Abstract
The gut microbiota influences development1,2,3 and homeostasis4,5,6,7 of the mammalian immune system, and is associated with human inflammatory8 and immune diseases9,10 as well as responses to immunotherapy11,12,13,14. Nevertheless, our understanding of how gut bacteria modulate the immune system remains limited, particularly in humans, where the difficulty of direct experimentation makes inference challenging. Here we study hundreds of hospitalized—and closely monitored—patients with cancer receiving haematopoietic cell transplantation as they recover from chemotherapy and stem-cell engraftment. This aggressive treatment causes large shifts in both circulatory immune cell and microbiota populations, enabling the relationships between the two to be studied simultaneously. Analysis of observed daily changes in circulating neutrophil, lymphocyte and monocyte counts and more than 10,000 longitudinal microbiota samples revealed consistent associations between gut bacteria and immune cell dynamics. High-resolution clinical metadata and Bayesian inference allowed us to compare the effects of bacterial genera in relation to those of immunomodulatory medications, revealing a considerable influence of the gut microbiota—together and over time—on systemic immune cell dynamics. Our analysis establishes and quantifies the link between the gut microbiota and the human immune system, with implications for microbiota-driven modulation of immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary Information files. The data used in our study are organized in Excel-compatible comma-separated value files as Supplementary Tables (data-tables.zip). All sequencing data have been made available publicly, and the NCBI SRA accession numbers are listed in the Supplementary Tables. Metadata and processed sequencing data are made available on a public repository via Figshare: meta data, https://doi.org/10.6084/m9.figshare.12016986.v4; samples, https://doi.org/10.6084/m9.figshare.12016983.v4; 16S counts, https://doi.org/10.6084/m9.figshare.12016989.v3; and 16S taxonomy, https://doi.org/10.6084/m9.figshare.12016992.v1.
Code availability
All of the steps of the analyses that were performed in this study are described in detail to allow reproduction of the results. Relevant analysis code is available publicly at https://github.com/jsevo/wbcdynamics_microbiome.
References
Mazmanian, S. K., Liu, C. H., Tzianabos, A. O. & Kasper, D. L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122, 107–118 (2005).
Gomez de Agüero, M. et al. The maternal microbiota drives early postnatal innate immune development. Science 351, 1296–1302 (2016).
Olin, A. et al. Stereotypic immune system development in newborn children. Cell 174, 1277–1292 (2018).
Tan, T. G. et al. Identifying species of symbiont bacteria from the human gut that, alone, can induce intestinal Th17 cells in mice. Proc. Natl Acad. Sci. USA 113, E8141–E8150 (2016).
Deshmukh, H. S. et al. The microbiota regulates neutrophil homeostasis and host resistance to Escherichia coli K1 sepsis in neonatal mice. Nat. Med. 20, 524–530 (2014).
Ivanov, I. I. et al. Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe 4, 337–349 (2008).
Geva-Zatorsky, N. et al. Mining the human gut microbiota for immunomodulatory organisms. Cell 168, 928–943 (2017).
Lloyd-Price, J. et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 569, 655–662 (2019).
Markey, K. A. et al. The microbe-derived short-chain fatty acids butyrate and propionate are associated with protection from chronic GVHD. Blood 136, 130–136 (2020).
Azzouz, D. et al. Lupus nephritis is linked to disease-activity associated expansions and immunity to a gut commensal. Ann. Rheum. Dis. 78, 947–956 (2019).
Routy, B. et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359, 91–97 (2018).
Gopalakrishnan, V. et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359, 97–103 (2018).
Vétizou, M. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084 (2015).
Matson, V. et al. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science 359, 104–108 (2018).
Tanoue, T. et al. A defined commensal consortium elicits CD8 T cells and anti-cancer immunity. Nature 565, 600–605 (2019).
Brandi, G. & Frega, G. Microbiota: overview and implication in immunotherapy-based cancer treatments. Int. J. Mol. Sci. 20, 2699 (2019).
Xin Yu, J., Hubbard-Lucey, V. M. & Tang, J. The global pipeline of cell therapies for cancer. Nat. Rev. Drug Discov. 18, 821–822 (2019).
Morjaria, S. et al. Antibiotic-induced shifts in fecal microbiota density and composition during hematopoietic stem cell transplantation. Infect. Immun. 87, e00206-19 (2019).
Peled, J. U. et al. Microbiota as predictor of mortality in allogeneic hematopoietic-cell transplantation. N. Engl. J. Med. 382, 822–834 (2020).
Taur, Y. et al. Reconstitution of the gut microbiota of antibiotic-treated patients by autologous fecal microbiota transplant. Sci. Transl. Med. 10, eaap9489 (2018).
Staffas, A. et al. Nutritional support from the intestinal microbiota improves hematopoietic reconstitution after bone marrow transplantation in mice. Cell Host Microbe 23, 447–457. (2018).
Savani, B. N. et al. Absolute lymphocyte count on day 30 is a surrogate for robust hematopoietic recovery and strongly predicts outcome after T cell-depleted allogeneic stem cell transplantation. Biol. Blood Marrow Transplant. 13, 1216–1223 (2007).
Scheiermann, C., Frenette, P. S. & Hidalgo, A. Regulation of leucocyte homeostasis in the circulation. Cardiovasc. Res. 107, 340–351 (2015).
Thompson, P. A. et al. Umbilical cord blood graft engineering: challenges and opportunities. Bone Marrow Transplant. 50 (Suppl 2), S55–S62 (2015).
Gabrilove, J. L. et al. Effect of granulocyte colony-stimulating factor on neutropenia and associated morbidity due to chemotherapy for transitional-cell carcinoma of the urothelium. N. Engl. J. Med. 318, 1414–1422 (1988).
Belkaid, Y. & Hand, T. W. Role of the microbiota in immunity and inflammation. Cell 157, 121–141 (2014).
Schirmer, M. et al. Linking the human gut microbiome to inflammatory cytokine production capacity. Cell 167, 1125–1136 (2016).
McLoughlin, K., Schluter, J., Rakoff-Nahoum, S., Smith, A. L. & Foster, K. R. Host selection of microbiota via differential adhesion. Cell Host Microbe 19, 550–559 (2016).
Hooper, L. V., Littman, D. R. & Macpherson, A. J. Interactions between the microbiota and the immune system. Science 336, 1268–1273 (2012).
Palm, N. W. et al. Immunoglobulin A coating identifies colitogenic bacteria in inflammatory bowel disease. Cell 158, 1000–1010 (2014).
Henke, M. T. et al. Ruminococcus gnavus, a member of the human gut microbiome associated with Crohn’s disease, produces an inflammatory polysaccharide. Proc. Natl Acad. Sci. USA 116, 12672–12677 (2019).
Okba, A. M. et al. Neutrophil/lymphocyte ratio and lymphocyte/monocyte ratio in ulcerative colitis as non-invasive biomarkers of disease activity and severity. Auto Immun. Highlights 10, 4 (2019).
Choi, S.-J. et al. High neutrophil-to-lymphocyte ratio predicts short survival duration in amyotrophic lateral sclerosis. Sci. Rep. 10, 428 (2020).
Gao, Y. et al. Neutrophil/lymphocyte ratio is a more sensitive systemic inflammatory response biomarker than platelet/lymphocyte ratio in the prognosis evaluation of unresectable pancreatic cancer. Oncotarget 8, 88835–88844 (2017).
Hergott, C. B. et al. Peptidoglycan from the gut microbiota governs the lifespan of circulating phagocytes at homeostasis. Blood 127, 2460–2471 (2016).
Smith, P. M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013).
Balmer, M. L. et al. Microbiota-derived compounds drive steady-state granulopoiesis via MyD88/TICAM signaling. J. Immunol. 193, 5273–5283 (2014).
Ze, X., Duncan, S. H., Louis, P. & Flint, H. J. Ruminococcus bromii is a keystone species for the degradation of resistant starch in the human colon. ISME J. 6, 1535–1543 (2012).
Foster, K. R., Schluter, J., Coyte, K. Z. & Rakoff-Nahoum, S. The evolution of the host microbiome as an ecosystem on a leash. Nature 548, 43–51 (2017).
Fu, Y.-Y. et al. T cell recruitment to the intestinal stem cell compartment drives immune-mediated intestinal damage after allogeneic transplantation. Immunity 51, 90–103 (2019).
Gerber, G. K. The dynamic microbiome. FEBS Lett. 588, 4131–4139 (2014).
Jobin, C. Precision medicine using microbiota. Science 359, 32–34 (2018).
The Integrative HMP (iHMP) Research Network Consortium. The integrative human microbiome project. Nature 569, 641–648 (2019).
Walter, J., Armet, A. M., Finlay, B. B. & Shanahan, F. Establishing or exaggerating causality for the gut microbiome: lessons from human microbiota-associated rodents. Cell 180, 221–232 (2020).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Murali, A., Bhargava, A. & Wright, E. S. IDTAXA: a novel approach for accurate taxonomic classification of microbiome sequences. Microbiome 6, 140 (2018).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Pinheiro, J. C., Bates, D. M., DebRoy, S. S. & Sarkar, D. nlme: Linear and Nonlinear Mixed Effects Models. R package version 3.1-150 (2013).
Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. B 58, 267–288 (1996).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825 (2011).
Salvatier, J., Wiecki, T. V. & Fonnesbeck, C. Probabilistic programming in Python using PyMC3. PeerJ Comput. Sci. 2, e55 (2016).
Hoffman, M. D. & Gelman, A. The No-U-turn sampler: adaptively setting path lengths in Hamiltonian Monte Carlo. J. Mach. Learn. Res. 15, 1593–1623 (2014).
Franzosa, E. A. et al. Species-level functional profiling of metagenomes and metatranscriptomes. Nat. Methods 15, 962–968 (2018).
Acknowledgements
We thank M. Lipsitch, S. B. Andersen, K. R. Foster, J. K. Sia, E. G. Pamer, K. Coyte, S. Mitschka and the members of the Xavier lab for helpful discussion and comments on the manuscript. This work was supported by the National Institutes of Health (NIH) grants U01 AI124275, R01 AI137269 and U54 CA209975 to JBX, by the MSKCC Cancer Center Core Grant P30 CA008748, the Parker Institute for Cancer Immunotherapy at Memorial Sloan Kettering Cancer Center, the Sawiris Foundation, the Society of Memorial Sloan Kettering Cancer Center, MSKCC Cancer Systems Immunology Pilot Grant and Empire Clinical Research Investigator Program. M.S. received funding from the Burroughs Wellcome Fund Postdoctoral Enrichment Program, the Damon Runyon Physician-Scientist Award, and the Robert Wood Johnson Foundation. T.M.H. is investigator in the Pathogenesis of Infectious Diseases from the Burroughs Wellcome Fund, and funded via an award from Geoffrey Beene Foundation, and NIH RO1 AI093808. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
J.S. and J.B.X. wrote the manuscript. J.S. and J.B.X. designed the analyses with expert help from R.N. J.U.P. and Y.T. contributed to the clinical data preparation, B.P.T. provided the 16S data-processing pipelines, K.A.M., M.S., A.S., S.M., M.F., M.S.P., T.M.H., M.-A.P. and M.R.M.v.d.B. provided clinical context and helped with variable selection, N.J.C., M.L., L.B., A.B. and A.D.S. provided clinical and other data from Duke, A.D. provided the shotgun processing pipelines. E.F., L.A.A. and R.J.W. processed patients’ stool samples, including for 16S sequencing, shotgun metagenomics and qPCR quantification of total 16S rRNA gene. All authors contributed to the writing and interpretation of the results.
Corresponding authors
Ethics declarations
Competing interests
M.R.M.v.d.B. and J.U.P. received financial support from Seres Therapeutics. M.-A.P. has received honoraria from AbbVie, Bellicum, Bristol-Myers Squibb, Incyte, Merck, Novartis, Nektar Therapeutics, and Takeda, research support for clinical trials from Incyte, Kite (Gilead) and Miltenyi Biotec, and serves on data and safety monitoring boards for Servier and Medigene and scientific advisory boards for MolMed and NexImmune.
Additional information
Peer review information Nature thanks Henrik Nielsen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Blood cell counts over time.
a, WBC counts and platelet counts per graft source over the first 100 days post HCT per day relative to HCT from N = 2,235 adult patients (detailed demographics in supplementary Table 1); lines: mean, shaded: ± standard deviations. b, Data exclusion diagram.
Extended Data Fig. 2 FMT increases WBC counts.
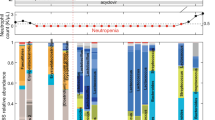
a, HCT patient who received an autologous faecal microbiota transplant (auto-FMT, dashed red line) that restored commensal microbial families and ecological diversity in the gut microbiota, with concurrent cell counts of peripheral neutrophils, lymphocytes and monocytes and immunomodulatory drug administrations. b, Total WBC counts in 24 enrolled patients (10 control, 14 treated) post-neutrophil engraftment; vertical lines indicate randomization dates. c, Weekly mean WBC counts aligned to the randomization date (FMT-treated: red, control: black). Line: mean per week, shaded region: 95% CI. d, Coefficient estimates (mean vs. mean + FMT effect) from linear mixed effects models of total WBC counts over time indicate an auto-FMT-induced increase of WBCs (βFMT: P = 7 × 10−14). e–g, Respectively: neutrophil, lymphocyte and monocyte count trajectories of 24 FMT trial patients. Thin lines: raw data (blue: post-FMT); thick black: mean per day, thick blue: mean+post-FMT coefficient. Means and confidence intervals (shaded region) without (black) and after FMT (blue), as well as the coefficient estimate for FMT treatment and its P value from a linear mixed effects model relating cell counts over time to the FMT treatment (Methods).
Extended Data Fig. 3 Results of the feature selection stage 1 regression.
a–c, Stage 1 regression on neutrophil, lymphocyte, and monocyte dynamics, respectively, on patients without microbiome data. Coefficients from tenfold cross-validated elastic net regression daily changes in neutrophils. gr: intercept; TCD: T cell depleted graft (ex-vivo) by CD34+ selection; PBSC: peripheral blood stem cells; BM: bone marrow; cord: umbilical cord blood; NONABL: Nonmyeloablative; REDUCE: reduced-intensity conditioning regimen; F: female; N: patients, n: samples (daily changes in neutrophils).
Extended Data Fig. 4 Additional coefficients, posterior convergence evaluation and validation.
a–c, Additional posterior coefficient estimates of medications, additional genera and HCT metadata from the Bayesian stage 2 regression, see also Fig. 3. REDUCE: reduced-intensity conditioning regimen; NONABL: non-myeloablative conditioning regimen. F: female. d–f, posterior sampling convergence. Histograms of the ranked posterior draws from the model of neutrophil, lymphocyte and monocyte dynamics, respectively, in PBSC patients (ranked over all chains), plotted separately for each chain show no substantial differences between chains. g–i, Predictors of WBC dynamics using data from patients treated at Duke. Heatmaps indicate the slope coefficients from individual univariate regressions of microbiome and clinical predictors with changes in neutrophils, lymphocytes and monocyte, and for comparison the corresponding coefficients signs from the Bayesian multiple linear regressions in stage 2 of the analysis of WBC dynamics in MSK patients (Fig. 3). Pvalues were adjusted for multiple hypothesis testing using Bonferroni correction: ***P < 0.001, **P < 0.01, *P < 0.05; P > 0.05: n.s. Sign of coefficients from MSK PBSC patients for comparison. j, Equivalent validation analysis from patients treated at Duke using partial least squares regression of microbiome and clinical predictors identified in stage 2 of our analysis on daily changes in neutrophils, lymphocytes and monocyte.
Extended Data Fig. 5 Validation using absolute instead of relative abundance bacterial genus data.
a–d, Validation analysis of the main model using absolute bacterial abundances as predictors instead of relative abundances in Fig. 3. Results show inferred coefficients and P values from multiple linear regressions. One regression per analysed WBC type dynamics, that is, neutrophil, lymphocyte and monocyte daily log-changes, was conducted, and coefficients for medications (a), WBC feedbacks (b) metadata (c) and total genus abundances (d) are shown. This was only possible for only a subset of the data used in the main analysis for which we obtained absolute bacterial abundance estimates (Methods), n: samples, N: patients.
Extended Data Fig. 6 Jointly inferred association network between WBC and bacterial genus dynamics.
Strong regularization yields few non-zero coefficients and antibiotics dominate the dynamics.
Extended Data Fig. 7 Jointly inferred association network between WBC and bacterial genus dynamics with reduced regularization.
Reducing regularization strength (Methods) indicates potential bidirectional feedbacks, for example, between lymphocytes and [Ruminococcus] gnavus group (highlighter green boxes, and cartoon).
Extended Data Fig. 8 Functional analysis of microbiota samples.
To distinguish samples predicted to increase rates of WBCs, a microbiota potency score was calculated from posterior coefficients (Fig. 3, Methods) and the relative abundance of taxa in samples. Bars show linear discriminant analysis (LDA) scores of MetaCyc pathway profiles from 124 shotgun sequenced samples that distinguished positive and negative potency samples the most (LDA-score magnitude in the 95th percentile). Highlighted pathways are discussed in the main text. For each pathway, we tested whether pathway presence was enriched (depleted) in positive (negative) potency samples using one-sided Fisher’s exact test; ***P < 0.001, **P < 0.01, *P < 0.05.
Extended Data Fig. 9 Abundance profiles of bacterial genera across analysed samples.
a, The relative non-zero abundance of Staphylococcus is inversely related to microbiome alpha diversity, bold line: regression line from a linear model of the mean of the log10 Staphylococcus relative abundance, shaded: 95% confidence intervals (n = 1,381 samples with non-zero Staphylococcus abundances). b, Abundance profiles of the two genera, Faecalibacterium and Ruminococcus 2, most strongly associated with WBC increase; number of times detected (left) and log10 abundance distribution when above detection (right).
Extended Data Fig. 10 Survival analysis and confirmation of model results with different priors.
a, Kaplan–Meier plot of patient 3-year survival with sufficient available blood data (Supplementary Information, Extended Data Fig. 1). b, posterior association coefficients do not depend on the choice of prior for σ in the main Bayesian model. Plotted are the posterior means from our main analysis against the equivalent inference with an inverse Gamma prior (alpha = 1, beta = 1).
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-5 and Supplementary Methods.
Supplementary Data
This zip file contains Data Tables.
Rights and permissions
About this article
Cite this article
Schluter, J., Peled, J.U., Taylor, B.P. et al. The gut microbiota is associated with immune cell dynamics in humans. Nature 588, 303–307 (2020). https://doi.org/10.1038/s41586-020-2971-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2971-8
This article is cited by
-
Lactobacillus plantarum L168 improves hyperoxia-induced pulmonary inflammation and hypoalveolarization in a rat model of bronchopulmonary dysplasia
npj Biofilms and Microbiomes (2024)
-
Microbiota–gut–brain axis and its therapeutic applications in neurodegenerative diseases
Signal Transduction and Targeted Therapy (2024)
-
Accelerated intestinal wound healing via dual electrostimulation from a soft and biodegradable electronic bandage
Nature Electronics (2024)
-
Insights into gut microbiomes in stem cell transplantation by comprehensive shotgun long-read sequencing
Scientific Reports (2024)
-
Microplastics dampen the self-renewal of hematopoietic stem cells by disrupting the gut microbiota-hypoxanthine-Wnt axis
Cell Discovery (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.