heterogeneity within medulloblastoma subgroups Cancer Cell 31, 737–754 (2017).
Schwalbe, E. C. et al. Novel molecular subgroups for clinical classification and outcome prediction in childhood medulloblastoma: a cohort study. Lancet Oncol. 18, 958–971 (2017).
Sharma, T. et al. Second-generation molecular subgrouping of medulloblastoma: an international meta-analysis of Group 3 and Group 4 subtypes. Acta Neuropathol. https://doi.org/10.1007/s00401-019-02020-0 (2019).
Chizhikov, V. V. et al. Lmx1a regulates fates and location of cells originating from the cerebellar rhombic lip and telencephalic cortical hem. Proc. Natl Acad. Sci. USA 107, 10725–10730 (2010).
Englund, C. et al. Unipolar brush cells of the cerebellum are produced in the rhombic lip and migrate through developing white matter. J. Neurosci. 26, 9184–9195 (2006).
Capper, D. et al. DNA methylation-based classification of central nervous system tumours. Nature 555, 469–474 (2018).
Rusch, M. et al. Clinical cancer genomic profiling by three-platform sequencing of whole genome, whole exome and transcriptome. Nat. Commun. 9, 3962 (2018).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome BMC Bioinformatics 12, 323 (2011).
Gaujoux, R. & Seoighe, C. A flexible R package for nonnegative matrix factorization. BMC Bioinformatics 11, 367 (2010).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Pöschl, J. et al. Genomic and transcriptomic analyses match medulloblastoma mouse models to their human counterparts. Acta Neuropathol. 128, 123–136 (2014).
Tamayo, P. et al. Metagene projection for cross-platform, cross-species characterization of global transcriptional states. Proc. Natl Acad. Sci. USA 104, 5959–5964 (2007).
Acknowledgements
P.A.N. is a Pew–Stewart Scholar for Cancer Research (Margaret and Alexander Stewart Trust) and recipient of The Sontag Foundation Distinguished Scientist Award. P.A.N. was also supported by the National Cancer Institute (R01CA232143-01), American Association for Cancer Research (NextGen Grant for Transformative Cancer Research), The Brain Tumour Charity (Quest for Cures), the American Lebanese Syrian Associated Charities (ALSAC), and St Jude. M. L. Suvà was supported by grants from the Howard Goodman Fellowship at MGH, the Merkin Institute Fellowship at the Broad Institute of MIT and Harvard, the Wang Family Fund, the V Foundation for Cancer Research, the Swiss National Science Foundation Sinergia program, and the Alex’s Lemonade Stand Foundation. M. L. Suvà is also recipient of The Sontag Foundation Distinguished Scientist Award. B.E.B. is the Bernard and Mildred Kayden Endowed MGH Research Institute Chair and an American Cancer Society Research Professor. This research was supported by a Pioneer Award from the NIH Common Fund and National Cancer Institute (DP1CA216873). V.H. is supported by a Human Frontier Science Program long-term fellowship (LT000596/2016-L). L.B. is supported by a Future Leaders Award from The Brain Tumour Charity (GN-000518). M.G.F. was supported by a Career Award for Medical Scientist from Burroughs Wellcome Fund, a K12 Paul Calabresi Career Award for Clinical Oncology (K12CA090354), a Harvard Brain Cancer SPORE—Career Enhancement Program Award, the National Institutes of Health (3P30 CA006516-53S6), The Cure Starts Now Foundation, Solving Kids’ Cancer/The Bibi Fund, The Andruzzi Foundation and Alex’s Lemonade Stand Foundation. I.S., D.K. and D.L. were supported by the Austrian National Bank (OeNB Jubiläumsfonds Project 15173). M.N.R. is supported by the ALSF, PBTF, AKBTC and CBJOLF. We are indebted to the Flow Cytometry Core Laboratory (Department of Developmental Neurobiology, St Jude) and the Core Flow Cytometry and Cell Sorting Shared Resource Facility (St Jude). From St Jude, we explicitly acknowledge the Hartwell Center, the Biorepository, members of the Clinical Genomics team, the Diagnostic Biomarkers Shared Resource in the Department of Pathology, and the Center for In Vivo Imaging and Therapeutics. We thank S. Pounds (Department of Biostatistics, St Jude) for valuable discussions and B. Stelter for assistance with artwork.
Author information
Authors and Affiliations
Contributions
Study design: V.H., K.S.S., L.B., M.G.F., B.E.B., M. L. Suvà and P.A.N. Generation of human transcriptome data: L.B., M.G.F., M. L. Shaw, A.B., J.C.D., A. Groves, L.M., H.R.W., A.R.R, M.E.S., J. H., R.A.A., J.G., D.K., D.L., R.G. and A.H. Generation of mouse transcriptome data: L.B., C.R., T.N.P., J.L.H., Y.T. and J.E. Analysis of human transcriptome data: V.H. and K.S.S. Analysis of mouse transcriptome data: V.H., K.S.S., R.A.C. and C.G. Generation and analysis of genome data: V.H., K.S.S., L.B., T.S., D.F., A.S., S.M.P, A. Gajjar and G.W.R. Immunohistochemistry experiments: L.B. and B.A.O. RNA in situ hybridization: H.R.W. and M.E.S. Procurement of patient and PDX samples: L.B., M.G.F., L.G., J.L.H., M.D., K.L.L., J.M.R., R.J.W.-R., X.-N.L., A.P., T.C., C.D., C.H., A. Gajjar, B.A.O., I.S. and G.W.R. Project support: S.L.P., M.N.R., O.R.-R. and A. R. Manuscript preparation (with feedback from all authors): V.H., K.S.S., L.B., M.G.F., B.E.B., M. L. Suvà and P.A.N. Study supervision and funding: M.G.F., B.E.B., M. L. Suvà and P.A.N.
Corresponding authors
Ethics declarations
Competing interests
B.E.B. discloses financial interests in Fulcrum Therapeutics, 1CellBio, HiFiBio, Arsenal Biosciences, Cell Signaling Technologies and Nohla Therapeutics. A.R. is a founder and equity holder of Celsius Therapeutics and an SAB member of ThermoFisher Scientific and Syros Pharmaceuticals.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Xing Fan and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Characteristics of the MB single-cell cohort.
a, Haematoxylin and eosin-stained sections from all St Jude single-cell samples (n = 12). Tumours demonstrated large cell/anaplastic morphology (LCA, top), classic morphology (middle), or desmoplastic/nodular morphology (D/N, bottom). Scale bars, 50 µm. b, Detailed characterization of the PDX single-cell dataset. Subgroup prediction scores35 derived by DNA methylation profiling are indicated in the top panel (light shade, low probability; dark shade, high probability). The heat map shows expression levels of previously described subgroup-specific marker genes19 in 946 PDX-derived single-cells. c, Heat map shows expression levels of previously described subgroup-specific marker genes19 in 7,788 tumour-derived single-cells. d, Heat maps show pairwise correlation of aggregated scRNA-seq data (top) and bulk DNA methylation data (bottom) of all patient (n = 25) and PDX (n = 11) samples. For each PDX sample, the patient sample with the highest correlation coefficient is indicated by a black circle. e, Scatter plots show expression scores for published subgroup-specific gene sets for all single cells in the patient cohort (n = 7,788). Cells from WNT and SHH subgroups score only for their respective gene set. Some overlap is observed between cells from Group 3 and 4 subgroups and their respective gene sets, warranting the combined analysis of these subgroups in this study.
Extended Data Fig. 2 Copy-number analysis distinguishes malignant from non-malignant single cells.
a–e, Heat maps show scRNA-seq-derived copy-number profiles of every cell in each sample (y axis) along the genome (x axis) for WNT (a), SHH (b), Group 3 (c) and Group 4 (d) patient MBs as well as PDX samples (e). Copy-number profiles derived from array-based DNA methylation profiling from the same sample are shown above. CNVs are observed in 21/25 patient tumour samples (all except MUV34, MUV41, SJ577 and SJ625). Generally, we observe a high concordance between single-cell and DNA methylation array-derived copy-number profiles. Genetic subclones at the level of broad copy-number changes are detected in samples SJ99 and BCH825. Cells without detected CNVs from samples that showed CNVs in the majority of cells are indicated for samples in which at least four non-malignant cells were detected (BCH807 and SJ454). Amplifications of the MYC and MYCN oncogenes detected by DNA methylation array are indicated.
Extended Data Fig. 3 Unsupervised clustering and detection of expressed SNVs in MB single-cells.
a, t-SNE visualization of the entire single-cell dataset (n = 8,924 cells). WNT (blue), SHH (red), Group 3 (yellow) and Group 4 (green) patient samples are indicated. PDX models are shown in pink. Non-neoplastic oligodendrocytes and immune cells are included for comparison. Generally malignant cells are expected to cluster by patient sample, whereas non-malignant cells are expected to cluster by cell type. Only few cells from different samples cluster with oligodendrocytes (n = 22) or immune cells (n = 6) and were classified as non-malignant. No additional clusters of cells from different samples were identified, indicating the absence of additional non-malignant cell populations in our dataset. b, Identical t-SNE visualization as in a, coloured by copy-number state. CNVs were detected in most single cells, facilitating their classification as malignant. A small number of cells did not show CNVs, even though CNVs were detected in the majority of cells from the respective sample (n = 38). These cells were classified as non-malignant. Most cells with without CNVs clustered with normal oligodendrocytes (n = 21), supporting their initial classification as non-malignant. Remaining cells without CNVs did not form clusters and likely represent poor-quality cells. c, Identical t-SNE visualization as in a, coloured by detected mutant and wild-type transcripts. Cells classified as non-malignant are depleted for mutant transcripts (P < 0.01, binomial test), supporting their initial classification. d, Heat map shows detected mutant and wild-type transcripts for 39 variants (columns) in each cell (n = 1,780, rows) of the WNT MB dataset. If both mutant and wild-type transcripts are detected in a single cell, only the mutant transcript is shown. Variants were initially detected by genome sequencing and subsequently quantified in the scRNA-seq data. Sample BCH807 was not subjected to genome sequencing, and the CTNNB1 variant was manually detected by examining scRNA-seq alignments. Mutations are detected almost exclusively in single cells from samples in which they were detected by genome sequencing, illustrating the high specificity of single-cell variant detection. e, Heat map shows mutant and wild-type transcripts for 15 variants in each cell (n = 1,135, rows) of the SHH MB dataset. Sample SJ454 was not subjected to genome sequencing, and the TP53 mutation was manually identified by examining scRNA-seq alignments. f, Heat map shows mutant and wild-type transcripts for 28 variants in each cell (n = 3,172, rows) of the Group 3/4 MB samples that were subjected to genome sequencing.
Extended Data Fig. 4 Single-cell mapping of mouse cerebellar development.
a–c, Two-dimensional representation of the cerebellar (CB) scRNA-seq dataset by t-SNE. Each dot represents one cell. In a, colours represent 13 different embryonic and early postnatal time points. In b, colours indicate the differentiation score across the entire dataset. In c, colours indicate cell types identified by Louvain clustering using the top 3,000 overdispersed genes. The main CB lineages were assigned on the basis of published lineage markers. d, Annotation of 18 CB cell types based on the expression of lineage specific marker genes shown as violin plot. Violin plots represent kernel density estimation showing the distribution shape of the data. e, Lineage tree reconstruction using partition-based graph abstraction. The abstracted graph shows all cell types (nodes) as identified in c and d. The size of the nodes is related to the number of cells in the defined cell type. The width of edges connecting cell types reflects the probability of the path. f, Radar plot showing CCA coefficients between each mouse CB cell type and human MB subgroup scRNA-seq.
Extended Data Fig. 5 Characterization of WNT MB single-cell programs.
a, Expression scores for individual programs identified by unsupervised NMF analysis in each sample. Cells are ordered as in Fig. 2a (n = 1,780). Metaprograms WNT-A, WNT-B, WNT-C, and WNT-D were identified by hierarchical clustering of individual programs. b, Heat maps show pairwise correlation (left), principal component analysis (PCA, centre), and expression scores for NMF-derived metaprograms (right) for 301 cells from WNT MB sample MUV44. The ordering of cells (rows) is maintained between the heat maps. A two-dimensional representation of the same cells using t-SNE is shown on the far right (coloured by expression scores for each metaprogram). This analysis shows that the same programs and cell populations that are identified by the NMF analysis are also supported by PCA and t-SNE clustering. Furthermore, no additional programs and cell populations are identified (starting from PC5, components are less informative). c, Scatter plot shows isometric projection of average gene expression levels for cells with highest expression score for WNT-B (undifferentiated, proliferating), WNT-C (neuron-like), or WNT-D (undifferentiated, post-mitotic). WNT-B metaprogram genes are indicated in red, WNT-C metaprogram genes are indicated in green, and WNT-D metaprogram genes are indicated in blue. Genes that are higher in both undifferentiated cell populations compared to neuron-like cells are indicated in black. d, Images show RNA in situ hybridization experiments of five marker genes representative for the four WNT MB metaprograms in two samples of the single-cell cohort. Results confirm expression of these genes independently of the scRNA-seq experiments.
Extended Data Fig. 6 Characterization of SHH MB single-cell programs.
a, Expression scores for individual programs identified by unsupervised NMF analysis in each sample. Cells are ordered as in Fig. 3a (n = 1,135). Metaprograms SHH-A, SHH-B, and SHH-C were identified by hierarchical clustering of individual programs. b, Heat maps show pairwise correlation (left), PCA (centre), and expression scores for NMF-derived metaprograms (right) for 493 cells from SHH MB sample SJ577. The ordering of cells (rows) is maintained between the heat maps. A two-dimensional representation of the same cells using t-SNE is shown on the far right (coloured by expression scores for each metaprogram). This analysis shows that the same programs and cell populations that are identified by the NMF analysis are also supported by PCA and t-SNE clustering. Furthermore, no additional programs and cell populations are identified (starting from PC3, components are less informative). c, Pairwise correlations between the expression profiles of 303 single-cells (rows, columns) from two SHH PDX samples (RCMB18 and RCMB24) (left). Expression scores for each of the NMF-derived metaprograms SHH-A, SHH-B, and SHH-C (columns) (right). Cells are ordered as in the left panel (rows). d, Heat maps show the relative expression of the 60 genes representing the metaprograms SHH-B and SHH-C (rows), across 303 cells for RCMB18 and RCMB24. Cells are sorted by the difference between the two scores. Cells positive for the cell cycle program (SHH-A) are indicated by red bars. Similar cell populations as in the primary samples (undifferentiated GNP-like and differentiated neuron-like cells) are identified in RCMB18. No differentiated cells are identified in RCMB24.
Extended Data Fig. 7 Cross-species mapping of SHH MB origins.
a, Heat map shows average expression levels of 29 GNP-associated genes (rows) in cell types identified in the mouse CB dataset (columns). Genes are ordered by their relative expression in GNPs. b, Left, the relative expression of orthologous genes in a in all cells from the single-cell cohort (n = 7,745; columns). Cells are ordered by increasing GNP CCA cosine correlation coefficients. Cells expressing high levels of GNP-associated genes are predominantly from SHH tumours. Right, the relative expression of the same genes in the bulk microarray cohort (n = 392). c, d, Heat maps as in a and b, but showing 30 genes associated with the UBC/GN intermediate cell type. e, Two-dimensional representation of GNPs/granule neurons from the cerebellar atlas by t-SNE. Each dot represents one cell (n = 35,013). Colours represent the assigned cerebellar cell types (left), as well as the expression of Atoh1 and Neurod1 (middle and right). f, Box plots of select granule lineage marker genes in the mouse CB cohort (left), MB single-cell cohort (middle) and MB bulk microarray cohort (right). g, Box plot of patient age associated with infant and adult/child subtypes of SHH MB. h, Box plot of the number of coding mutations associated with SHH MB subtypes. The median is shown as a thick line; box limits are 25th and 75th percentiles; whiskers denote 1.5 times the interquartile range. i, Expression of Barhl1 (left) and Pde1c (right) at P4 during CB development. In situ hybridization data were obtained from the Allen Developing Mouse Brain Atlas (© 2008 Allen Institute for Brain Science. Allen Developing Mouse Brain Atlas http://developingmouse.brain-map.org). j, Radar plot showing the CCA cosine correlation coefficients between each mouse CB cell type and the MB single-cell cohort from cells scoring highest for metaprograms SHH-B (GNP-like cells) and SHH-C (granule neuron-like cells).
Extended Data Fig. 8 Characterization of Group 3/4 MB single-cell programs.
a, Top, Group 3/4 subtype prediction scores derived by DNA methylation profiling32 (light shade, low probability; dark shade, high probability). Expression scores for individual programs identified by unsupervised NMF analysis in each sample are indicated in the bottom. Cells are ordered as in Fig. 4a (n = 4,873). Metaprograms Group 3/4-A, Group 3/4-B, and Group 3/4-D were identified by hierarchical clustering of individual programs. b, Expression scores across 4,873 single cells (columns) for each of the NMF-derived metaprograms Group 3/4-A, Group 3/4-B, and Group 3/4-C (rows). Cells are ordered as in a. c, Heat maps show pairwise correlation (left), PCA (centre) and expression scores for NMF-derived metaprograms (right) for 400 cells from Group 3 MB sample SJ617. The ordering of cells (rows) is maintained between the heat maps. A two-dimensional representation of the same cells using t-SNE is shown on the far right (coloured by expression scores for each metaprogram). This analysis shows that the same programs and cell populations that are identified by the NMF analysis are also supported by PCA and t-SNE clustering. Furthermore, no additional programs and cell populations are identified (starting from PC4 components are less informative). d, Pairwise correlations between the expression profiles of 643 single cells (rows, columns) from nine patient-derived xenograft models (Med114FH, Med2112FH, Med211FH, Med411FH, RCMB20, Icb1299, Icb1572, Med2312FH and DMB006). Left, Group 3/4 subtype prediction scores derived by DNA methylation profiling. Right, expression score for the NMF-derived metaprograms Group 3/4-A, Group 3/4-B and Group 3/4-C (columns). e, Heat maps show the relative expression of the 60 genes representing the metaprograms Group 3/4-B and Group 3/4-C (rows) across 140 cells for RCMB20 and DMB006. Cells are sorted by the difference between the two scores. Cells positive for the cell cycle program (Group 3/4-A) are indicated by red bars. Group 3 PDX samples are predominantly undifferentiated, with the exception of Med2312FH, which is predominantly differentiated (classified by DNA methylation array as an intermediate Group 3/4 sample). This parallels the high frequency of MYC amplifications in our Group 3 PDX cohort (5/8). Group 4 PDX sample DMB006 is also predominantly differentiated. These results are supportive of the cellular compositions detected in primary Group 3/4 samples.
Extended Data Fig. 9 Analysis of Group 3/4 intermediate samples and pan-subgroup comparison.
a, Scatter plot of the metaprogram Group 3/4-C (x axis) and Group 3/4-B (y axis) expression scores for Group 3 and Group 4 bulk MBs3 (yellow and green dots, respectively; n = 248). Samples that score similarly for both programs are classified as intermediate samples (n = 49) b, Representative MYC and TUJ1 immunohistochemistry images of seven Group 3/4 samples. Four of these samples are shown at higher magnification in Fig. 5b (SJ17, SJ617, SJ625, SJ723). c, Two-dimensional representation of 740 Group 3/4 MB samples analysed by DNA methylation profiling using t-SNE3. Eight subtypes are delineated by curved lines. Samples are coloured by their predicted subgroup35. d, Heat map showing expression of transcripts coding for ribosomal proteins (n = 75, rows). Cells positive for the cell cycle programs, and cells classified as neuron-like cells are indicated on top. Cells are ordered as in Fig. 6b (n = 7,745). e, Heat map showing relative expression levels of genes that are specific to neuron-like cells and are shared between multiple subgroups (n = 134, rows). Cells are ordered as in d. f, Heat map shows the relative expression of UBC-specific genes in Fig. 6d (n = 30; rows) in the bulk expression array cohort (n = 392; columns). Samples are ordered by increasing CCA cosine correlation coefficient.
Extended Data Fig. 10 Cross-species mapping of Group 4 MB origins.
a, Top, expression of TBR1 and EOMES in bulk Group 4 MB expression array data (n = 149). Middle, Group 3/4 DNA methylation-based subtype annotations for each sample. Bottom, CCA scores from comparison of bulk MB expression data and UBCs and GluCN late populations from the cerebellar single-cell dataset. b, t-SNE visualization shows clustering of glutamatergic populations correlated with Group 4 MBs. c, Box plot of CCA cosine correlation coefficients from comparison of bulk MB expression data and UBCs, according to Group 3/4 subtypes. The median is shown as a thick line; box limits are 25th and 75th percentiles; whiskers denote 1.5 times the interquartile range. d, e, Left, in situ hybridization data for Tbr1 (d) and Eomes (e) in the developing mouse cerebellum at the indicated time point. Data were obtained from the Allen Developing Mouse Brain Atlas (© 2008 Allen Institute for Brain Science. Allen Developing Mouse Brain Atlas http://developingmouse.brain-map.org). Right, expression of Tbr1 (d) and Eomes (e) in the mouse single-cell dataset according to the t-SNE structure shown in b. f, Radar plot showing CCA cosine correlation coefficients between each mouse CB cell type and Group 3 MB (top) or Group 4 MB (bottom) cells scoring highest for metaprograms Group 3/4-B or Group 3/4-C. g, Graphical summary of subgroup-specific cellular hierarchies identified in MB.
Supplementary information
Supplementary Table 1
Cohort details.
Supplementary Table 2
Transcriptional programs.
Supplementary Table 3
Comparison of neuronal-like cells between MB subgroups.
Rights and permissions
About this article
Cite this article
Hovestadt, V., Smith, K.S., Bihannic, L. et al. Resolving medulloblastoma cellular architecture by single-cell genomics. Nature 572, 74–79 (2019). https://doi.org/10.1038/s41586-019-1434-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1434-6
This article is cited by
-
miR-124-3p and miR-194-5p regulation of the PI3K/AKT pathway via ROR2 in medulloblastoma progression
Cancer Gene Therapy (2024)
-
Heterogeneity and tumoral origin of medulloblastoma in the single-cell era
Oncogene (2024)
-
Compartments in medulloblastoma with extensive nodularity are connected through differentiation along the granular precursor lineage
Nature Communications (2024)
-
Developmental basis of SHH medulloblastoma heterogeneity
Nature Communications (2024)
-
De novo identification of expressed cancer somatic mutations from single-cell RNA sequencing data
Genome Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.