Abstract
Oestrogen depletion in rodents and humans leads to inactivity, fat accumulation and diabetes1,2, underscoring the conserved metabolic benefits of oestrogen that inevitably decrease with age. In rodents, the preovulatory surge in 17β-oestradiol (E2) temporarily increases energy expenditure to coordinate increased physical activity with peak sexual receptivity. Here we report that a subset of oestrogen-sensitive neurons in the ventrolateral ventromedial hypothalamic nucleus (VMHvl)3,4,5,6,7 projects to arousal centres in the hippocampus and hindbrain, and enables oestrogen to rebalance energy allocation in female mice. Surges in E2 increase melanocortin-4 receptor (MC4R) signalling in these VMHvl neurons by directly recruiting oestrogen receptor-α (ERα) to the Mc4r gene. Sedentary behaviour and obesity in oestrogen-depleted female mice were reversed after chemogenetic stimulation of VMHvl neurons expressing both MC4R and ERα. Similarly, a long-term increase in physical activity is observed after CRISPR-mediated activation of this node. These data extend the effect of MC4R signalling — the most common cause of monogenic human obesity8 — beyond the regulation of food intake and rationalize reported sex differences in melanocortin signalling, including greater disease severity of MC4R insufficiency in women9. This hormone-dependent node illuminates the power of oestrogen during the reproductive cycle in motivating behaviour and maintaining an active lifestyle in women.
Similar content being viewed by others
Main
To establish that female mice rely on VMHvl ERα signalling to maximize their spontaneous physical activity, we ablated ERα in the VMHvl or arcuate nucleus (ARC) of adult Esr1fl/fl female mice using stereotaxic delivery of AAV-Cre-GFP (VMHvlERαKO and ARCERαKO mice). Control female littermates received similarly targeted AAV-GFP (VMHvlcontrol or ARCcontrol mice). Reduced ambulatory activity was observed in VMHvlERαKO female mice during the dark cycle (that is, the active period) that corresponded with a modest increase in body weight, a reduction of Ucp1 expression in interscapular brown adipose tissue (iBAT) and unchanged food intake (Fig. 1a and Extended Data Fig. 1a–e). Although we have shown that ARCERαKO female mice exhibit a surprisingly high bone mass phenotype4, no changes in activity, body weights or food intake were noted in this cohort (Fig. 1a and Extended Data Fig. 1b). Normal food consumption, particularly in ARCERαKO female mice, suggests that the anorexigenic effects of oestrogen are mediated by sites other than the ARC10 or masked by the type of mouse strains and/or institutional housing conditions used here. When considered together with other ERα-knockout mouse models, our data demonstrate a requirement for ERα in the VMHvl to maximize the physical activity levels of adult female mice.
a, Body weight (*P = 0.0310) at 12 weeks after infection, x-ambulatory activity during the dark period (**P = 0.0080) and food intake during the dark period in VMHvlERαKO (n = 10), ARCERαKO (n = 12) and control (grey, n = 7 and 5) female mice. NS, not significant. b, ERα and pS6(S244/S247) co-expression (arrows) in proestrus and oestrus (representative of five mice). Scale bars, 100 μm. c, Number of pS6(S244/S247)-labelled VMHvl (***P = 0.0005) and ARC cells in vehicle-treated (Veh.) (n = 4) or oestradiol-benzoate-treated (EB) (n = 5) female mice. d, Enrichment of peptide ligand-binding receptors (red). Dashed line, Benjamini–Hochberg-adjusted P < 0.05. e, VMHvl Mc4r expression in proestrus (P; n = 5), male (n = 6, *P = 0.0189) and oestrus (O; n = 5, *P = 0.0163) mice. f, Mc4r and Esr1 expression in oestrus and proestrus (representative of five mice). Scale bars, 100 μm. g, Counts-per-million (CPM)-normalized coverage tracks of ERE-containing ERα-binding sites (pink boxes) within Mc4r and Nmur2 loci in sub-cortical nuclei of vehicle- and oestradiol-benzoate-treated gonadectomized mice (n = 3; MACS2, q < 0.01). phyloP60wayPlacental track from UCSC shows sequence conservation (Cons.) of ERα-binding sites in placental mammals. Data are mean ± s.e.m., scatter plots or box plots. In box plots, whiskers indicate the minimum and maximum values, the edges of the box are 25th and 75th percentiles, and the centre line indicates the mean. a, c, Unpaired two-tailed Student’s t-test. e, One-way ANOVA with Holm–Šidák multiple comparisons test. Anatomical abbreviations are included in Extended Data Table 1.
Hormone responsiveness of VMHvlERα neurons was visualized across the oestrous cycle by monitoring phosphorylated ribosomal protein S6 (pS6) expression during oestrus (low E2) and proestrus (high E2). VMHvl pS6 signals increase substantially during proestrus or after an injection of oestradiol benzoate in ovariectomized (OVX) female mice (Fig. 1b, c and Extended Data Fig. 2a), but were negligible during oestrus, in female mice lacking ERα or in intact, untreated male mice (Extended Data Fig. 2b, c), underscoring a complete dependence of this pS6 response on both oestrogen and ERα. Oestrogen induction of pS6 in VMHvlERα neurons occurs through a classical genomic mechanism that begins slowly starting at 2 h after treatment. By contrast, no hormone-dependent induction of pS6 was detected in adjacent ARCERα neurons (Fig. 1c and Extended Data Fig. 2a). The fact that VMHvlERα neurons respond highly to oestrogen, but not to fasting11, suggests that fluctuating hormones rather than hunger stimulate these neurons, setting the stage for behavioural changes across the reproductive cycle.
MC4R levels are controlled by oestrogen
Candidate mediators of ERα signalling were identified after profiling the VMHvl transcriptome in OVX female mice treated with vehicle or oestradiol benzoate (Fig. 1d). Among differentially expressed genes we noted enrichment of peptidergic G-protein-coupled receptors (Mc4r, Nmur2, Npy1r and Ghsr) and known oestrogen-dependent genes (Greb1, Pgr4), some of which are linked to locomotor activity (MP:0003313, adjusted P = 3.19 × 10−4) (Extended Data Fig. 2d, e). We focused on Mc4r given its expression in the VMH12, its role in locomotor behaviour13,14 and observed sex differences in Mc4r loss-of-function mutations in mice13,15 and humans9,16. Mc4r was induced in female mice during proestrus but not in oestrus nor in intact male mice (Fig. 1e). We confirmed increased Mc4r expression in VMHvl neurons during proestrus or after oestradiol benzoate treatment that colocalized with ERα (Esr1) and Rprm, a VMHvl female-specific marker7 (Fig. 1f and Extended Data Fig. 2f–i).
We further established that Mc4r is a direct transcriptional target of ERα using CUT&RUN (Cleavage Under Targets and Release Using Nuclease), a technique that detects transient in vivo binding events within heterogeneous tissues17. As expected, hormone-dependent ERα–chromatin interactions were detected in Greb1 and Pgr (Extended Data Fig. 3a, b). The high sensitivity afforded by CUT&RUN enabled the detection of two conserved ERα-binding sites within the Mc4r locus (Fig. 1g and Extended Data Fig. 3c, d). The first, located −210 kb downstream of the transcript, contains a canonical oestrogen-response element (ERE) consensus sequence. The second, in the proximal promoter, consists of an ERE half-site and a site for the trans-acting transcription factor 1 (Sp1) that together coordinate the oestrogen-dependent global regulation of ERα-target genes18. An ERE was detected +200 kb upstream of Nmur2, consistent with its upregulation during proestrus (Fig. 1g and Extended Data Figs. 2e, 3e). These data establish a direct molecular link between ERα and MC4R and imply that oestrogen dynamically regulates the responsiveness of VMHvlERα neurons to neuropeptides.
VMHvlMC4R neurons project to arousal centres
ERα and MC4R coexpression, assessed using mice expressing a Cre-dependent reporter under the the control of Mc4r-t2a-cre (Ai14Mc4r), revealed that VMHvlERα/MC4R neurons are a subset of the VMHvlERα population (Fig. 2a). This near-perfect concordance of Ai14Mc4r and ERα and stage-dependent Mc4r induction was not detected in the medial amygdala (MeA) (Fig. 2a and Extended Data Fig. 4a, b). ERα was undetected in the paraventricular hypothalamus (PVH), a primary site that couples MC4R with food intake (Fig. 2a).
a, ERα and Ai14 expression and quantification in the PVH (n = 4), VMHvl (n = 9, ****P < 0.0001) and MeA (n = 4, **P = 0.0057) of Ai14Mc4r female mice. Scale bars, 100 μm. b, Labelling vector and map of major VMHvlMC4R projections. c, VMHvl projections to anterior (top) and posterior (bottom) regions. Images representative of bilateral VMHvl targeting (n = 3 mice). Scale bars, 200 µm. d, Semi-quantitative comparison of VMHvlMC4R and VMHvlERα projection19 intensities. Clusters I and II indicate unique anatomical targets of VMHvlERα and VMHvlMC4R neurons, respectively. Anatomical abbreviations are included in Extended Data Table 1. A description of the box plots is provided in the legend of Fig. 1c. a, One-way ANOVA with Holm–Šidák multiple comparisons test.
We then asked how afferent VMHvlMC4R neuron projections, labelled by Cre-dependent, membrane-targeted YFP (mYFP), compared with the broader VMHvlERα population19 (Fig. 2b). Overall, although many (~84%) of the same major projections reported for VMHvlERα neurons were identified, robust targeting to the ARC and MeA was not observed (Fig. 2c, d cluster I and Extended Data Fig. 4c, d). Unexpectedly, VMHvlMC4R neurons projected to the dorsal CA1 and the adjacent subiculum (Fig. 2c, d cluster II), a hippocampal region that controls locomotor speed in mice20 and contains ‘speed cells’, the firing rate of which correlates with velocity21. Expected projections from VMHvlMC4R neurons to the midbrain pre-motor periaqueductal grey (PAG) region showed a unique pattern restricted to the lateral and dorsolateral PAG columns (PAGdl/l) associated with escape behaviours22, while conspicuously avoiding the ventrolateral PAG (PAGvl), which is involved in freezing and defensive behaviours23 (Extended Data Fig. 4e). VMHvlMC4R neurons also projected to the hindbrain pontine region containing a cluster of nuclei that mediate sexual receptivity and locomotor arousal24,25.
Activating VMHvlMC4R node offsets oestrogen loss
To assess the functional output of VMHvlERα/MC4R neurons, we stimulated this population using Cre-dependent DREADDs (Designer Receptors Exclusively Activated by Designer Drugs; AAV-DIO-hM3Dq-mCherry) injected bilaterally into the VMHvl of Mc4r-t2a-cre mice and control littermates (Fig. 3a). Administration of clozapine N-oxide (CNO) during the inactive period (light) substantially increased spontaneous physical activity in female and male VMHvlMC4R::hM3Dq mice, but not in VMHvlCre− controls (Fig. 3b and Extended Data Fig. 5a). Responses to a single injection of CNO lasted approximately 5 h in VMHvlMC4R::hM3Dq mice, with the distance travelled increasing by 700%, concomitant with a precipitous decrease in immobile behaviour (Extended Data Fig. 5b).
a, ERα and mCherry in a VMHvlMC4R::hM3Dq female mouse. Scale bars, 200 µm (top) and 10 µm (bottom). b, Spontaneous activity during zeitgeber time (ZT)3–ZT8 in VMHvlMC4R::hM3Dq mice (female, n = 5, ****P < 0.0001; male, n = 5, ****P < 0.0001) and VMHvlCre− (female, n = 5; male, n = 4) with or without CNO injection. c, Left, Thermography images of VMHvlCre− (left) and VMHvlMC4R::hM3Dq (right) female mice. Right, iBAT surface temperature analysis of VMHvlCre− (left) and VMHvlMC4R::hM3Dq (right) female mice, 30 and 45 min after saline (Sal.), CNO or CL-316,243 (CL) injection (n = 4 VMHv1Cre− or 5 VMHvlMC4R::hM3Dq mice; VMHvlCre−, ***P < 0.0001; VMHvlMC4R::hM3Dq, **P = 0.0003). d, Food consumption normalized to body weight in female mice (n = 5 and 5, respectively) following saline or CNO injection during the light period (zeitgeber time (ZT)4–ZT9). e, Weight change over 24 h in female mice (n = 5 and 5, respectively) administered drinking water (H2O) or CNO-containing water (CNO, ****P < 0.0001). f, Dark period (ZT12–ZT24) activity in VMHvlMC4R::hM4Di (n = 8) and VMHvlCre− (n = 4) female mice administered water or DCZ-containing water. g, Total dark period distance, rearing and immobile time in VMHvlMC4R::hM4Di (n = 8) and VMHvlCre− (n = 4) female mice ± DCZ-containing water. ****P < 0.0001, **P = 0.0068, *P = 0.0213. h, Activity levels in intact (n = 16), OVX (n = 12, ****P < 0.0001) and OVX CNO-administered VMHvlMC4R::hM3Dq (n = 5, **P = 0.0023) mice. i, j, Body weight (i) and gonadal white adipocyte area (j) after the 8-day CNO treatment of OVX and HFD-fed mice (n = 5 and 5 for of VMHvlCre− and VMHvlMC4R::hM3Dq mice). Mice were fed a HFD for 6 weeks. Scale bars, 100 µm. Data are mean ± s.e.m., scatter plots or box plots. For definitions of the box plots, see the legend of Fig. 1c. b–f, h, Repeated-measures two-way ANOVA with Holm–Šidák multiple comparisons test. g, Unpaired two-tailed Student’s t-tests. h, One-way ANOVA with Holm–Šidák multiple comparisons test. i, Nested t-test.
Other than the increased movement, other metabolic functions were insensitive to VMHvlMC4R neuron stimulation. For example, compared to a β3-adrenergic agonist (CL-316,243), CNO did not elevate the iBAT temperature or Ucp1 expression in VMHvlMC4R::hM3Dq mice (Fig. 3c and Extended Data Fig. 5c, d). Glucose homeostasis was unchanged in CNO-treated VMHvlMC4R::hM3Dq mice; although the higher body weights inherent to the Mc4r-t2a-cre line increased fasting glucose (Extended Data Fig. 5e, f). Food intake was unaffected by stimulation during the light period and decreased modestly during the dark period (Fig. 3d and Extended Data Fig. 5g). Providing CNO in the drinking water over 24 h led to a nearly 10% drop in body weight in VMHvlMC4R::hM3Dq female mice with a corresponding increase in activity (Fig. 3e, Supplementary Video 1 and Extended Data Fig. 5h–j) and resulted in a 13% drop in body weight when extended over 8 days (Extended Data Fig. 5k). Conversely, targeting the VMHvl with inhibitory DREADDs (AAV-DIO-hM4Di-mCherry) increased sedentary behaviour during the dark period after the administration of the DREADD ligand, deschloroclozapine (DCZ) (Fig. 3f, g and Extended Data Fig. 5l, m). Thus, the marked changes in physical activity after chemogenetic or genetic manipulation of VMHvlERα/MC4R cells suggest that this neuron cluster is an essential generator of maximal physical activity in female mice and constitutes a potent node for promoting physical activity, which can be artificially engaged in both sexes.
Increased sedentary behaviour and metabolic decline are hallmarks of declining oestrogen during ageing. We investigated whether DREADD-activation of VMHvlMC4R neurons overrides these deleterious features in oestrogen-depleted OVX female mice. Stimulating VMHvlMC4R neurons over a short 24 h period fully restored physical activity parameters and promoted significant weight loss in OVX female mice (Fig. 3h and Extended Data Fig. 6a, b). Stimulating this node in OVX female mice fed a high-fat diet (HFD) (Extended Data Fig. 6c) reversed the overt metabolic impairment due to oestrogen depletion and chronic overnutrition. Fasting glucose and insulin tolerance improved notably after a single bout of CNO-induced activity in HFD-fed VMHvlMC4R::hM3Dq female mice (Extended Data Fig. 6d–f). Furthermore, chronic stimulation of VMHvlMC4R neurons in obese, sedentary OVX female mice resulted in rapid, marked weight loss, accompanied by lowered fasting blood glucose, a drop in cellular adiposity of gonadal fat and reduced plasma cholesterol (Fig. 3i, j and Extended Data Fig. 6g–i), all indices of improved metabolic health. Food intake was unaffected (Extended Data Fig. 6j). Hence, engagement of the VMHvlMC4R activity node reduces body weight in OVX female mice and improves metabolic health in the face of a dietary challenge and oestrogen depletion.
MC4R gene editing in VMHvl drives activity
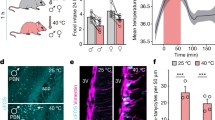
To determine whether melanocortin signalling itself regulates this VMHvl activity node, we initially confirmed that MT-II, a synthetic MC4R agonist, evoked FOS expression in female mice pretreated with oestradiol benzoate but not with vehicle (Extended Data Fig. 7a, b). We next used the Cre-dependent Mc4rloxTB allele26 in combination with the Sf1-cre transgene, which only overlaps with Mc4r expression in the VMH (Extended Data Fig. 7c) to restore Mc4r in the VMH of otherwise Mc4r-null mice (Mc4rSf1-cre). In response to oestradiol benzoate, this rescue approach increased Mc4r expression in VMHvlERα neurons, similar to wild-type (Mc4r+/+) female mice (Fig. 4a). Body weights were equivalent at weaning (Extended Data Fig. 7d). Consistent with the loss of PVH MC4R signalling, null Mc4rloxTB and rescued Mc4rSf1-cre female mice developed obesity, hyperphagia and increased body lengths26 compared with control littermates. However, restoring Mc4r in the VMHvl attenuated overt weight gain and sedentary behaviour in female but not male Mc4rSf1-cre mice (Fig. 4b, e and Extended Data Fig. 7d–g). Thus, our data solidify the role of MC4R signalling in the female VMHvl for promoting spontaneous activity.
a, Esr1 and Mc4r expression in Mc4r+/+, Mc4rloxTB and Mc4rSf1-cre female mice. Scale bars, 200 µm (left) and 100 µm (right). b, Body weights of 8-week-old female (**P = 0.0026) and male Mc4r+/+, Mc4rloxTB and Mc4rSf1-cre mice. c, Food intake in 8-week-old female mouse cohorts. d, Body length in 8-week-old female mouse cohorts. e, x-ambulatory activity of female mice during the light and dark periods (****P < 0.0001, *P = 0.0153). f, Mc4rCRISPRa targets the Mc4r promotor. g, Esr1 and Mc4r expression in control and Mc4rCRISPRa female and male mice. Scale bar, 200 µm. h, Home-cage activity in Mc4rCRISPRa (n = 6) and control (n = 5) female mice. i, Distances for the three most active runs of Mc4rCRISPRa and control female (n = 6 and 5, respectively, ****P < 0.0001) and male (n = 4 and 3, respectively) mice. j, Fraction of cortical bone volume of female mice 4 months after infection (P = 0.0129). CBV, cortical bone volume; TV, tissue volume. k, VMHvlERα/MC4R neurons integrate oestrogen and melanocortin signalling to generate a specialized hormone-dependent activity node in female mice. Data are mean ± s.e.m. or box plots, and number of mice analysed are indicated on or above each bar. For definitions of the box plots, see the legend of Fig. 1c. b, c, d, One-way ANOVA with Holm–Šidák multiple comparison test. e, i, Repeated-measures two-way ANOVA with Holm–Šidák multiple comparison test. j, Unpaired two-tailed Student’s t-tests.
To verify that MC4R signalling is an integral component of the hormone-responsive VMHvl activity node, CRISPR-mediated activation (CRISPRa) was used to increase Mc4r expression. Previously, in haploinsufficient Mc4r+/− mice, gene dosage and energy imbalance were normalized by CRISPRa targeting the PVH27. Here, wild-type female and male mice were stereotaxically injected with a dual vector system containing a guide RNA targeting the Mc4r promoter ERE half-site (AAV-Mc4r-Pr-sgRNA) and dCas9 tethered to the VP64 transcriptional activator (AAV-dCas9-VP64) to selectively upregulate Mc4r expression in the VMHvl (Fig. 4f). Control mice received dCas9–VP64 without sgRNA. Delivery of Mc4r-CRISPRa-viral vectors to the VMHvl was confirmed post mortem and revealed moderate but long-lived induction of Mc4r in both sexes (Fig. 4g and Extended Data Fig. 8a–c). Mc4rCRISPRa female mice travelled twice the distance in the dark compared to control mice, with increased movement persisting for at least 17 weeks after injection (Fig. 4h, i). Activity in Mc4rCRISPRa male mice also increased, and in both sexes the drop in sedentary behaviour in Mc4rCRISPRa mice was restricted to the nighttime, thus preserving regular diurnal activity patterns (Fig. 4i and Extended Data Fig. 8d, e). The lack of weight loss in female Mc4rCRISPRa mice may reflect the modest but significant increase in daily food intake in female mice. BAT activity was unchanged (Extended Data Fig. 8f–h). Weeks of increased physical activity (and mechanical loading) in Mc4rCRISPRa female mice increased cortical bone thickness and bone volume (Fig. 4j and Extended Data Fig. 8i). We noted that, under these conditions, Mc4rCRISPRa did not restore normal activity in OVX female mice (Extended Data Fig. 8j–l). Nevertheless, bypassing ERα and directly increasing Mc4r dosage in the VMHvl permanently increases spontaneous activity behaviour in intact female and male mice.
Concluding remarks
Here, we identify an oestrogen-sensitive VMHvlERα/MC4R node that maximizes daily patterns of spontaneous physical activity in female mice. MC4R is an essential intermediary component coupling oestrogen and energy expenditure as a direct ERα transcriptional target (Fig. 4k). Thus, as Mc4r expression increases during the preovulatory period, sensitivity to melanocortin increases in the VMHvl, resulting in spikes of oestrogen-dependent activity that were first described in 192428. Our findings explain how oestrogen drives an essential behavioural output during a critical point in the reproductive cycle.
As human gain-of-function MC4R variants reduce receptor turnover and protect against weight gain29, identifying endogenous signals that modulate MC4R expression becomes of interest. We identify oestrogen as a potent inducer of Mc4r expression. The high degree of conservation in consensus ERE-binding motifs in the mammalian MC4R locus suggests that oestrogen similarly upregulates human MC4R expression. MC4R agonists that elicit sexual behaviours in oestrogen-primed female rodents30 and enhance libido in premenopausal women suffering from hypoactive sexual desire disorder31 may act by directly targeting the VMHvlERα/MC4R node. Once engaged, VMHvlMC4R neurons project to regions of the central nervous system that are involved in reproductive behaviours, as well as sites in the hippocampal region that regulate the speed and orientation of locomotion20,21, and in hindbrain regions associated with arousal and motor output32. It remains to be determined whether these VMHvl outputs contribute to psychiatric disorders (for example, postpartum depression or premenstrual dysphoric disorder) that coincide with periods of hormonal fluctuations. Conversely, curtailment of MC4R expression after oestrogen depletion might underlie the increased sedentary lifestyle associated with menopause33.
Despite the pronounced increase in physical activity in Mc4rCRISPRa female mice, body weights remained stubbornly constant in the face of small increases in daily food intake. Whereas DREADD activation of VMHvlMC4R neurons reduced body weight rapidly in oestrogen-depleted OVX female mice, this rate of weight loss was not sustainable (Extended Data Fig. 7g). Collectively, these results reinforce the notion that adaptive responses limit the extent of exercise-induced weight loss34. Nonetheless, increasing physical activity reduces the risk of metabolic- and age-related co-morbidities, including heart disease, frailty, cancer and infectious diseases35. As such, the extremely durable increase in spontaneous physical activity achieved by the non-transgenic Mc4rCRISPRa approach provides a unique preclinical model to explore the motivational aspects and long-term health benefits of an active lifestyle. Our findings underscore the benefits of oestrogen in minimizing sedentary behaviour and provoke further discussion about hormone replacement therapies in postmenopausal women.
Methods
Mice
All experiments were conducted in accordance with institutional guidelines and approved protocols for animal care and use at the University of California San Francisco (UCSF) and Cold Spring Harbor Laboratory. Mice were housed on a 12 h:12 h light:dark cycle (lights on, 06:00; lights off, 18:00) and had ad libitum access to standard chow (LabDiet, 5058) or high-fat diet (Research Diets, D12492). Mc4rloxTB mice and the Ai14fl/fl reporter mice were purchased from Jackson Laboratories and maintained on a C57BL/6J background. Mc4r-t2a-cre mice36 were a gift from B. Lowell and were maintained on a C57BL/6J background. Esr1fl/fl mice were maintained on a mixed background, and Sf1-cre mice37 were maintained on a C57BL/6N in the laboratory as previously described3,4. Wild-type mice used for CRISPRa studies were on a pure C57BL/6J background. For Mc4r rescue experiments, Sf1-cre was contributed through female mice. CUT&RUN experiments were performed on adult male (8–12 weeks of age) gonadectomized C57Bl6/J wild-type mice obtained from Jackson Laboratories. Three weeks after gonadectomy, animals were injected subcutaneously with either corn oil (vehicle) or 5 µg of oestradiol benzoate and euthanized after 4 h. For each biological replicate, brain dissections were pooled from five animals.
Stereotaxic injections
AAV2-Cre-GFP and AAV2-GFP were purchased from the Vector Core at the University of North Carolina at Chapel Hill. AAV2-hM3Dq-mCherry and AAV2-hM4Di-mCherry were gifts from B. Roth and viral preparations were purchased from Addgene (viral prep no. 44361-AAV2, http://n2t.net/addgene:44361, RRID: Addgene_44361; and viral prep no. 44362-AAV2, http://n2t.net/addgene:44362, RRID: Addgene_44362; Addgene)38. For axonal tracing, AAV2-CAGs-FLEX-membrane-YFP-WPRE.hGH was re-engineered to fuse YFP with a C-terminal farnesylation tag to enhance the membrane labelling39. AAV2 was prepared using a standard polyethylene-glycol gradient followed by a caesium-chloride density-gradient centrifugation protocol40 to reach a titre of (1 × 1013 genome copies per ml). AAVdj-dCas9-VP64 and AAVdj-Prm-Mc4r-sgRNA were generated by the Stanford Gene Vector and Virus Core and details of vector constructs are as previously described27. Adult mice were secured in a Model 1900 stereotaxic frame (David Kopff Instruments) and 250–600 nl of virus was injected bilaterally at the following coordinates. For the VMHvl, anterior–posterior: bregma −1.48 mm, mediolateral: bregma ±0.85 mm, dorsoventral: skull −5.9 mm. For the ARC, anterior–posterior: bregma −1.58 mm, mediolateral: bregma ±0.25 mm, dorsoventral: skull −5.8 mm.
For all surgeries regardless of viral vectors used, mice recovered for at least 2 weeks before any metabolic or behavioural assays. For projection labelling, mice were allowed to express the reporter for 5−8 weeks before tissue collection. At the conclusion of the experiments, mice were euthanized and the brains were collected to confirm proper targeting. Any mice absent of correctly targeted fluorescent protein expression were excluded from subsequent analyses. Water-soluble CNO (HB6149; Hello Bio) was administered by intraperitoneal injection (0.3 mg kg−1 in sterile saline) or in the drinking water (0.25 mg ml−1). CNO-laden drinking water was replaced every 48 h. Water-soluble DCZ (HB9126; Hello Bio) was administered in the drinking water (0.1 mg ml−1)41.
Oestrous cycle staging and oestradiol benzoate treatment
Reproductive stages in female mice were determined by comparing relative amounts of leukocytes, epithelial cells and cornified epithelial cells collected by vaginal lavage. Stage assessments were made daily between ZT3 and ZT5. Brains from oestrus or proestrus female mice were collected between ZT7 and ZT10 and processed for immunofluorescence, in situ hybridization (ISH) or qPCR.
Adult female mice (>8 weeks old) were ovariectomized. Oestradiol benzoate (Cayman Chemical, 10006487) was dissolved in DMSO and diluted in sesame oil (Sigma, S3547). Mice received a subcutaneous injection of either 1 ug oestradiol benzoate in 150 µl sesame oil or 150 µl of sesame oil with an equivalent amount of DMSO. Control mice received a subcutaneous injection of 150 µl of sesame oil with an equivalent amount of DMSO. To minimize changes in VMH gene expression or signal transduction associated with fear and/or anxiety, mice were handled daily in a manner that simulated injection for at least 5 days before oestradiol benzoate or vehicle treatment and tissue collection. For FOS analyses, mice were treated with 400 µg MT-II (Bachem) by intraperitoneal injection, and brains were collected 1–1.5 h later.
RNA sequencing and qPCR
Brains from OVX female mice treated with oestradiol benzoate (n = 4) or vehicle (n = 3) were rapidly dissected into ice-cold PBS with 0.1% DEPC. Coronal brain sections (250 µm thick) were cut on a vibratome and transferred to glass slides so that the VMH could be visualized and manually microdissected. Isolated tissue was flash-frozen and stored at −80 °C. RNA was prepared using the RNeasy Micro kit (Qiagen). Sequencing libraries were constructed using the TRIO RNA-seq Library Preparation kit (TECAN) using 15 ng of input RNA. Equal amounts of each sample library were multiplexed and sequenced (50-bp single-end reads) on a single flow cell lane HiSeq 4000 (Illumina). Demultiplexed reads were aligned to the mouse genome (mm10) using HISAT242 v.2.1.0 and counted using HTSeq43 v.0.9.1. Finally, differential gene expression testing was performed using DESeq244 v.1.14.1.
Isolated RNA, prepared as described above, was converted to cDNA using the SuperScript III reverse transcriptase (Invitrogen). qPCR was performed using a BioRad CFX instrument with Maestro software v.4.1.2433.1219. Target genes were amplified using specific primers (Mc4r forward, 5′-GCCAGGGTACCAACATGAAG-3′ and reverse, 5′-ATGAAGCACACGCAGTATGG-3′; Nmur2 forward, 5′-CCTCCTTCCTCTTCTACATCCT-3′ and reverse, 5′-AGTCACTTTGTCTGCCTCAA-3′; Esr1 forward, 5′-GAACGAGCCCAGCGCCTACG-3′ and reverse, 5′-TCTCGGCCATTCTGGCGTCG-3′; and Ucp1 forward, 5′-CACGGGGACCTACAATGCTT-3′ and reverse, 5′-TAGGGGTCGTCCCTTTCCAA-3′). Ct values were normalized to cyclophilin B (Ppib; forward primer, 5′-TGGAGAGCACCAAGACAGACA-3′ and reverse primer, 5′-TGCCGGAGTCGACAATGAT-3′) and relative expression levels were quantified using the comparative Ct method. Individual values, representing the VMHvl or iBAT from one mouse are the average of two technical replicates.
CUT&RUN
ERα CUT&RUN was performed on 400,000 nuclei isolated from BNSTp, POA and MeA tissue using density gradient centrifugation45. In brief, tissue was homogenized 15× with a loose pestle in a glass homogenizer containing homogenization medium (250 mM sucrose, 25 mM KCl, 5 mM MgCl2, 20 mM Tricine KOH, 1 mM DTT, 0.15 mM spermine, 0.5 mM spermidine, 1× Roche EDTA-free protease inhibitor cocktail, pH 7.8). Then, 0.3% IGEPAL CA-630 was added, and the tissue was further dounced 5× with a tight pestle. After douncing, the homogenate was filtered through a 40-µm strainer and mixed 1:1 with 50% OptiPrep solution (Millipore Sigma) prepared in dilution buffer (150 mM KCl, 30 mM MgCl2, 120 mM Tricine KOH, pH 7.8). The homogenate was underlaid with 5 ml each of 30% and 40% OptiPrep solution, and centrifuged at 10,000g for 18 min at 4 °C in an ultracentrifuge. Then, 2 ml of nucleus-containing solution was removed from the 30–40% OptiPrep interface by direct tube puncture. After the isolation of the nuclei, 0.4% IGEPAL CA-630 was added to improve binding to concanavalin A magnetic beads (Bangs Laboratories BP531). CUT&RUN was performed on brain nuclei, according to the standard protocol17. Nuclei were washed twice in wash buffer (20 mM HEPES, pH 7.5, 150 mM NaCl, 0.1% BSA, 0.5 mM spermidine, 1× protease inhibitor cocktail) and incubated overnight on a nutator with ERα antibody (Millipore Sigma, 06-935), diluted 1:100 in antibody buffer (wash buffer containing 2 mM EDTA). Nuclei were washed twice in wash buffer, and around 700 ng ml−1 protein-A-MNase (pA-MNase) was added. After 1 h incubation on a nutator at 4 °C, the nuclei were washed twice in wash buffer and placed in a metal heat block on ice. pA-MNase digestion was initiated by 2 mM CaCl2. After 90 min, pA-MNase activity was stopped by mixing 1:1 with 2× stop buffer (340 mM NaCl, 20 mM EDTA, 4 mM EGTA, 50 µg ml−1 RNase A, 50 µg ml−1 glycogen). Digested fragments were released by incubating at 37 °C for 10 min, followed by centrifuging at 16,000g for 5 min at 4 °C. DNA was purified from the supernatant by phenol–chloroform extraction.
CUT&RUN library preparation
CUT&RUN libraries were prepared using the SMARTer ThruPLEX DNA-seq Kit (Takara Bio), with the following PCR conditions: 72 °C for 3 min, 85 °C for 2 min, 98 °C for 2 min, (98 °C for 20 s, 67 °C for 20 s, 72 °C for 30 s) for 4 cycles, (98 °C for 20 s, 72 °C for 15 s) for 10 cycles. Samples were size-selected with AMPure XP beads (1.5× right-sided and 0.5× left-sided) to remove residual adapter dimers and large DNA fragments. Individually barcoded libraries were multiplexed and sequenced with paired-end 75-bp reads on an Illumina NextSeq, using the High Output Kit
CUT&RUN data processing
Paired-end reads were trimmed with cutadapt46 v.3.2.0 to remove low-quality base calls (-q 30) and adapters. Trimmed reads were aligned to mm10 using Bowtie247 v.2.4.2 with the following flags: -dovetail -very-sensitive-local -no-unal -no-mixed -no-discordant -phred33. After alignment, duplicate reads were removed using Picard v.2.21.6 (http://broadinstitute.github.io/picard/) MarkDuplicates (REMOVE_DUPLICATES = true). Deduplicated reads were filtered by mapping quality (MAPQ > 40) using samtools48 v.1.11.0 and fragment length (<120 bp) using deepTools v.3.5.0 alignmentSieve49. After filtering, peaks were called using MACS2 v.2.2.7.1 callpeak50 with a q-value threshold of 0.01 and min-length set to 25. Peaks shown in Fig. 1 were called in 2 out of 3 replicates. Individual replicate BAM files were normalized by CPM and converted to bigwig tracks, using deepTools bamCoverage (-bs 1, -normalize using CPM). CPM-normalized bigwig tracks for individual oestradiol benzoate and vehicle samples (n = 3 per condition) were plotted using Gviz51.
ISH
For colorimetric ISH, antisense Mc4r probes were PCR-amplified (forward primer, 5′-ACTCTGGGTGTCATAAGCCTGT-3′ and reverse primer, 5′-TCTGTCCCCCACTTAATACCTG-3′) from hypothalamic cDNA libraries, and in vitro transcribed with incorporation of digoxigenin-UTP (Roche) using the T7 or SP6 Riboprobe kit (Promega). The 20-µm sections from fixed tissue were labelled and detected by chromogenic immunohistochemistry as previously described3. Fluorescent ISH was performed using RNAScope (ACD, Multiplex Fluorescent v.2) according to the manufacturer’s protocol using the following probes: Esr1 (478201), Mc4r (319181-C2) and Rprm (466071).
Immunofluorescence staining and histology
Fixed central nervous system tissue was cryosectioned (20 µm) and stained overnight with primary antibodies against: ERα (EMD Millipore, C1355, polyclonal rabbit, 1:750 dilution or Abcam, 93021, monoclonal mouse, 1:100 dilution), pS6(S244/S247) (RPS6) (Invitrogen, 44-923G, polyclonal rabbit, 1:500 dilution), FOS (Santa Cruz, SC-52, polyclonal rabbit, 1:500 dilution) or red fluorescent protein (RFP; Rockland, 600-401-379, polyclonal rabbit, 1:1,000 dilution). For detection, sections were labelled with species-appropriate secondary Alexa-Fluor-coupled antibodies (Invitrogen, A11029 and A11037, 1:1,000 dilution for both). Widefield images were acquired using a Nikon microscope and NIS-Elements v.3.22.15. Confocal images were acquired at the UCSF Nikon Imaging Center using a Nikon CSU-22 with EMCCD camera and MicroManager v.2.0gamma. Images were processed and quantified using ImageJ FIJI v.1.52i and the Cell Counter plugin v.2.
Fixed gonadal white adipose tissue was paraffin-embedded, sectioned (5 µm) and stained with haematoxylin and eosin by the Gladstone Histology and Light Microscopy core. Brightfield images were thresholded to define adipocyte borders, and the adipocyte area was quantified using ImageJ FIJI.
Micro-computed tomography
After perfusion fixation, femurs from Mc4rCRISPRa and control female mice were isolated. Volumetric bone density and bone volume were measured by micro-computed tomography as previously described4.
Metabolic and activity monitoring
Indirect calorimetry and food intake were measured in CLAMS chambers (Comprehensive Laboratory Animal Monitoring System, Columbus Instruments) and analysed using CLAX v.2.2.15. Any spilled food that was not consumed was accounted for at the conclusion of the 4-day period spent in CLAMS.
Ambulatory activity in mice that received chemogenetic or CRISPRa manipulations was recorded using infrared cameras and quantified using the ANY-maze behavioural tracking system (Stoelting, v.6.33). Before any measurements, mice were acclimatized to single housing in the ANY-maze chambers for at least 3 days. For CRISPRa studies, activity tracking was continuously monitored for at least five 24-h periods.
Interscapular skin temperatures were measured using a FLIR-E4 handheld infrared camera (FLIR Systems) and the FLIR Tools analysis software v.5.13.18031.2002 as previously described52. Female mice were lightly anaesthetized in groups of four or five in an anaesthesia induction chamber and images were captured at baseline, 30 min and 60 min after intraperitoneal injection of saline, CNO (0.3 mg kg−1) or CL-316,243 (3 mg kg−1).
Blood glucose and lipid assays were performed after a 6-h fast (starting around ZT2) during which mice were housed in clean cages with ad libitum water access. For glucose- and insulin-tolerance tests, fasted mice were intraperitoneally injected with glucose (1 g kg−1) or insulin (1 U kg−1), respectively. Tail-blood samples were collected at baseline and every 15 min after glucose or insulin injection. Blood glucose levels were quantified using a handheld glucometer (Roche, Accu-Check Compact). For triglyceride and cholesterol measurements, plasma was isolated from tail-blood and measured (3 µl in technical duplicates) using the Cholesterol Quantitation Kit (Sigma, MAK043) or the Triglyceride Quantification Colorimetric/Fluorometric Kit (Sigma, MAK266).
Statistics
Statistical tests, excluding RNA-sequencing analyses, were performed using Prism 8 (GraphPad). A description of the test and results are provided in Extended Data Table 2. Multiple-comparisons correction for one-way, two-way and repeated-measures ANOVA were performed using the Holm–Šidák post hoc test. Unless otherwise noted, data are mean ± s.e.m. or box plots in which whiskers represent minimum and maximum values, edges of the box are 25th and 75th percentiles, and the centre line indicates the mean. Sample sizes were based on previous work from our laboratory; however, no specific statistical calculation was performed to determine sample size. For chemogenetic manipulation, mice were drawn at random from a pool of littermate mice containing a roughly equal mix of cre+ and cre− genotypes; both mice were injected with the same Cre-dependent AAV construct; partitioning into control and experimental groups was therefore determined by genotype. For AAV-Cre, AAV-mYFP and CRISPRa injections mice of identical genotypes were drawn at random from littermate pools to receive functional or control virus injections. Physical activity and food intake parameters were objectively measured using the ANY-maze and/or CLAMS automated systems. Measurements of BAT temperature and mRNA expression as well as cortical bone parameters were made by experimenters blinded to the type of AAV received and/or genotype of the mice studied. The large differences in body weights among Mc4r-control, -null and -rescue mice precluded blinding of the genotype of these mice.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
References
Mauvais-Jarvis, F., Clegg, D. J. & Hevener, A. L. The role of estrogens in control of energy balance and glucose homeostasis. Endocr. Rev. 34, 309–338 (2013).
Carr, M. C. The emergence of the metabolic syndrome with menopause. J. Clin. Endocrinol. Metab. 88, 2404–2411 (2003).
Correa, S. M. et al. An estrogen-responsive module in the ventromedial hypothalamus selectively drives sex-specific activity in females. Cell Rep. 10, 62–74 (2015).
Herber, C. B. et al. Estrogen signaling in arcuate Kiss1 neurons suppresses a sex-dependent female circuit promoting dense strong bones. Nat. Commun. 10, 163 (2019).
Martinez de Morentin, P. B. et al. Estradiol regulates brown adipose tissue thermogenesis via hypothalamic AMPK. Cell Metab. 20, 41–53 (2014).
Xu, Y. et al. Distinct hypothalamic neurons mediate estrogenic effects on energy homeostasis and reproduction. Cell Metab. 14, 453–465 (2011).
van Veen, J. E. et al. Hypothalamic estrogen receptor alpha establishes a sexually dimorphic regulatory node of energy expenditure. Nat. Metab. 2, 351–363 (2020).
Farooqi, I. S. et al. Dominant and recessive inheritance of morbid obesity associated with melanocortin 4 receptor deficiency. J. Clin. Invest. 106, 271–279 (2000).
Qi, L., Kraft, P., Hunter, D. J. & Hu, F. B. The common obesity variant near MC4R gene is associated with higher intakes of total energy and dietary fat, weight change and diabetes risk in women. Hum. Mol. Genet. 17, 3502–3508 (2008).
Thammacharoen, S., Lutz, T. A., Geary, N. & Asarian, L. Hindbrain administration of estradiol inhibits feeding and activates estrogen receptor-α-expressing cells in the nucleus tractus solitarius of ovariectomized rats. Endocrinology 149, 1609–1617 (2008).
Villanueva, E. C. et al. Complex regulation of mammalian target of rapamycin complex 1 in the basomedial hypothalamus by leptin and nutritional status. Endocrinology 150, 4541–4551 (2009).
Mountjoy, K. G., Mortrud, M. T., Low, M. J., Simerly, R. B. & Cone, R. D. Localization of the melanocortin-4 receptor (MC4-R) in neuroendocrine and autonomic control circuits in the brain. Mol. Endocrinol. 8, 1298–1308 (1994).
Ste Marie, L., Miura, G. I., Marsh, D. J., Yagaloff, K. & Palmiter, R. D. A metabolic defect promotes obesity in mice lacking melanocortin-4 receptors. Proc. Natl Acad. Sci. USA 97, 12339–12344 (2000).
Chen, A. S. et al. Role of the melanocortin-4 receptor in metabolic rate and food intake in mice. Transgenic Res. 9, 145–154 (2000).
Huszar, D. et al. Targeted disruption of the melanocortin-4 receptor results in obesity in mice. Cell 88, 131–141 (1997).
Sina, M. et al. Phenotypes in three pedigrees with autosomal dominant obesity caused by haploinsufficiency mutations in the melanocortin-4 receptor gene. Am. J. Hum. Genet. 65, 1501–1507 (1999).
Skene, P. J., Henikoff, J. G. & Henikoff, S. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat. Protoc. 13, 1006–1019 (2018).
Lin, C. Y. et al. Whole-genome cartography of estrogen receptor α binding sites. PLoS Genet. 3, e87 (2007).
Lo, L. et al. Connectional architecture of a mouse hypothalamic circuit node controlling social behavior. Proc. Natl Acad. Sci. USA 116, 7503–7512 (2019).
Fuhrmann, F. et al. Locomotion, theta oscillations, and the speed-correlated firing of hippocampal neurons are controlled by a medial septal glutamatergic circuit. Neuron 86, 1253–1264 (2015).
Góis, Z. H. T. D. & Tort, A. B. L. Characterizing speed cells in the rat hippocampus. Cell Rep. 25, 1872–1884 (2018).
Evans, D. A. et al. A synaptic threshold mechanism for computing escape decisions. Nature 558, 590–594 (2018).
Tovote, P. et al. Midbrain circuits for defensive behaviour. Nature 534, 206–212 (2016).
Carter, M. E. et al. Tuning arousal with optogenetic modulation of locus coeruleus neurons. Nat. Neurosci. 13, 1526–1533 (2010).
Coimbra, B. et al. Role of laterodorsal tegmentum projections to nucleus accumbens in reward-related behaviors. Nat. Commun. 10, 4138 (2019).
Balthasar, N. et al. Divergence of melanocortin pathways in the control of food intake and energy expenditure. Cell 123, 493–505 (2005).
Matharu, N. et al. CRISPR-mediated activation of a promoter or enhancer rescues obesity caused by haploinsufficiency. Science 363, eaau0629 (2019).
Slonaker, J. R. The effect of pubescence, oestruation and menopause on the voluntary activity in the albino rat. Am. J. Physiol. 68, 294–315 (1924).
Lotta, L. A. et al. Human gain-of-function MC4R variants show signaling bias and protect against obesity. Cell 177, 597–607 (2019).
Pfaus, J. G., Shadiack, A., Van Soest, T., Tse, M. & Molinoff, P. Selective facilitation of sexual solicitation in the female rat by a melanocortin receptor agonist. Proc. Natl Acad. Sci. USA 101, 10201–10204 (2004).
Clayton, A. H. et al. Bremelanotide for female sexual dysfunctions in premenopausal women: a randomized, placebo-controlled dose-finding trial. Womens Health 12, 325–337 (2016).
Chandler, D. J. et al. Redefining noradrenergic neuromodulation of behavior: impacts of a modular locus coeruleus architecture. J. Neurosci. 39, 8239–8249 (2019).
Duval, K. et al. Effects of the menopausal transition on energy expenditure: a MONET Group Study. Eur. J. Clin. Nutr. 67, 407–411 (2013).
O’Neal, T. J., Friend, D. M., Guo, J., Hall, K. D. & Kravitz, A. V. Increases in physical activity result in diminishing increments in daily energy expenditure in mice. Curr. Biol. 27, 423–430 (2017).
Duggal, N. A., Niemiro, G., Harridge, S. D. R., Simpson, R. J. & Lord, J. M. Can physical activity ameliorate immunosenescence and thereby reduce age-related multi-morbidity? Nat. Rev. Immunol. 19, 563–572 (2019).
Garfield, A. S. et al. A neural basis for melanocortin-4 receptor-regulated appetite. Nat. Neurosci. 18, 863–871 (2015).
Dhillon, H. et al. Leptin directly activates SF1 neurons in the VMH, and this action by leptin is required for normal body-weight homeostasis. Neuron 49, 191–203 (2006).
Krashes, M. J. et al. Rapid, reversible activation of AgRP neurons drives feeding behavior in mice. J. Clin. Invest. 121, 1424–1428 (2011).
Cai, D., Cohen, K. B., Luo, T., Lichtman, J. W. & Sanes, J. R. Improved tools for the Brainbow toolbox. Nat. Methods 10, 540–547 (2013).
Hu, Y. et al. Differential effects of unfolded protein response pathways on axon injury-induced death of retinal ganglion cells. Neuron 73, 445–452 (2012).
Nagai, Y. et al. Deschloroclozapine, a potent and selective chemogenetic actuator enables rapid neuronal and behavioral modulations in mice and monkeys. Nat. Neurosci. 23, 1157–1167 (2020).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Mo, A. et al. Epigenomic signatures of neuronal diversity in the mammalian brain. Neuron 86, 1369–1384 (2015).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137.
Hahne, F. & Ivanek, R. Visualizing genomic data using Gviz and Bioconductor. Methods Mol. Biol. 1418, 335–351 (2016).
Crane, J. D., Mottillo, E. P., Farncombe, T. H., Morrison, K. M. & Steinberg, G. R. A standardized infrared imaging technique that specifically detects UCP1-mediated thermogenesis in vivo. Mol Metab. 3, 490–494 (2014).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Acknowledgements
We thank C. Paillart and T. McMahon for technical assistance with the running and data acquisition for the CLAMS and ANY-maze systems; all members of the Ingraham laboratory for their comments and discussions; O. Yabut for expertise in imaging; and C. Vaisse for insights on MC4R signalling. This work was supported by funding to H.A.I. (R01DK121657, R01AG062331, UCSF Women’s Reproductive Health RAP Award and GCRLE Senior Scholar Award 0320), to W.C.K. (American Heart Association Postdoctoral Fellowship 16POST27260361), to R.R. (K12GM081266 IRACDA Program), to B.G. (2T32GM065094, F31MH124365), to N.M. (UCSF Mary Ann Koda-Kimble Innovation Seed Award, UCSF Catalyst Program), to X.D. (R01EY030138, Research to Prevent Blindness and Klingenstein‐Simons Neuroscience Fellowship), to S.M.C. (K01 DK098320, UCLA Women’s Health Center, UL1TR001881), to C.B.H. (F32 DK107115-01A1, AHA Postdoctoral Fellowship 16POST29870011), to N.A. (R01CA197139, R01 MH109907), to J.T. (R01MH113628, SFARI600568). We acknowledge the mouse metabolic core funded by P30 DK098722-01.
Author information
Authors and Affiliations
Contributions
W.C.K. designed experiments, analysed data and wrote the manuscript. R.R. performed thermal and glucose homeostasis analyses in mice. B.G. optimized, performed and analysed the CUT&RUN method for ERα binding in neurons. N.M. provided CRISPRa viral vectors and expert advice. A.N.R. performed histology experiments and quantification of expression data. A.M.P.-R. aided with chemogenetic data acquisition and analyses. C.B.H. analysed bone and plasma lipid data. S.M.C. designed experiments, provided animal models and analysed data. K.T. and X.D. provided the AAV-DIO-mYFP vector. N.A. provided key unpublished reagents related to CRISPRa constructs and helped to guide studies. J.T. optimized CUT&RUN method for ERα binding in neurons, performed analyses and wrote the manuscript. H.A.I. designed experiments, analysed data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Miguel López and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 VMHvlERαKO does not affect energy intake or physical activity during the light phase but does decrease iBAT thermogenic gene expression.
a, Representative image of a successful hit confirmed post-mortem by loss of ERα expression (red) in the VMHvl (white arrow) with corresponding expression of GFP driven by the AAV2-Cre viral vector. Scale bars, 100 µm. b, Food intake and X-ambulatory parameters obtained in light period for VMHvlErαKO (n = 10) and ARCERαKO (n = 12) female cohorts compared to controls (n = 7 and 5). c, d, Equivalent fat mass in VMHvlERαKO (n = 9), ARCERαKO (n = 11), and control (n = 7 and 6) (c) and oxygen consumption rates in VMHvlERαKO (n = 12), ARCERαKO (n = 11), and control (n = 6 and 6) (d) female mice measured by EchoMRI and indirect calorimetry in CLAMS, respectively. e, Quantification of BAT thermogenic gene expression levels by qPCR in VMHvlERαKO (n = 6), ARCERαKO (n = 7), and control (n = 6 and 3) female mice (VMHvlERαKO Ucp1: unpaired two-tailed Student's t-test, t9 = 2.599, P = 0.0288).
Extended Data Fig. 2 Induction of pS6 and Mc4r in the VMHvl depends on oestrogen signalling through ERα.
a, Immunofluorescence microscopy images of ERα and pS6(S244/S247) in the VMHvl (left panels) and ARC (right panels) of Esr1fl/fl and OVX female mice 4 h post oestradiol benzoate treatment. b, Immunofluorescence microscopy images of ERα (red) and pS6(S244/S247) (green) in the VMHvl in conditional knockout (Esr1Nkx2-1Cre) 4OVX female mice following 4 h after oestradiol benzoate treatment. c, Immunofluorescence microscopy images of ERα (red) and pS6(S244/S247) (green) in the VMHvl of intact adult male mice following 4 h post vehicle or oestradiol benzoate treatment. Bar graph to the right shows increased number of pS6(S244/S247)-positive neurons in the VMHvl of male mice after oestradiol benzoate treatment as done for OVX female mice (unpaired two-tailed t-test, t6 = 8.569, P = 0.0001). d, Mammalian Phenotype Ontology terms most significantly enriched and top five significantly enriched Reactome Pathways among DEGs in the VMHvl (vehicle versus oestradiol benzoate). e, qPCR analysis of the indicated target genes in VMHvl from oestrus female mice, proestrus female mice, and male mice; data points represent values from individual mice (one-way ANOVA Nmur2: F2,15 = 8.469, P = 0.0035, post hoc: oestrus (E) versus proestrus (P) P = 0.0454, proestrus versus male P = 0.0030, and oestrus versus male P = 0.1438; Esr1: F2,11 = 10.18, P = 0.0031, post hoc: oestrus versus proestrus P = 0.1650, proestrus versus male P = 0.0374, and oestrus versus male P = 0.0033). Holm–Šidák multiple comparisons test. f, Mc4r expression levels in VMHvl from OVX oestradiol-benzoate-treated (n = 6) female mice normalized to vehicle treatment (n = 3) (unpaired two-tailed Student's t-test t6 = 6.519, P = 0.0006). g, ISH showing Mc4r expression (red arrows) in the VMHvl of an oestrus female, a proestrus female, and an intact male. h, Representative ISH (Mc4r, red arrows) and immunofluorescent (pS6, yellow arrows) staining in the VMHvl (dashed circle) from OVX female mice treated with vehicle for 4 h, oestradiol benzoate for 2 h, or oestradiol benzoate for 4 h. i, Full size images showing bilateral expression of Mc4r and Rprm in intact female mice staged for oestrus and proestrus. Rprm expression is unchanged in both oestrous stages. Data presented as box plots (see Fig. 1c legend for description). Micrographs are representative of data from 5 mice.
Extended Data Fig. 3 ERα-binding sites in oestradiol-benzoate-sensitive target genes contain conserved ERE consensus sequences.
CUT&RUN CPM-normalized coverage track showing oestradiol-benzoate-specific ERα binding sites containing EREs (pink boxes) within the Greb1 locus (a; 3/3 replicates) and Pgr locus (b; 3/3 replicates), and in the Mc4r promoter (c; 1 of 3 replicates) in 400,000 sub-cortical brain nuclei collected from vehicle and oestradiol benzoate (5 µg) treated gonadectomized mice. Below each track the location and sequence conservation of full (a, b) ERE and half (c) SP1/ERE consensus sites in target gene loci indicated by pink and green boxes. d, e, Location and sequence conservation of ERE consensus sites within Mc4r and Nmur2 loci corresponding to ERα binding sites presented in Fig 1g. For all panels the genomic intervals containing ERE/SP1 sites are located within the ERα binding sites identified by CUT&RUN.
Extended Data Fig. 4 Limited induction of Mc4r expression in the MeA by ERα signalling, and additional neuroanatomical targets of VMHvlMC4R neurons.
a, Representative coronal brain images of Ai14Mc4r female mice stained for ERα (green) and Ai14 (magenta) in the MeA used for quantification shown in Fig 2a. b, Additional ISH comparing Mc4r induction in the VMHvl and MEA in oestrus and proestrus female mice. c, Representative mYFP reporter expression in additional neuroanatomical regions. Scale bars, 200 µm. d, Heat map from Fig. 2d rearranged to compare VMHvlMC4R and VMHvlERα projection intensity in target regions along the anterior-posterior axis. e, VMHvlMC4R projections to the PAG preferentially target the PAGdl/l and PAGdm. Scale bars, 200 µm.
Extended Data Fig. 5 VMHvlMC4R neurons function specifically to drive physical activity.
a, Distance travelled over time in female and male mice following a single injection of CNO (repeated-measures two-way ANOVA interaction effect female: F39,312 = 11.96, P < 0.0001; male: F39,312 = 6.898, P < 0.0001). Holm–Šidák multiple comparisons test. b, Total time spent immobile in intact female and male VMHvlCre− controls (n = 5/4) and VMHvlMC4R::hM3Dq mice (n = 5/5) (repeated-measures two-way ANOVA female interaction effect F1,8 = 33.89, P = 0.0004, post hoc P < 0.0001 and male interaction effect F1,8 = 96.79, P = 0.0005, post hoc P < 0.0001). Holm–Šidák multiple comparisons test. c, Thermal imaging of BAT surface temperatures for VMHvlCre− (left mouse) and VMHvlMC4R:: hM3Dq (right mouse) female mice treated with CL-316,243. d, No differences were observed in Ucp1 mRNA in the BAT from VMHvlCre− and VMHvlMC4R::hM3Dq mice collected 1.5 h after a single CNO injection. e, Body weights for female VMHvlCre− (n = 5) and VMHvlMC4R::hM3Dq (n = 6, baseline measurement includes 1 mouse with mistargeted injection) mice prior to glucose tolerance test (GTT). f, GTT glucose levels for intact female cohorts treated with saline or CNO and total area under the curve (AUC) (unpaired two-tailed Student's t-test t8 = 2.824, P = 0.0223). g, Body weight normalized food consumption in female mice (n = 5/5) following Sal/CNO injection during light dark period (ZT12–ZT17) (repeated-measures two-way ANOVA Dark period interaction effect F1,8 = 3.502, P = 0.0982, post hoc P = 0.0489). Holm–Šidák multiple comparisons test. h, Sustained physical activity increase across light/dark periods in VMHvlMC4R::hM3Dq (n = 5) female mice administered CNO-H2O as compared to VMHvlCre− female mice (n = 5) or during exposure to plain drinking water (H2O). i, j, Cumulative distance travelled (i) and number of rearing events (j) during light/dark periods following administration of CNO or water during the light stage (i repeated-measures two-way ANOVA F1,8 = 15.8, P = 0.0041, post hoc P = 0.0006 and j, repeated-measures two-way ANOVA F1,8 = 15.8, P = 0.0041, post hoc P = 0.0006). Holm–Šidák multiple comparisons test. k, Starting body weights and weight change during continuous administration of CNO-H2O for intact female mice. l, ERα and mCherry expression in the VMHvl following targeted injection of Cre-dependent AAV-hM4Di-mCherry into a female Mc4r-t2a-cre mouse. m, Number of rearing episodes during the dark period in VMHvlMC4R::hM4Di (n = 8) and VMHvlCre− (n = 4) intact female mice following administration of plain or DCZ-laden drinking water. Data are mean ± s.e.m. or box plots (described in Fig. 1c legend).
Extended Data Fig. 6 Increased physical activity and improvement in metabolic health markers in OVX VMHvlMC4R::hM3Dq female mice in response to acute and chronic CNO.
a, Distance travelled over time in OVX female VMHvlCre− (n = 5) and VMHvlMC4R::hM3Dq (n = 5) mice during administration of plain H2O or CNO-H2O (repeated-measures two-way ANOVA interaction effect F11,88 = 5.265, P < 0.0001). b, Total dark period rearing events in intact and OVX female mice administered plain H2O or CNO-H2O drinking water (repeated-measures two-way ANOVA interaction effect F1,8 = 60.31, P < 0.0001 post hoc P < 0.0001). c, Body weights (repeated-measures two-way ANOVA time effect F2,24 = 49.51, P < 0.0001; genotype effect F2,24 = 33.50, P < 0.0001) and fasting glucose levels (repeated-measures two-way ANOVA time effect F2,26 = 6.456, P = 0.0053; genotype effect F1,26 = 10.11, P = 0.0038) in female mice after OVX and subsequent HFD feeding. d, Blood glucose (left) and AUC (right) during ITT in chow-fed OVX female mice following 6-hour fast and saline/CNO treatment. e, Blood glucose (left, repeated-measures two-way ANOVA interaction effect F1,8 = 7.791, P = 0.0235, post hoc: VMHvlMC4R::hM3Dq saline versus CNO, T15 P = 0.0009 and T60 P = 0.0318) and AUC (right, repeated-measures ANOVA with mixed-effects model, note: one cre+ female with missed injection was included in saline but not CNO treated groups, interaction effect F1,8 = 7.791, P = 0.0235, post hoc: VMHvlMC4R::hM3Dq saline versus CNO P = 0.0007) values during 90 min ITT test on HFD-fed OVX female mice performed 6 hours post fasting and post injection with saline or CNO. f, Blood glucose levels following a 6 h fast in OVX female mice maintained on Chow/HFD following a single saline or CNO injection (repeated-measures ANOVA with mixed-effects model, note: one cre+ female with missed injection was included in saline but not CNO treated groups, Chow: treatment effect F1,17 = 5.038, P = 0.0384, post hoc P = 0.0179; and HFD: interaction effect F1,17 = 20.47, P = 0.0019, post hoc P = 0.0073). g, Percentage change in body weight in HFD-fed OVX female mice (n = 5/5) continuously administered CNO-laden drinking water (repeated-measures two-way ANOVA interaction effect F7,64 = 4.583, P = 0.0003. h, Fasting blood glucose levels in OVX/HFD mice before and after 8 days of chronic CNO (repeated-measures ANOVA with mixed-effects model, note: one cre+ female with missed injection was included in saline but not CNO treated groups, interaction effect F1,17 = 5.180, P = 0.0361 post hoc: VMHvlMC4R::hM3Dq Pre versus Post P = 0.0156). i, Plasma cholesterol levels before (Pre) and after (Post) 8 days of continuous CNO-H2O exposure (repeated-measures ANOVA with mixed-effects model, note: one cre+ female with missed injection was included in saline- but not CNO-treated groups, interaction effect F1,8 = 5.502, P = 0.0470, post hoc P = 0.0203). j, Average daily food intake during 8 days of continuous CNO-H2O exposure (points represent separate daily measurements of average consumption per mouse). Data are mean ± s.e.m. or box plots (described in Fig. 1c legend). a–h, ANOVA with Holm–Šidák multiple comparisons test.
Extended Data Fig. 7 Additional metabolic and expression data for conditional Mc4r-rescue mice.
a, FOS expression (arrows) in the VMHvl and PVH of female mice treated with oestradiol benzoate ± MT-II. b, FOS+ cells in oestradiol benzoate + MT-II (n = 5) compared to vehicle (Veh) + MT-II (n = 3, **P = 0.0037) and oestradiol benzoate + vehicle (n = 4, **P = 0.0046) treated female mice (one-way ANOVA F2,9 = 14.00, P = 0.0017). c, Mc4r and Sf1 expression patterns from Genotype-Tissue Expression Project intersect specifically in the hypothalamus (blue arrows) and not in peripheral tissues (red arrows). Transcripts/million (TPM) expression presented as box plots with centre line at median, box edges are 25th and 75th percentiles, and whiskers are 1.5x interquartile range. d, Equivalent body weights within cohorts of female and male Mc4r+/+, Mc4rloxTB and Mc4rSf1-cre mice at weaning. e, Percentage lean (one-way ANOVA F2,31 = 101.4, P < 0.0001, post hoc: Mc4r+/+ versus Mc4rloxTB P < 0.0001, Mc4r+/+ versus Mc4rSf1-cre P < 0.0001, and Mc4rSf1-cre versus Mc4rloxTB P = 0.0720) and % fat (one-way ANOVA F2,31 = 104.2, P < 0.0001, post hoc: Mc4r+/+ versus Mc4rloxTB P < 0.0001, Mc4r+/+ versus Mc4rSf1-cre P < 0.0001, and Mc4rSf1-cre versus Mc4rloxTB P=0.0769) body composition analysis (EchoMRI) in adult female mice of each genotype. f, Oxygen consumption (VO2) as a function of body weight in adult female mice. g, Body weights in 13-week-old Mc4r+/+, Mc4rloxTB, and Mc4rSf1-cre female mice (one-way ANOVA F2,32 = 226.6, post hoc: Mc4r+/+ versus Mc4rloxTB P < 0.0001, Mc4r+/+ versus Mc4rSf1-cre P < 0.0001, and Mc4rSf1-cre versus Mc4rloxTB P=0.0029). Data presented as mean ± s.e.m. or scatterplots of values from individual mice. Number of mice analysed are indicated on each bar. b, d-g, Holm–Šidák multiple comparisons test.
Extended Data Fig. 8 Expression and physical activity levels in male and female Mc4rCRISPRa mice.
a, mCherry expression in Mc4rCRISPRa female hypothalamus. b, Fluorescent ISH images from Mc4rCRISPRa female (left) and male (right) showing Esr1 and Mc4r expression. Images are reproduced and expanded from Fig. 4g to show limited induction of Mc4r outside of the VMHvl target region. c, Dark phase (ZT12–ZT24) physical activity levels (distance per 12 h) as a function of Mc4r or mCherry mRNA expression in microdissected VMHvl from control and Mc4rCRISPRa female mice. d, Home-cage activity in Mc4rCRISPRa (n = 4) and control (n = 3) male mice. e, Time spent immobile during the 12 hour dark phase in control and Mc4rCRISPRa female (unpaired two-tailed Student's t-test, t9 = 2.015, P = 0.0747) and male Mc4rCRISPRa mice (see Fig. 4 for number of mice per group) (unpaired two-tailed Student's t-test, t5 = 3.245, P = 0.0228). f, Mc4rCRISPRa (n = 6) and control (n = 7) female body weights during ad lib feeding. g, Normalized daily food intake in Mc4rCRISPRa (n = 6) and control (n = 7) female mice (unpaired two-tailed Student's t-test, t11 = 2.409, *P = 0.0347). h, BAT surface temperatures in female control (n = 4) and Mc4rCRISPRa (n = 5) mice, repeated measurements at 30- and 60-min post-anaesthesia. i, Cortical bone thickness for female cohorts (unpaired two-tailed t-test, t6 = 2.957, P = 0.0254). j, Body weights in control (n = 4) and Mc4rCRISPRa (n = 6) female mice at wk 9 and at wk 17 after eight weeks of OVX. k, Distance travelled over 24 hours in OVX control and OVX Mc4rCRISPRa compared to intact female mice (blue). l, Total dark phase distance in intact (n = 5), OVX control (n = 7), and OVX Mc4rCRISPRa (n = 8) female mice. Data are presented as scatterplots of values from individual mice, mean ± s.e.m., or as box plots (described in the legend of Fig. 1c).
Supplementary information
Supplementary Video 1
CNO administered in the drinking water stimulates VMHvlMC4R neurons to increase physical activity. Video recording of VMHvlMC4R::hM3dq (top) and VMHvlCre− (bottom) female mice following addition of CNO-laden drinking water (0.25 mg ml−1) during the inactive, lights-on period. Recordings have been sped up 20×.
Rights and permissions
About this article
Cite this article
Krause, W.C., Rodriguez, R., Gegenhuber, B. et al. Oestrogen engages brain MC4R signalling to drive physical activity in female mice. Nature 599, 131–135 (2021). https://doi.org/10.1038/s41586-021-04010-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-04010-3
This article is cited by
-
Artificial ovaries constructed from biodegradable chitin-based hydrogels with the ability to restore ovarian endocrine function and alleviate osteoporosis in ovariectomized mice
Reproductive Biology and Endocrinology (2023)
-
Chemogenetics for cell-type-specific modulation of signalling and neuronal activity
Nature Reviews Methods Primers (2023)
-
The ventromedial hypothalamic nucleus: watchdog of whole-body glucose homeostasis
Cell & Bioscience (2022)
-
Sex-specific multi-level 3D genome dynamics in the mouse brain
Nature Communications (2022)
-
Treating menopause — MHT and beyond
Nature Reviews Endocrinology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.