Abstract
APOE4 is the strongest genetic risk factor for Alzheimer’s disease1,2,3. However, the effects of APOE4 on the human brain are not fully understood, limiting opportunities to develop targeted therapeutics for individuals carrying APOE4 and other risk factors for Alzheimer’s disease4,5,6,7,8. Here, to gain more comprehensive insights into the impact of APOE4 on the human brain, we performed single-cell transcriptomics profiling of post-mortem human brains from APOE4 carriers compared with non-carriers. This revealed that APOE4 is associated with widespread gene expression changes across all cell types of the human brain. Consistent with the biological function of APOE2,3,4,5,6, APOE4 significantly altered signalling pathways associated with cholesterol homeostasis and transport. Confirming these findings with histological and lipidomic analysis of the post-mortem human brain, induced pluripotent stem-cell-derived cells and targeted-replacement mice, we show that cholesterol is aberrantly deposited in oligodendrocytes—myelinating cells that are responsible for insulating and promoting the electrical activity of neurons. We show that altered cholesterol localization in the APOE4 brain coincides with reduced myelination. Pharmacologically facilitating cholesterol transport increases axonal myelination and improves learning and memory in APOE4 mice. We provide a single-cell atlas describing the transcriptional effects of APOE4 on the aging human brain and establish a functional link between APOE4, cholesterol, myelination and memory, offering therapeutic opportunities for Alzheimer’s disease.
Similar content being viewed by others
Main
Late-onset Alzheimer’s disease (AD) accounts for more than 95% of the disease, with approximately 50 million cases worldwide. The heritability of AD is estimated to be between 50% and 80%, suggesting that genetics is the most dominant risk factor after age. Genome-wide association studies have identified genetic variants that are associated with an increased risk of AD1. The E4 allele variant of the APOE gene (APOE4) shows the strongest association with AD1,2,3. One copy of the APOE4 allele, which differs from the major allele (APOE3) by a single amino acid change from Cys112 to Arg112, increases the risk of developing AD three- to fourfold, whereas two copies increase the risk by eight- to twelvefold2. APOE4 is associated with increased amyloid-β deposition, hyperphosphorylation and aggregation of tau, and accelerated cognitive decline4,5,6,7. However, the mechanisms by which APOE4 mediates these effects are not fully understood. The APOE4 allele is present in 40–50% of individuals with AD8; thus, elucidating the molecular and cellular pathways that underlie APOE4-associated pathogenesis could reveal therapeutic opportunities for a large portion of the AD population.
APOE is a lipid and cholesterol transporter. The APOE4 polymorphism alters the accessibility of the lipid-binding region, which interferes with its cholesterol- and lipid-transport ability7. Alois Alzheimer noted lipid inclusions in glial cells when first describing AD9. Recent in vitro studies showed that APOE4 promotes the accumulation of unsaturated triglycerides and lipid droplets in induced pluripotent stem cell (iPS)-cell-derived astrocytes10 and increases cholesterol sequestration in astrocytes leading to changes in the brain matrisome11. Despite these longstanding associations, how cholesterol and lipid abnormalities impair learning and memory in the APOE4 brain, and whether such effects can be mitigated, remains unclear.
Insight into APOE4-mediated pathogenesis is complicated by the fact that APOE is a soluble factor expressed by many cell types12,13. The APOE genotype has a widespread impact on cell-autonomous and non-autonomous biological processes14. To investigate this complexity, we sought to generate an extensive cell-type-specific reference of the molecular processes influenced by APOE4 in the post-mortem human brain. We profiled the prefrontal cortex (PFC) from APOE4 carriers (APOE3/4 or APOE4/4) and non-carriers (APOE3/3) using single-nucleus RNA-sequencing (snRNA-seq) and then complemented this transcriptomic approach with a phenotypic analysis of post-mortem human brain sections and isogenic iPS cell models, and with studies in humanized APOE-knockin mice. Our results reveal that APOE4 (1) alters cholesterol biosynthesis, trafficking and localization in human and mouse oligodendrocytes; and (2) is associated with endoplasmic reticulum (ER) stress and reduced myelination. We establish that pharmacologically facilitating cholesterol transport improves myelination and learning and memory in old APOE4/4 mice, therefore providing new insights into the mechanistic connection between APOE4, myelination and AD pathogenesis.
APOE4 single-nucleus profiling
To determine the effect of APOE4 on gene expression in the aging human brain, we studied a cohort of 32 individuals (12 APOE3/3, 12 APOE3/4 and 8 APOE4/4 carriers) from the Religious Order Study (ROS) or the Rush Memory and Aging Project (MAP), collectively known as ROSMAP15. The APOE3/3 and APOE3/4 subgroups were balanced by AD pathological diagnosis (n = 6 each for positive and negative diagnoses of AD) and gender (3 male and 3 female individuals each; Extended Data Fig. 1a and Supplementary Table 1). All of the individuals with AD presented high levels of both amyloid and neurofibrillary tangle (Tau) pathology, and there were no differences in age or post-mortem intervals (PMIs) between APOE3/3 and APOE3/4 groups and AD status (Extended Data Fig. 1b). All of the available APOE4/4 carriers had a diagnosis of AD, and 5 out of 8 were female. We obtained post-mortem tissue samples extracted from the PFC of each individual and performed snRNA-seq using the 10x Genomics Chromium platform. We report a total of 178,789 (164,741 after quality control) single-nucleus transcriptomes (Extended Data Fig. 1c).
Cellular diversity of the human PFC
Cell types and subtypes were annotated through two rounds of graph-based clustering analysis (Extended Data Fig. 1c). The first round identified excitatory (NRGN+) and inhibitory (GAD1+) neurons (SYT1+), astrocytes (AQP4+), oligodendrocytes (MBP+), oligodendrocyte precursor cells (OPCs) (VCAN+), microglia (CSF1R+) and a heterogeneous population of vascular cell types (Extended Data Fig. 1d). Annotations were supported by expression patterns of curated marker genes16,17 (Extended Data Fig. 1e and Supplementary Table 2). Subclustering analysis of each cell type population identified T cells (CD247+), pericytes (PDGFRB+), endothelial cells (FLT1+), smooth muscle cells (CALD1+) and fibroblasts (ABCA9+) (Extended Data Fig. 1f). A total of 11 cell types supported by gene marker expression and significant enrichment (false-discovery rate (FDR) < 0.01, permutation test) of marker sets (Extended Data Fig. 1g) were annotated. Cell type expression profiles were strongly correlated between individual donors (average Pearson r = 0.94 and r = 0.65 for high- and low-abundance cell types, respectively; Extended Data Fig. 1h,i and Supplementary Table 3). Low-abundance cells (T cells, endothelial cells, pericytes, fibroblasts and smooth muscle cells) were not detected in a small fraction of participants (average fraction across cell types = 0.1) and had relatively low cell counts when detected (9.3 versus 850.2 cells on average across cell types and participants for low- and high-abundance cells, respectively) (Extended Data Fig. 1j,k). High-abundance cell types were well represented across all of the donors, independent of AD diagnosis or genotypic background, with reproducible proportions in most cases (Extended Data Fig. 1l,m). Thus, downstream analysis focused on only high-abundance cell types.
APOE4 pathway-level alterations
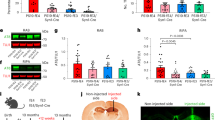
To broadly characterize the potential transcriptomic effects of APOE4 on molecular processes, we considered Gene Ontology biological processes and performed differential pathway activity analysis. We aggregated gene expression values into pathway activity scores and estimated the effect of APOE4 on aggregate scores. Activity scores summarize the overall gene expression levels for genes in a given pathway, improving the power to detect subtle changes in pathway activity (Methods). We compared APOE3/4 and APOE4/4 versus APOE3/3 scores using a multivariate linear model controlling for sex, PMI, amyloid, neurofibrillary tangles and age at death. A total of 486 candidate APOE4-perturbed molecular processes were identified (P < 0.05), including cell-type-specific (404 pathways, affected only in one cell type) and broadly dysregulated (82 pathways, affected in at least two cell types) processes (Fig. 1a and Supplementary Table 4).
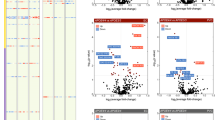
a, Top Gene Ontology biological processes with expression changes associated with APOE4 (nominal P < 0.05, linear model, APOE3/3 versus APOE3/4 and APOE4/4). Red indicates APOE4 upregulation and blue indicates downregulation relative to APOE3. The top 20 pathways are shown in order of P value. Unique alterations indicate evidence (P < 0.05) of pathway alteration in a single cell type. Shared alterations indicate evidence in multiple cell types. Ast, astrocytes; Ex, excitatory neurons; In, inhibitory neurons; Mic, microglia; Oli, oligodendrocytes. Bold text highlights pathways discussed in the Article. b, APOE-associated pathways with expression changes associated with APOE4 (nominal P < 0.05, linear model, APOE3/3 versus APOE3/4 and APOE4/4). Evidence of alteration (P < 0.05) is indicated (X). Pathways were manually classified into three categories (colour key). c, Brain-specific lipid-associated pathways (Methods) with APOE4-associated expression changes (nominal P < 0.05, linear model, APOE3/3 versus APOE3/4 and APOE4/4).
APOE4-dysregulated processes included upregulation of inflammatory and immune-related pathways in several cell types, including NF-кB signalling in excitatory neurons, inhibitory neurons and OPCs; T cell receptor and cytokine signalling in astrocytes; and tumour-necrosis-factor-mediated signalling in microglia and excitatory neurons (Fig. 1a). We also observed downregulation of synaptic-related processes, including ion channel activity, excitatory postsynaptic potential and synaptic plasticity (Fig. 1a). Notably, these alterations not only occurred in neurons (for example, voltage-gated calcium channel activity), but also in oligodendrocytes (for example, long-term neuronal synaptic plasticity). In APOE4 excitatory neurons, we identified upregulation of ERK1/2 signalling (such as MAPK1, MAP2K1 and MAP2K2) and endosomal-related transport (such as RAB21 and SNX12) genes (Supplementary Table 4). All of these effects are consistent with the reported roles of APOE4 in inflammation, synaptic impairment, MAPK1 signalling and downstream APP transcription and early endosomes7,18. We also identified decreased glycosyltransferase activity (for example, ALG6, ALG3) in APOE4 neurons and widespread alterations to amyloid-β metabolism genes in excitatory neurons, oligodendrocytes and OPCs (amyloid-β formation, for example, BIN1, ROCK1; Fig. 1a). DNA damage response was also altered in APOE4 microglia and OPCs, suggesting APOE4 may promote DNA damage in specific cell types. Altered amyloid-β processing and DNA damage have been associated with APOE4 and AD19,20,21.
Finally, APOE4 was associated with cellular stress and energy metabolism in multiple cell types. Chaperone-mediated protein folding was increased in most cell types (for example, DNAJB14 and HSPA1A), and ATF-mediated unfolded protein response was increased in neurons and oligodendrocytes. Cholesterol biosynthesis was increased in APOE4 oligodendrocytes (for example, DHCR24 and LBR), lipid storage in OPCs (such as PPARA) and glycogen metabolism in microglia and astrocytes (such as UGP2 and PGM1), whereas acetyl-CoA metabolism was decreased in astrocytes (for example, ACAT1 and ACSS1; Fig. 1a and Supplementary Table 4). Thus, dysregulation of energy production, transport and use may be central pathogenic mechanisms of APOE4.
Overall, our analysis confirms established APOE4 neurobiology7, reveals disturbances and assigns cell type specificity to known APOE4-associated alterations. A cell-type-specific molecular atlas of APOE4 pathway-level effects is provided in Supplementary Tables 4–6.
APOE4 affects lipid pathways in the brain
We next performed a targeted pathway analysis focusing on specific molecular pathways that are probably affected by APOE4 by integration of previous literature (Methods). We created a curated database of 193 brain-expressed and APOE-related pathways comprising 8,759 unique genes (Extended Data Fig. 2a). Most pathways exhibited cell-type-specific activity, yet a subset was active across all major cell types, suggesting the potential for both cell-type-specific and broad effects of APOE alterations (Extended Data Fig. 2b). By comparing pathway activity scores of APOE3/4 and APOE4/4 versus APOE3/3, we found that the APOE4 genotype affects 22 out of the 193 pathways (Fig. 1b and Supplementary Table 5). As expected, associated pathways included amyloid-β and protein processing, lipid metabolism and synaptic-related processes. However, additional alterations in cholesterol efflux and transport were recovered by this targeted analysis. Given the reported impairment of APOE4’s lipid-transport function22, and our finding of associations with lipid metabolism (Fig. 1a,b), we performed a third, more narrowly targeted analysis focusing on only brain-specific lipid-related processes. We reasoned that this could uncover molecular processes that mediate the effects of APOE4. This analysis further prioritized 17 lipid-related processes affected by APOE4, revealing cell-type-specific alterations in cholesterol, steroid, phospholipid, glycolipid, fatty acid and triglyceride metabolism (Fig. 1c and Supplementary Table 6).
APOE4 alters cholesterol in oligodendrocytes
In APOE4 oligodendrocytes, genes encoding canonical enzymes associated with cholesterol biosynthesis (such as DHCR24, HMGCS1, SQLE and MVK) were confirmed to be upregulated by differential expression and gene set enrichment analysis (Fig. 1a–c and Extended Data Fig. 2c,d). Our previous single-cell analysis uncovered AD-associated oligodendrocyte subpopulations, highlighting a role of myelin-related processes in AD pathogenesis12, consistent with reports of hypomyelination and white-matter changes in AD23,24. Given this emerging association, our experimental follow-up focused on investigating the relationship between APOE4 and cholesterol in oligodendrocytes.
APOE4 was associated with increased expression of cholesterol-related genes in oligodendrocytes in a dose-dependent manner (APOE4/4 > APOE3/4 > APOE3/3, P = 0.01, Pearson’s correlation; Fig. 2a), similar to APOE4’s dose-dependent effect on AD risk, suggesting that APOE4 modulation of cholesterol in oligodendrocytes may contribute to AD. However, cholesterol biosynthesis gene expression in APOE3/3 individuals displayed a bimodal trend, with higher expression in AD (Fig. 2a), suggesting potential modulation of cholesterol pathways by AD pathology. To test whether the effects of APOE4 were independent of the effects of AD, we performed pathology-stratified analyses focusing on lipid-associated pathways. Cholesterol biosynthesis gene expression increased in individuals without AD pathology (Extended Data Fig. 2e,f; lipid-storage pathways were also increased) and there was a non-significant increase in individuals with AD (Extended Data Fig. 2g), indicating an APOE4 effect independent of AD pathology. Individuals with AD versus control individuals also exhibited similar increases when controlling for APOE3/3 and APOE3/4 backgrounds, suggesting an effect of AD pathology independent of APOE4 (Extended Data Fig. 2h–j). Notably, the combined effect of the APOE4 genotype and AD pathology appeared to be additive. Individuals with APOE3/3, with and without pathology, exhibited the lowest cholesterol biosynthesis gene expression, and those with both APOE4 and AD pathology showed the highest cholesterol biosynthesis gene expression (Extended Data Fig. 2k). Together, this suggests that APOE4 has a pathology-independent effect on lipid and cholesterol metabolism and that increased cholesterol biosynthesis might be a convergent pathway of both APOE4 and AD pathology.
a, Dose-dependent association between APOE4 and the aggregated expression of cholesterol biosynthesis genes in post-mortem oligodendrocytes (APOE4/4 > APOE3/4 > APOE3/3, Pearson correlation coefficient (pcc) = 0.45, P = 0.01, two-sided). b, The curation process of cholesterol-related genes. Pathway databases were filtered using the terms cholesterol/lipid storage, transport or synthesis. BP, biological processes. c, Cholesterol-related genes differentially expressed in APOE3/3 versus APOE3/4 and APOE4/4 post-mortem oligodendrocytes (snRNA-seq data, FDR-adjusted P < 0.05, negative binomial mixed model). d, The detection levels of four cholesteryl esters quantified by MS analysis of post-mortem human corpus callosum from APOE3 individuals (APOE3/3, female, n = 3) and APOE4 carriers (APOE4/4, female, n = 3). Data points represent the relative abundance per individual. n = 3 individuals per genotype. The numbers at the top label cholesteryl ester species, specifying the carbon atoms in tail:number of double bonds in tail. P values were calculated using two-sided unadjusted Wilcoxon tests (all Wilcoxon tests are Wilcoxon rank-sum tests). Box plots show the median (centre line), and the interquartile range (IQR) defines the box. The upper whisker extends to the largest value no further than 1.5× IQR from the end of the box. The lower whisker extends to the smallest value at most 1.5× IQR from the end of the box. e, Representative images of BODIPY–cholesterol staining and anti-myelin basic protein (MBP) immunoreactivity in the human post-mortem PFC (BA10) from APOE4 carriers compared with APOE3/3 individuals (left). Scale bar, 10 μm. Cholesterol localization was analysed for four individuals for each APOE genotype. Right, the total BODIPY–cholesterol signal and the percentage of BODIPY–cholesterol signal within 1 μm of an MBP-positive axon quantified using Imaris. Data points represent individuals. Data are mean ± s.e.m. P values were calculated using unpaired two-tailed Student’s t-tests.
APOE4 raises cholesteryl esters in the human brain
Genes associated with cholesterol and lipid synthesis, storage or transport were differentially expressed in APOE3/4 and APOE4/4 versus APOE3/3 post-mortem oligodendrocytes (FDR < 0.05, negative binomial mixed model; Fig. 2b,c). Genes associated with cholesterol biosynthesis and droplet formation (DHCR24, LPIN2, IRS2, NR1H2, LBR, LPIN1 and MBTPS1) were upregulated, whereas genes associated with cholesterol transport were downregulated (PCYT1B, SEC23A, PRKN, SCP2 and LPCAT3; Fig. 2c). To investigate the abundance of cholesterol species in APOE3/3 individuals and APOE4 carriers, we performed mass spectrometry (MS)-based lipidomic profiling on frozen post-mortem corpus callosum tissue from individuals with APOE4/4 (all female; n = 3) and APOE3/3 (all female; n = 3). Four species of cholesteryl esters, a form of cholesterol associated with storage in lipid droplets, were detected in the human corpus callosum (20:4, 18:2, 22:6 and 18:1 (carbon atoms in tail:number of double bonds in tail); Fig. 2d). Corpus callosum from individuals with APOE4/4 exhibited a trend towards higher levels of all four detected cholesteryl ester species relative to non-carriers (APOE3/3, all female, P ≥ 0.7), suggesting that cholesteryl ester may be elevated in APOE4 brain tissue.
We further analysed a separate lipidomic dataset generated from the PFC of 314 individuals, including APOE3/3 and APOE4 individuals with and without AD pathology. Although this larger dataset was not designed to specifically measure cholesterol species, we detected evidence of increased cholesteryl ester species (CH18:1) in APOE4 carriers (15, APOE3/4 and APOE4/4 male and female individuals) versus non-carriers (105, APOE3/3 male and female individuals) in the absence of AD pathology (P = 0.046, Wilcoxon rank-sum test) (Extended Data Fig. 3a–d). Although additional well-powered lipidomic analyses designed to quantify cholesteryl esters in post-mortem tissue are needed, these further support an association between APOE4 and brain cholesterol, potentially indicating that there is increased cholesterol storage in lipid droplets (Supplementary Tables 7 and 8).
We found that APOE4 carriers have decreased localization of cholesterol along neurofilaments and increased intracellular accumulation or storage of cholesterol compared with brains from age-matched APOE3/3 individuals. We stained post-mortem PFC samples from individuals with late-stage AD who are APOE4 carriers (APOE3/4; n = 4) and non-carriers (APOE3/3; n = 7) with BODIPY–cholesterol and antibodies against the lipid-droplet-associated protein PLIN1. BODIPY–cholesterol and PLIN1 staining exhibited a high degree of overlap, indicating the accumulation of BODIPY–cholesterol in lipid droplets (Extended Data Fig. 4a). Punctate, perinuclear PLIN1-stained lipid droplets appeared in both APOE3/3 and APOE3/4 tissue; staining was more prominent around distinct nuclei in APOE4 carriers (Extended Data Fig. 4a). In APOE3/3 PFC tissue, BODIPY–cholesterol staining was instead most intense along neurofilament tracts, where it formed ribbon-like staining patterns adjacent to neurofilament and MBP staining (Fig. 2e). In the white-matter tracts from APOE3/3 PFCs, 59% of BODIPY–cholesterol was within 1 μm of neurofilament staining (Fig. 2e). In APOE4 carriers, the localization of BODIPY–cholesterol along neurofilaments significantly (P = 0.0106) decreased from 58% to only 19% of the total signal (Fig. 2e). APOE4 carriers also exhibited a significantly (P = 0.0220) higher mean intensity of BODIPY–cholesterol staining across multiple PFC sections, indicating an increase in overall cholesterol (Fig. 2e). To further corroborate altered localization of cholesterol, we performed transmission electron microscopy (TEM) analysis of the corpus callosum of six-month-old APOE3/3 and APOE4/4 targeted-replacement (APOE-TR) mice, quantifying the number of lipid droplets per μm2 surface area. APOE4/4-TR mice had a significantly (P = 0.0048) increased density of small lipid-droplet-like structures in the corpus callosum compared with age-matched APOE3/3-TR mice (Extended Data Fig. 4b), suggesting that APOE4 brains have increased lipid and cholesterol storage.
Our single-nucleus transcriptomic analysis revealed altered cholesterol biosynthesis, storage and transport in APOE4 oligodendrocytes and, to a lesser extent, in microglia and astrocytes (Fig. 1a–c). To determine whether these cell types contributed to the observed differences in cholesterol accumulation, we co-stained PFC white matter tissue from APOE4 carriers with BODIPY–cholesterol and cell-type-specific markers for oligodendrocytes (OLIG2), astrocytes (GFAP) and microglia (IBA1). IBA1+ microglia and GFAP+ astrocytes exhibited minimal (less than 15% of cells) BODIPY–cholesterol accumulation (Extended Data Fig. 4c). By contrast, the area within a 2 μm radius of OLIG2+ nuclei exhibited significantly (P < 0.0001) more BODIPY–cholesterol staining, with 77% of OLIG2+ nuclei surrounded by intense BODIPY–cholesterol signal, suggesting that cholesterol may aberrantly accumulate in APOE4 oligodendrocytes (Extended Data Fig. 4c). PLIN1 immunoreactivity around OLIG2+ nuclei in APOE3/4 human PFC from individuals with AD also exhibited a minimal but non-significant increase compared with APOE3/3 (P = 0.1521, n = 3 per genotype) (Extended Data Fig. 4d,e). These findings indicate that cholesterol accumulates in APOE4 oligodendrocytes.
APOE4 affects cellular storage of cholesterol
Genetic diversity may confound the effects of APOE4 observed on human post-mortem brain tissue. We therefore used isogenic APOE3/3 and APOE4/4 iPS cells generated by CRISPR–dCas9 editing14. In brief, an iPS cell line from a cognitively normal APOE3/3 individual was edited to APOE4/4 homozygous. Using a reciprocal strategy, an iPS cell line from a APOE4/4 patient with AD was also edited to APOE3/3. These isogenic iPS cell sets were simultaneously differentiated into oligodendroglia using established protocols (Methods). Similar to human post-mortem oligodendrocytes, iPS-cell-derived oligodendroglia transcribed mRNAs and expressed proteins specific to oligodendrocytes, including MOG, MBP, PLP1, PLLP and MYRF (Extended Data Fig. 5a,b). After completing the differentiation protocol, 65–81% of cells were immunoreactive for MBP and 69–82% were immunoreactive for PLP1 across cultures (Extended Data Fig. 5c). To further validate iPS-cell-derived oligodendroglia, gene expression was compared with pseudobulk signatures of major cell types from snRNA-seq analysis of the post-mortem human brain. Principal component analysis confirmed that iPS-cell-derived oligodendroglia cluster with post-mortem human oligodendrocytes and OPCs (Extended Data Fig. 5d and Supplementary Table 9), and a pairwise comparison of cells identified that iPS-cell-derived oligodendroglia have the most similarity to OPCs and oligodendrocytes (Extended Data Fig. 5e and Supplementary Table 9). Similar to in vivo human oligodendrocytes, iPS-cell-derived oligodendroglia exhibited robust upregulation of myelin-associated (Extended Data Fig. 5f) and cholesterol-associated (Extended Data Fig. 5g,h and Supplementary Tables 9 and 10) genes. APOE mRNA and protein were expressed at low to moderate levels in iPS-cell-derived oligodendroglia, with no significant (P = 0.2635) difference between isogenic APOE3/3 and APOE4/4 iPS-cell-derived oligodendroglia (Extended Data Fig. 5i,j). Post-mortem oligodendrocytes had low but detectable APOE expression, consistent with previous single-cell resolution reports25 (Extended Data Fig. 5k,l). This demonstrates that iPS-cell-derived oligodendroglia are transcriptionally similar to ex vivo human oligodendrocytes and OPCs, providing a reliable proxy for investigating the effect of APOE4 on human oligodendrocytes.
MS-based lipidomic profiling of APOE3/3 and APOE4/4 isogenic iPS-cell-derived oligodendroglia detected 88 lipid species that were increased (adjusted P < 0.05) in APOE4/4 oligodendroglia compared with APOE3/3 controls, and only one species decreased (Fig. 3a, Extended Data Fig. 6 and Supplementary Table 11). In agreement with the lipidomic analysis of post-mortem human brain, cholesteryl esters were the most abundantly altered lipid class, with 15 species increased in APOE4/4 oligodendroglia compared with APOE3/3 (adjusted P < 0.05; Fig. 3a). We also observed changes to phosphatidic acids, diglycerides, triglycerides and phosphatidylcholines; suggesting that APOE4/4 broadly alters cellular lipid composition. Notably, diglycerides and triglycerides are major components of lipid droplets, suggesting an increase in lipid droplet biosynthesis in APOE4/4 oligodendrocytes. We also observed an increase in lysophosphatidylcholine and its derivative lysophosphatidic acid, two lipid classes that are known to promote brain inflammation and demyelination26 (Extended Data Fig. 6 and Supplementary Table 11). SOAT1—which encodes the enzyme responsible for cholesterol esterification—was differentially expressed in isogenic APOE4/4 versus APOE3/3 oligodendroglia (Extended Data Fig. 7a). SOAT2 was not detected in iPS-cell-derived oligodendroglia for either genotype. Similarly, CYP46A1, a hydroxylation enzyme that facilitates brain cholesterol trafficking and clearance, was downregulated in APOE4/4 compared with APOE3/3 oligodendroglia (Extended Data Fig. 7a and Supplementary Table 12), suggesting that cholesterol membrane trafficking in APOE4/4 oligodendroglia may be reduced.
a, Detected lipid-species concentrations from MS-based lipidomic profiling of iPS-cell-derived APOE3/3 and APOE4/4 oligodendroglia. Cholesteryl ester species are highlighted in yellow. The number of differentially (adjusted P < 0.05) detected lipid species for each lipid class is shown. Cholesteryl esters are the most frequently differentially detected class, with 15 species upregulated in APOE4 oligodendroglia. b, Representative images of filipin (cholesterol) and WGA-membrane staining in APOE3/3 and APOE4/4 iPS-cell-derived oligodendroglia. The arrows highlight altered cholesterol localization in APOE4/4 iPS-cell-derived oligodendroglia. Filipin intensity was quantified for the cell membrane (localized with WGA) and intracellular compartment (between membrane and nucleus; n = 6 replicates from independent experiments). Scale bars, 50 μm. Data are mean values from independent biological replicates. P values were calculated using unpaired two-tailed Student’s t-tests. c, Representative images of APOE3/3 and APOE4/4 iPS-cell-derived BODIPY–cholesterol-stained oligodendroglia. Data are mean values from independent biological replicates (n = 4 per genotype). P values were calculated using unpaired two-tailed Student’s t-tests. Scale bar, 24 μm. d, Representative images of APOE4/4 iPS-cell-derived oligodendroglia after addition of 1 μg ml−1 BODIPY–cholesterol to the cell culture medium of live cells for 2 h. Cells were counterstained for markers of the endosome (EEA1), lysosome (LAMP1) and ER (calreticulin). The percentage of BODIPY–cholesterol particles overlapping with EEA1 (n = 5), LAMP1 (n = 5) or calreticulin (n = 4) immunoreactivity was quantified using Imaris. SAD, sporadic AD. Scale bars, 24 μm. Data are mean values. Data points represent independent biological replicates. P values were calculated using one-way analysis of variance (ANOVA).
To further characterize APOE4-mediated differences in cholesterol storage, we stained isogenic APOE3/3 and APOE4/4 oligodendroglia with the cholesterol dye filipin, BODIPY–cholesterol and lipid-droplet-associated PLIN1. In APOE3/3 oligodendroglia, the majority (63 ± 12%) of filipin staining accumulated around the membrane, co-localizing with WGA-membrane staining (Fig. 3b). The remaining filipin signal (37 ± 13%) accumulated intracellularly in APOE3/3 oligodendroglia (Fig. 3b). Isogenic APOE4/4 oligodendroglia exhibited the opposite staining pattern, with 79% of filipin staining appearing intracellular and 22% localized to the membrane (Fig. 3b). To assess cholesterol uptake and intracellular trafficking, we added fluorescent BODIPY–cholesterol to the oligodendroglia cell culture medium. Similar to the post-mortem human brain, BODIPY–cholesterol staining co-localized with PLIN1 immunoreactive areas (Extended Data Fig. 7b). APOE4/4 oligodendroglia exhibited approximately tenfold higher BODIPY–cholesterol staining (P = 0.0253 and 0.0386) than isogenic APOE3/3 controls across two isogenic sets generated from different individuals (Fig. 3c). Both sets of APOE4/4 oligodendroglia also contained more BODIPY droplets per cell (P = 0.0345 and P = 0.027) compared with the APOE3/3 controls, suggesting that there is an increased number of lipid droplets in APOE4/4 oligodendroglia (Extended Data Fig. 7c). APOE4 oligodendroglia exhibited significantly (P = 0.0069) more BODIPY–cholesterol co-localized with Lysotracker-Red staining compared with APOE3/3 oligodendroglia, suggesting that there is lysosomal accumulation of cholesterol in APOE4 oligodendroglia (Extended Data Fig. 7d). However, the increased BODIPY–cholesterol in APOE4 oligodendroglia was not entirely localized to lysosome, suggesting that cholesterol may be accumulating in other organelles. These results demonstrate that APOE4 oligodendroglia have increased intracellular cholesterol storage, confirming and extending results seen in post-mortem human PFC.
Cholesterol localizes in the ER causing stress
Accumulation of lipid droplets is a common hallmark of ER stress27,28. Post-mortem human APOE4/4 oligodendrocytes exhibited upregulation of the ATF6-mediated ER stress pathways compared with APOE3/3 controls with increased expression of multiple heat-shock proteins and of ATF6—a key regulator of the unfolded protein response. (Extended Data Fig. 7e and Supplementary Table 4). Immunohistochemistry analysis of ATF6 showed increased nuclear ATF6 in APOE4/4 oligodendroglia compared with isogenic APOE3/3 controls (Extended Data Fig. 7f). Similarly, genes associated with unfolded protein response (CALR, HSP90B1 and HSPA5) were upregulated in APOE4/4 oligodendroglia compared with APOE3/3 isogenic controls (Extended Data Fig. 7g). Concurrently, genes associated with ER cholesterol transport (PCYT1B, SEC23A, SCP2 and LPCAT3) were downregulated in APOE4/4 oligodendrocytes (Fig. 2c), suggesting that ER stress and impaired cholesterol transport could have a central role in the observed intracellular cholesterol accumulation. To examine this possibility, we added BODIPY–cholesterol to the medium of iPS-cell-derived APOE4/4 oligodendroglia and counterstained for canonical markers of the endosome (EEA1), lysosome (LAMP1) and ER (calreticulin). Approximately 2% of BODIPY–cholesterol particles localized to the endosome, around 18% to the lysosome and about 80% to the ER, indicating that the majority of intracellular cholesterol accumulating in APOE4/4 oligodendroglia is retained in the ER (Fig. 3d). Together, these results demonstrate that APOE4 is associated with reduced localization of cholesterol in the plasma membrane of oligodendrocytes, increased cholesteryl esters and lipid droplets, and upregulation of ATF6-mediated ER stress response.
Cholesterol and APOE4 affect myelination
Oligodendrocytes myelinate the central nervous system and the bioavailability of cholesterol is rate-limiting for the formation and maintenance of myelin29. We reasoned that altered cholesterol localization and homeostasis in APOE4 oligodendrocytes may impair myelination. Compared with APOE3/3 post-mortem human oligodendrocytes, APOE3/4 and APOE4/4 post-mortem human oligodendrocytes exhibited downregulation of myelin-associated genes (PLLP, MYRF, MAG, OPALIN, MOG and PLP1; Fig. 4a and Supplementary Table 13). This pattern persisted in cells from individuals without AD pathology, suggesting that these myelin-associated genes may be influenced by the APOE genotype and not confounded by pathology (Extended Data Fig. 8a and Supplementary Table 13). Decreased myelin-associated gene expression occurred concomitantly with increased expression of genes associated with cholesterol homeostasis, including MVK, FDPS, ABCG1, IDI1, LDLR, INSIG1, SREBF2, SQLE, DHCR7, DHCR24, FDFT1 and LSS (Fig. 4a).
a, log2[mean gene expression in APOE3/4 and APOE4/4/mean gene expression in APOE3/3] for differentially expressed (adjusted P < 0.05, Wilcoxon test computed by wilcoxauc(); Methods) myelin-associated or cholesterol-associated genes in human post-mortem oligodendrocytes. b, Black Gold II staining area μm2 of myelinated axons in the PFC (BA10) of APOE3/3 and APOE3/4 individuals. n = 3 individuals per genotype. The area positive for Black Gold II staining was quantified using ImageJ, with the same intensity thresholds for each image and group. Scale bar, 250 μm. Data points represent individuals, and the bars represent the mean value per genotype. P values were calculated using unpaired two-tailed Student’s t-tests. c, MBP and neurofilament (SMI311) immunoreactivity in the PFC (BA10) from APOE4 carriers and non-carriers. n = 6 individuals per genotype. The area positive for MBP and neurofilament was quantified using ImageJ with the same intensity thresholds for each image and group. Three images were quantified for each individual. Data points represent individuals mean values, and the bars represent the mean value per genotype. P values were calculated using unpaired two-tailed Student’s t-tests. Scale bar, 50 μm. d, TEM analysis of sections from human corpus callosum of APOE4 carriers (n = 3 individuals, n = 58 axons) and APOE3/3 individuals (n = 3 individuals, n = 44 axons). Scale bars, 500 μm. The g-ratio was calculated using ImageJ, measuring the inner axonal diameter divided by the diameter of outer myelin band. Data points represent axons. P values were calculated using Wilcoxon tests. e, TEM analysis of the corpus callosum of APOE3/3-TR (n = 3) and APOE4/4-TR (n = 3) mice at 6 months of age. The g-ratio was quantified using ImageJ as described in d. Scale bars, 500 μm. Data points represent axons. P values were calculated using Wilcoxon tests. The number of myelinated axons per image was counted using the Cell Counter tool in ImageJ. Data points represent the average per animal, and the bars represent the mean value per genotype. P values were calculated using unpaired two-tailed Student’s t-tests.
To examine whether reduced myelin-associated gene expression in APOE4 carriers translated to decreased myelin, we used Black Gold II staining to visualize myelinated axons in post-mortem human PFC tissue from individuals with APOE3/4 (n = 3) compared with APOE3/3 (n = 3). APOE3/3 human PFC showed multiple dense, parallel, axonal tracks suggestive of robust levels of myelination (Fig. 4b). By contrast, PFC from APOE3/4 had a significant (P = 0.0031) reduction in the area positive for Black Gold II staining, and axonal fasciculation was largely absent, suggesting a decrease in axonal myelination (Fig. 4b). To further compare myelination between APOE4 carriers and APOE3/3 individuals, we immunostained PFC tissue with two different anti-MBP antibodies using two separate cohorts of post-mortem human tissue. In APOE3/3 carriers, MBP staining was robust and closely associated with neurofilaments that largely formed parallel tracts of axons (Fig. 4c and Extended Data Fig. 8b). By contrast, PFC white matter regions from APOE4 carriers exhibited significantly (P = 0.0027, P = 0.0294) decreased MBP immunoreactivity compared with APOE3/3 non-carriers, with fewer MBP regions surrounding neurofilament-positive axons (Fig. 4c and Extended Data Fig. 8b). Together, these results suggest that there are reduced myelin levels in the brains of humans carrying APOE4.
To assess ultrastructural myelin differences and integrity, we performed TEM analysis of human post-mortem corpus callosum samples from APOE4/4 (n = 3; mean age, 77 ± 7 years; male; mean PMI, 24 ± 15 h) and APOE3/3 (n = 3; mean age, 86 ± 2 years; male and female; mean PMI, 29 ± 8 h) individuals with AD pathology. Individuals with APOE4/4 had reduced levels of axonal myelination relative to individuals with APOE3/3, as indicated by fewer and thinner dark electron-dense bands encircling neuronal axons (Fig. 4d). To quantify the relative myelin levels per axon, we calculated the g-ratio (the inner axonal diameter divided by the outer diameter with myelin sheath) for 150 neurons per condition. APOE4 carriers had higher g-ratios (P < 0.0001), suggesting that the corpus callosum from older individuals with APOE4/4 contains fewer myelinated axons and thinner overall myelin sheaths compared with individuals with APOE3/3 (Fig. 4d).
To control for covariates inherent to post-mortem human samples, we also quantified myelin levels in inbred APOE-TR mice (9 months old, male and female mice; n = 4 per genotype). The hippocampus from APOE4/4-TR mice exhibited less MBP immunoreactivity (P = 0.044), suggesting reduced myelin levels compared with APOE3/3-TR mice (Extended Data Fig. 8c). Western blot analysis of lysates prepared from mouse cortex revealed reduced total MBP protein levels (P = 0.0112) in APOE4/4-TR mice compared with APOE3/3-TR control mice (Extended Data Fig. 8d). TEM analysis of the corpus callosum of 6-month-old APOE3/3-TR mice showed densely packed myelinated axons, with an average of 170 myelinated axons per field of view. APOE4/4-TR mice had fewer myelinated axons (P = 0.0265, 140 myelinated axons in average per field of view, n = 3 mice) and higher g-ratios compared with APOE3/3-TR mice (P < 0.0001; Fig. 4e). To investigate whether the reduced myelination in APOE4/4-TR mice is an age-related phenotype, we performed TEM analysis of young two-month-old mice. Similar to six-month-old mice, two-month-old APOE3/3-TR mice exhibited densely packed myelinated axons, whereas two-month-old APOE4/4-TR mice exhibited a more heterogeneous and sparse distribution of myelinated axons, reflected by an increased g-ratio (Extended Data Fig. 8e). Together, these findings suggest that APOE4/4-TR mice have a reduced number of myelinated axons and thinner myelin sheaths compared with APOE3/3-TR mice—a phenotype that may precede age-related neurodegeneration.
A key question is whether APOE4 expression in oligodendrocytes is sufficient to decrease myelination or requires the involvement of additional cells and/or factors. To investigate this, we used an in vitro myelination assay using isogenic iPS-cell-derived oligodendroglia co-cultured with NGN2-induced neurons derived from the same iPS cells (iNeurons) in a three-dimensional extracellular matrix that enables cells to interact with each other and self-assemble into tissue-like structures (Extended Data Fig. 9a). After 2 weeks in these 3D co-cultures, MBP-positive oligodendroglia spread and engaged neurofilament-positive axons (Extended Data Fig. 9b). After 6 weeks in culture, myelin-associated proteins O4 and MBP encircled neurofilaments, mimicking biological processes critical to myelination (Extended Data Fig. 9c). GFP-labelled oligodendroglia cells aligned with neurofilament-positive axons in co-cultures containing iNeurons (Extended Data Fig. 9d). High-magnification imaging confirmed that the cells surrounding neurofilaments were positive for MBP (Extended Data Fig. 9e and Supplementary Video 1), and TEM showed axonal segments surrounded by electron-dense membrane rings at a low frequency, suggesting the presence of myelinated axons (Extended Data Fig. 9f). Given that these co-cultures exhibit key components of myelination, such as axonal engagement and canonical marker expression, we sought to use this model to assess the effect of APOE4 on myelin-associated phenotypes in human cells.
We established oligodendroglia–iNeuron co-cultures from two different APOE4/4 and APOE3/3 isogenic sets of iPS-cell-derived cells. MBP-immunoreactive areas sporadically appeared after 3 weeks in APOE3/3 co-cultures (Extended Data Fig. 9g). After 6 weeks, both APOE3/3 and APOE4/4 co-cultures contained similar (P > 0.9999) levels of neurofilament-positive signal, indicating a similar number of axons (Extended Data Fig. 9h). APOE3/3 co-cultures exhibited robust MBP immunostaining with approximately 60% of MBP signal surrounding (within 1 μm) neurofilament-positive axons (Fig. 5a). However, both sets of isogenic APOE4/4 co-cultures exhibited significantly (P = 0.0003 and P = 0.0193) less (<20%) MBP staining localized with neurofilament, suggesting that APOE4/4 oligodendrocytes were producing less MBP and engaging less with neuronal processes than isogenic APOE3/3 co-cultures (Fig. 5a).
a, Representative co-culture images of isogenic iPS-cell-derived APOE4/4 and APOE3/3 oligodendroglia and NGN2-induced neurons after 6 weeks of culture. n = 5 biological replicates. Scale bar, 50 μm. Co-cultures were prepared using two different isogenic iPS cell sets created with reciprocal editing strategies from different individuals. MBP immunoreactivity localized within 1 μm of neurofilament immunoreactivity was quantified using Imaris. Data points represent mean values (n = 4 images, independent biological replicates). Bars represent group mean values. P values were calculated using unpaired two-tailed Student’s t-tests. I, isogenic; P, parental. b, Genetic mix-and-match experiment in which oligodendrocytes and neurons were co-cultured under the following four permutations (oligodendroglia with iNeuron): APOE3/3 with APOE3/3; APOE3/3 with APOE4/4; APOE4/4 with APOE4/4; and APOE4/4 with APOE3/3. n = 5 biological replicates. The percentage of MBP immunoreactivity localized within 1 μm of neurofilament staining was quantified using Imaris. Representative images of MBP, neurofilament and Hoechst staining. Scale bar, 50 μm. Data points represent the mean. n = 4 four images, independent experiments. Axonal MBP was quantified using ImageJ using the same thresholding settings for each image and group. P values were calculated using one-way ANOVA with Bonferroni correction.
The reduction in myelination associated with APOE4 could arise from autonomous defects in oligodendrocytes, or from more complex mechanisms involving paracrine signalling from neurons and other cell types. To gain insights into the cell-type-specific mechanisms of APOE4 on myelination, we performed a combinatorial experiment in which we co-cultured APOE4/4 oligodendroglia with APOE3/3 iNeurons and vice versa. APOE3/3 oligodendroglia co-cultured with isogenic APOE4/4 iNeurons exhibited a similar (P > 0.99) level of MBP staining localizing to neurofilaments as all APOE3/3 co-cultures, suggesting that APOE4/4 neurons do not impair the ability of APOE3/3 oligodendroglia to produce MBP or engage neuronal axons (Fig. 5b). By contrast, APOE4/4 oligodendroglia co-cultured with APOE3/3 iNeurons exhibited significantly (P = 0.0130) reduced MBP staining that localized with neurofilament staining (Fig. 5b). The levels of MBP surrounding neurofilament in APOE4/4 oligodendroglia co-cultured with APOE3 iNeurons were similar (P > 0.9999) to all APOE4/4 co-cultures, with only about 20% of neurofilament-positive axons surrounded by MBP staining (Fig. 5b). This suggests that APOE4 in oligodendroglia is sufficient to impair the ability of oligodendroglia to produce MBP and engage neuronal axons, processes that are critical for myelination. Consistent with lower MBP protein expression in APOE4 post-mortem oligodendrocytes (Extended Data Fig. 8b), APOE4/4 iPS-cell-derived oligodendroglia monocultures also exhibited significantly lower MBP immunoreactivity compared with isogenic APOE3/3 (Extended Data Fig. 9i).
Given that APOE is detectable in both APOE3/3 and APOE4/4 iPS cell oligodendroglia cultures at similar levels (Extended Data Fig. 5i,j), we reasoned that decreased myelination is probably the result of impaired protein function. To test whether APOE4 myelination defects are also observed in cells lacking APOE, we generated an APOE-knockout iPS cell line (APOE−/−), and differentiated oligodendroglia and iNeurons. We then created co-cultures of APOE−/− oligodendroglia and iNeurons and compared myelination after 6 weeks in culture with APOE3/3 and APOE4/4 iNeuron–oligodendroglia co-cultures. MBP immunoreactivity and axonal localization in the APOE−/− co-cultures were significantly (P = 0.0005) lower than APOE3/3 co-cultures and were similar to APOE4/4 co-cultures (Extended Data Fig. 9j). To test whether the addition of APOE3 protein could rescue the myelination phenotype, we added recombinant APOE3 protein (rAPOE3) to APOE4/4 iNeuron–oligodendroglia co-cultures for 6 weeks. The addition of rAPOE3 increased MBP expression and localization in APOE4/4 cultures to similar (P = 0.4545) levels as APOE3/3 co-cultures (Extended Data Fig. 9j). Thus, myelination defects are observed in APOE4/4 and APOE−/− oligodendrocytes and can be rescued by the addition of APOE3 protein. Taken together, these results suggest that the impaired ability of APOE4 to transport cholesterol may reduce myelination.
Aiding cholesterol transport increases myelin
APOE4 expression in oligodendrocytes promotes both cholesterol accumulation and impaired myelination. We therefore reasoned that inhibiting cholesterol biosynthesis and/or facilitating cholesterol transport in APOE4/4 oligodendrocytes may reduce intracellular accumulation and improve myelination. To test this hypothesis, we used small molecules that inhibit cholesterol biosynthesis (simvastatin and atorvastatin) or facilitate cholesterol transport (2-hydroxypropyl-β-cyclodextrin; cyclodextrin). Cyclodextrin is a molecule that is known to reduce intracellular cholesterol accumulation in Niemann–Pick disease type C30. APOE4/4 iPS-cell-derived oligodendroglia were treated with each drug for 2 weeks, and intracellular cholesterol was subsequently quantified by BODIPY–cholesterol staining. APOE4/4 oligodendroglia cultured in the presence of cholesterol biosynthesis inhibitors (either simvastatin or atorvastatin) exhibited more BODIPY–cholesterol staining compared with APOE3/3 oligodendroglia (P = 0.0085 and 0.0450) (Fig. 6a). However, APOE4/4 oligodendroglia treated with cyclodextrin exhibited reduced BODIPY–cholesterol staining to an extent that was not significantly different (P > 0.99) from that in APOE3/3 oligodendroglia (Fig. 6a). In cyclodextrin-treated APOE4/4 oligodendroglia, we observed a significant (P = 0.0092) decrease in the number of intracellular droplets of BODIPY–cholesterol, as well as a significant (P < 0.0001) decrease in neutral lipid droplets stained with BODIPY. This suggests that cyclodextrin reduced both intracellular cholesterol and accumulation of neutral lipids such as triacylglycerides (Fig. 6b and Extended Data Fig. 10a).
a, Representative BODIPY–cholesterol images of APOE4/4 iPS-cell-derived oligodendroglia treated with cyclodextrin (CD) (1 mM), simvastatin (1 μM) or atorvastatin (1μM). Scale bar, 50 μm. BODIPY–cholesterol-positive punctae normalized to the number of nuclei. Bars represent mean values. n = 7 biological replicates. P values were calculated using one-way ANOVA with Bonferroni correction. NS, not significant. b, MBP and neurofilament staining of cyclodextrin-treated co-cultures of iPS-cell-derived APOE4/4 oligodendroglia and neurons versus APOE3/3 co-cultures. Scale bars, 10 μm. Axonal MBP expression was quantified using Imaris. Bars represent mean values. n = 3 biological replicates. P values were calculated using one-way ANOVA with Bonferroni correction. c, BODIPY–cholesterol and OLIG2 staining in control (n = 5) or cyclodextrin-treated (n = 4) APOE4/4-TR mouse brain. The arrows highlight BODIPY–cholesterol accumulations around OLIG2-positive nuclei. Datapoints represent individual mice. The bars represent the mean value of the treatment group. P values were calculated using unpaired two-tailed Student’s t-tests. Scale bar, 50 μm. d, Representative corpus callosum TEM images from APOE4/4-TR mice treated with saline (n = 5 mice, n = 51 axons) or cyclodextrin (n = 4 mice, n = 56 axons) for 8 weeks. Scale bars, 500 nm. The g-ratio was calculated as described in Fig. 4d. P values were calculated using unpaired two-tailed Student’s t-tests. e, Schematic of the novel-object recognition task with control (n = 12) and cyclodextrin-treated (n = 14) APOE4/4-TR female mice. Preference was calculated by dividing the time that the animal explored the new object with the nose by the total time that the mouse interacted with either object (measured using Noldus EthoVision). Data points represent individual mice. Bars represent mean values. P values were calculated using unpaired two-tailed Student’s t-tests with Welch correction. f, Puzzle box test comparing control (n = 9) and cyclodextrin-treated (n = 10) APOE4/4-TR mice. The task performance was recorded using Noldus EthoVision. The data points represent the mean values per treatment group and the bars represent the s.e.m. P values were calculated using unpaired two-tailed Student’s t-tests. The schematics in e and f were generated using BioRender.
To determine whether the cyclodextrin-induced cholesterol reduction in APOE4/4 oligodendroglia affected myelination, we treated APOE4/4 oligodendroglia–iNeuron co-cultures with cyclodextrin for 6 weeks and then assessed myelination phenotypes (Fig. 6b). In all of the APOE3/3 co-cultures, a majority (62 ± 10%) of MBP staining was localized within 1 μm of neurofilament staining (Fig. 6b). By contrast, in untreated APOE4/4 co-cultures, less than a fifth (19 ± 7%) of MBP staining localized with neurofilament staining (Fig. 6b). Treating APOE4/4 co-cultures with cyclodextrin significantly (P = 0.0274) increased (by 43 ± 5%) MBP co-localization with neurofilament staining (Fig 6b) to similar levels (P = 0.0794) to those observed in APOE3/3 co-cultures (Fig. 6b).
We next assessed whether cyclodextrin could reduce aberrant cholesterol accumulation in oligodendrocytes and promote increased myelination in APOE4/4-TR mice in vivo. APOE4/4-TR mice were treated with subcutaneous injections of cyclodextrin or saline (control) for 8 weeks (n = 5 (control) and n = 4 (cyclodextrin-treated) mice). In the hippocampus of APOE4/4-TR cyclodextrin-treated mice, BODIPY–cholesterol droplets were reduced compared with the control mice (P = 0.0038; Fig. 6c). Moreover, in cyclodextrin-treated mice, we observed a significant increase (P = 0.0423) in BODIPY–cholesterol staining co-localizing within 1 μm of MBP staining, indicating that cyclodextrin may improve the trafficking of cholesterol to the myelin sheath (Extended Data Fig. 10b). MBP immunoreactivity was significantly increased (P = 0.0008) in APOE4/4-TR mice treated with cyclodextrin, suggesting that cyclodextrin may promote myelination (Extended Data Fig. 10b). Consistent with this, TEM-based ultrastructural analysis of corpus callosum from APOE4/4-TR mice revealed that cyclodextrin treatment significantly (P < 0.0001) reduced the g-ratio (n = 150 neurons for each genotype) versus saline-treated APOE4/4-TR mice, indicating an increased number of myelinated axons and thicker myelin sheaths (Fig 6d). Collectively, these data demonstrate that pharmacological enhancement of cholesterol transport can improve APOE4 myelination in vitro and in vivo.
Cyclodextrin improves cognition in APOE4 mice
To investigate whether the increased myelination observed in cyclodextrin-treated APOE4/4-TR mice is associated with functional improvements, we evaluated learning and memory using the novel-object recognition assay31 (Fig. 6e). The total distance travelled and time spent in the centre of the home cage (P = 0.1275 and P = 0.4478, respectively) was similar in control (n = 12) and cyclodextrin-treated (n = 14) APOE4/4-TR mice, suggesting that repeated cyclodextrin treatment did not affect general locomotion or induce anxiety-like behaviour (Extended Data Fig. 10c). However, when exposed to a new object, cyclodextrin-treated APOE4/4-TR mice exhibited significantly (P = 0.0466) increased preference for the new object compared with the control APOE4/4-TR mice, suggesting that cyclodextrin treatment may improve learning and memory (Fig. 6e). Furthermore, we used the puzzle box test—a test previously used to measure demyelination-associated cognitive dysfunction32—as an additional assessment of learning and executive function (Extended Data Fig. 10d). This test consists of an open, brightly lit arena connected by a tunnel to a covered goal box chamber with bedding (Fig 6f). On day 0, the mice were habituated by freely exploring the set-up for 5 min. Locomotion did not significantly vary between the treatment groups (Extended Data Fig. 10d). The next day (day 1), we blocked the tunnel with a piece of paper, placed a mouse in the arena and recorded the time required for the mouse to remove the paper and enter the goal box. The task was repeated for 4 days and the latency to enter the goal box between the first and last day was compared. Cyclodextrin-treated APOE4/4-TR mice exhibited a decrease in latency to enter the goal box (P = 0.02), whereas saline-treated APOE4/4-TR mice (control) did not improve between the trial days (P > 0.6; Fig. 6f). This demonstrates that cyclodextrin treatment improves learning and executive function in old APOE4/4-TR mice.
Discussion
By combining transcriptomic and lipidomic analysis of post-mortem human brains with functional studies using isogenic stem cell lines and knockin mouse models, we mechanistically dissected cell-type-specific effects of APOE4 from molecular pathways to cognitive behaviour. We found that APOE4 alters lipid and cholesterol homeostasis across multiple cell types in the human brain. Cholesterol gene expression and localization are dysregulated in oligodendrocytes, leading to impaired myelination. We observed that pharmacologically promoting cholesterol efflux increases myelination in vitro and in vivo, and improves cognition in APOE4/4-TR mice. These results establish a causal link between cholesterol dysregulation and myelination in APOE4 carriers, which may influence the onset and severity of cognitive decline in AD.
Developmental myelination defects might influence the age of onset of AD by increasing the vulnerability to amyloid-β, tau and neuroinflammatory toxicity that progressively accumulates in ageing33. Infant APOE4 carriers exhibit alterations in myelin volume and cognitive maturation trajectories relative to APOE3/3 carriers34,35. Reduced myelin volume and integrity often occur before the onset of dementia and are predictive of future cognitive impairment23,24,33,36. Thus, cognitive impairment could also be influenced by reduced myelin maintenance and repair in the adult AD brain. Pharmacological, dietary or lifestyle interventions focused on restoring cholesterol homeostasis and increasing myelin volume might increase cognitive reserves in individuals with APOE4.
Our results suggest that cholesterol transport could be an entry point to pharmacologically target impaired myelination, and that the APOE genotype can inform such a therapeutic approach. We demonstrate that facilitating cholesterol transport increases myelination and improves cognitive function in APOE4 mice. These effects are not observed when targeting biosynthesis, suggesting that APOE4 has a role in cholesterol trafficking and that increased biosynthesis might be a compensatory mechanism. Our integrative study therefore establishes a mechanistic link between APOE4, cholesterol transport and localization in oligodendrocytes, myelin impairment and cognitive performance. This finding offers therapeutic and diagnostic opportunities for Alzheimer’s disease.
Methods
ROSMAP participant selection
A total of 32 individuals were selected from the Religious Orders Study or the Rush Memory and Aging Project (ROSMAP)—two harmonized longitudinal cohort studies of ageing and dementia that include extensive post-mortem pathological evaluations and clinical data collected annually, as previously described15. No statistical methods were used to predetermine sample size. Details of clinical and pathological data collection methods have been previously reported37. To assess APOE4 effects in the context of AD pathology, 6 control participants with no or very low pathology (no pathology) and 6 age-matched participants with severe amyloid-β, tau pathology and cognitive decline (AD group) were selected independently for APOE3/3 and APOE3/4 carriers. For APOE3/3 and APOE3/4 carriers, participants were balanced between sexes (12 each) and matched for age (median age, 84.6 for APOE3/3 and 85.2 for APOE3/4). Informed consent and an Anatomical Gift Act were obtained from each participant, and the Religious Orders Study and Rush Memory and Aging Project were approved by an Institutional Review Board (IRB) of Rush University Medical Center. All of the participants signed a repository consent that allowed their data and biospecimens to be shared.
Isolation of nuclei from frozen post-mortem brain tissue
The protocol for the isolation of nuclei from frozen post-mortem brain tissue was adapted from a previous study12. All procedures were carried out on ice or at 4 °C. In brief, post-mortem brain tissue was homogenized in 700 µl homogenization buffer (320 mM sucrose, 5 mM CaCl2, 3 mM Mg(CH3COO)2, 10 mM Tris HCl pH 7.8, 0.1 mM EDTA pH 8.0, 0.1% IGEPAL CA-630, 1 mM β-mercaptoethanol and 0.4 U µl−1 recombinant RNase inhibitor (Clontech)) using a Wheaton Dounce tissue grinder (15 strokes with the loose pestle). The homogenized tissue was then filtered through a 40 µm cell strainer, mixed with an equal volume of working solution (50% OptiPrep density gradient medium (Sigma-Aldrich), 5 mM CaCl2, 3 mM Mg(CH3COO)2, 10 mM Tris HCl pH 7.8, 0.1 mM EDTA pH 8.0 and 1 mM β-mercaptoethanol) and loaded on top of an OptiPrep density gradient (750 µl 30% OptiPrep solution (30% OptiPrep density gradient medium, 134 mM sucrose, 5 mM CaCl2, 3 mM Mg(CH3COO)2, 10 mM Tris HCl pH 7.8, 0.1 mM EDTA pH 8.0, 1 mM β-mercaptoethanol, 0.04% IGEPAL CA-630 and 0.17 U µl−1 recombinant RNase inhibitor) on top of 300 µl 40% OptiPrep solution (40% OptiPrep density gradient medium, 96 mM sucrose, 5 mM CaCl2, 3 mM Mg(CH3COO)2, 10 mM Tris HCl pH 7.8, 0.1 mM EDTA pH 8.0, 1 mM β-mercaptoethanol, 0.03% IGEPAL CA-630 and 0.12 U µl−1 recombinant RNase inhibitor). The nuclei were separated by centrifugation (5 min, 10,000g, 4 °C). A total of 100 µl of nuclei was collected from the 30%/40% interphase and washed with 1 ml of PBS containing 0.04% BSA. The nuclei were centrifuged at 300g for 3 min (at 4 °C) and washed with 1 ml of PBS containing 0.04% BSA. The nuclei were then centrifuged at 300g for 3 min (4 °C) and resuspended in 100 µl PBS containing 0.04% BSA. The nuclei were counted and diluted to a concentration of 1,000 nuclei per µl in PBS containing 0.04% BSA.
Droplet-based snRNA-seq
For droplet-based snRNA-seq, libraries were prepared using the Chromium Single Cell 3′ Reagent Kits v3 according to the manufacturer’s protocol (10x Genomics). The generated snRNA-seq libraries were sequenced using the NextSeq 500/550 High Output v2 kits (150 cycles) or the NovaSeq 6000 S2 Reagent Kits.
snRNA-seq data preprocessing
Gene counts were obtained by aligning reads to the GRCh38 genome using Cell Ranger (v.3.0.2, 10x Genomics). To account for unspliced nuclear transcripts, reads mapping to pre-mRNA were counted. After quantification of pre-mRNA using the Cell Ranger count pipeline, the Cell Ranger aggr pipeline was used to aggregate all of the libraries (without equalizing the read depth between groups) to generate a gene-count matrix. The Cell Ranger (v.3.0) default parameters were used to call cell barcodes.
Quality control for cell inclusion
Outlier cells with less than 500 or more than 10,000 genes detected were excluded, and only genes detected in at least 10 cells were considered. The following quality measures were quantified for each cell: (1) the number of genes for which at least one read was mapped (indicative of library complexity); (2) the total number of counts; and (3) the percentage of reads mapped to mitochondrial genes (used to approximate the relative amount of endogenous RNA and commonly used as a measure of cell quality). Cells with a discriminatively high ratio of mitochondrial to non-mitochondrial read counts were excluded using unbiased k-means clustering-based binarization (k = 2). Nuclear-encoded protein-coding genes were considered for downstream analyses.
Clustering analysis and quality-control filtering
All dimensionality reduction, clustering and visualization analyses were performed using our computational analysis framework ACTIONet38, available at GitHub (https://github.com/shmohammadi86/ACTIONet, version ACTIONet-Legacy). In brief, for each round of clustering, singular value decomposition is performed for feature (gene) dimensionality reduction, and multiple rounds of matrix decomposition are performed to identify a lower dimensional cell state representation for each individual cell. This cell state representation is operationalized as a set of variables quantifying the relative contribution of latent cell state patterns learned from data to optimally describe the transcriptional heterogeneity of the whole dataset. This representation is used to build a cell network or embedding of which the structure captures transcriptomic state relationships at the single-cell level. This coupled decomposition and network-based analysis has been shown to recover biological cellular associations with improved performance relative to more conventional methods38. To avoid biased cell mixing due to independent sequencing batches or potential technical artifacts, a batch-correction step considering the sequencing batch as indicator vector was performed as part of the initial dimensionality reduction in all analyses. This step is achieved using ACTIONet’s function reduce.and.batch.correct.ace.Harmony, which internally uses the data integration procedure39 implemented in the software package Harmony (https://github.com/immunogenomics/harmony). Discrete groups of cells with similar transcriptomes (cell clusters) were identified by applying the Leiden graph-based clustering algorithm to the resulting network. The same methodology was applied in the second clustering round, using cell-type-annotated subsets of cells as input. As part of the quality-control filtering steps, clusters representing only cells from one individual, suspected to recover doublet cells, or to be composed of presumed low-quality cells, were excluded from downstream analyses. Doublet or low-quality cluster status was determined empirically on the basis of the examination of cell associations in 2D plots, the presence of mixed gene markers from distinct cell types and extreme quality-control metric values, the latter relative to those commonly observed in other subclusters of the same cell type. After applying the quality-control filtering steps, the dataset included 17,915 genes profiled in 164,741 nuclei.
Differential gene expression
Differential gene expression by negative binomial mixed model
Per-gene expression levels per cell type were modelled as a negative binomial model with random effects that model both within-individual and between-individual effects, as implemented in the R package NEBULA (v.1.2.1)40. In brief, NEBULA models gene counts as sampled from a negative binomial distribution, parameterized by a mean and variance.The mean parameter is modelled as an exponential scaled by a scaling factor (total library count per cell). The exponent is modelled as a linear combination of fixed and random effects (individual of origin), where the fixed effects are a linear combination of predictors: ~β0 × amyloid + β1 × nft + β2 × age_death + β3 × batch + β4 × APOE4, where the β4 coefficient and P value were computed with respect to the APOE4 variable of interest, when controlling for the remaining variables. Nebula results are reported in Supplementary Table 14.
Single-cell differential gene expression
Differential gene expression between groups of individual cells was estimated using a Wilcoxon rank-sum test, as implemented in the R package presto (v.1.0.0; https://github.com/immunogenomics/presto). Wilcoxon results are reported in Supplementary Table 13.
Pseudo-bulk differential gene expression
Individual-level normalized gene expression averages were fitted with the following linear model: gene expression ≈ β0 × APOE4 + β1 × amyloid + β2 × nft + β3 × age_death + β4 × msex + β5 × pmi. The linear models were fitted using the lmfit() function and t-statistics were computed using the eBayes() function from the Limma R package (v.3.50.3). Estimates are reported in Supplementary Table 15.
Pathway analyses
Databases
Both the APOE-associated and lipid-associated pathway databases were curated by filtering the union of pathways (downloaded from the Mayaan laboratory; https://maayanlab.cloud/Enrichr/#libraries) from Gene Ontology biological processes (2018), KEGG (2019), Reactome (2016) and HumanCyc (2016) pathways either for gene sets that contain the gene APOE or for gene sets of which the name contains at least one of the following terms: sterol, athero, cholest, LDL, HDL, lipoprotein, triglyceride, TAG, DAG, lipid, steroid, fatty acid and ceramide. Some pathways were renamed for clarity and owing to space constraints in Fig. 1. New names are documented in Supplementary Tables 4–6.
Pathway activity scores
Individual-cell-type-level normalized gene expression profile averages were first computed (normalized values were first computed using ACTIONet), followed by calculation of pathway activity scores, as previously implemented in the R package GSVA (v.1.42.0)41. In brief, GSVA estimates a normalized relative expression level per gene across samples. This expression level is then rank-ordered for each sample and aggregated into gene sets by calculating sample-wise enrichment scores using a Kolmogorov–Smirnov-like rank statistic. The following parameters were used to evaluate the GSVA function: mx.diff=TRUE, kcdf=c("Gaussian"), min.sz=5, max.sz=150. Activity scores computed this way accurately recovered cell type signatures that were not recovered by equally sized randomly sampled gene sets (data not shown), suggesting that this approach captures transcriptomically encoded biological signals in snRNA-seq data.
Differential pathway activity analysis
To minimize the discovery of false positive effects, activity scores were computed only after filtering out genes that were not expressed in a given cell type (defined as a nonzero detection rate ≥10%, see Supplementary Table 2 for a full list of expressed genes per cell type). For each pathway-cell type combination, pathway activity scores were modelled using a multivariate linear model as follows: pathway activity ~β0 × APOE4 + β1 × amyloid + β2 × nft + β3 × age_death + β4 × msex + β5 × pmi. Here, APOE4 is a binary variable, encoding the presence of at least one E4 allele. Amyloid and nft are averages of continuous measurements of pathology across 8 and 5 brain regions, respectively, as evaluated and reported by ROSMAP. Age_death is a continuous measurement indicating age at death, pmi represents the PMI (a continuous measurement) and sex is a binary variable encoding biological sex. β0 represents the additive effect of APOE4 (≥1 allele) on pathway activity, when accounting for other confounding variables that might be correlated with, and therefore explain, a portion of the variation observed in the activity of a pathway of interest. The linear models were fitted using the lmfit() function and t-statistics were computed using the eBayes() function from the Limma R package (v.3.50.3). To prioritize candidate APOE4-dysregulated processes to be subsequently thoroughly validated using independent data and experimental methods, processes with snRNA-seq evidence of association with APOE4 at a nominal P value of 0.05 were considered to be potential candidates (Fig. 1). This procedure resulted in prioritized candidate pathways for all major cell types. β0 estimates with confidence intervals related to the analysis in Fig. 1 and Extended Data Fig. 2 are reported in Supplementary Tables 4–6, including statistics, and nominal and FDR-adjusted P values.
Pathway dysregulation by gene set enrichment analysis
For a subset of highly relevant pathways, a second statistical analysis was performed to further corroborate the reproducibility of results across the methods. To this end, Nebula (above) was used to model changes in gene expression associated with APOE4 in oligodendrocytes. Next, for each gene, the following score was computed: s = −log10(P) × sign(β4), and genes were rank-ordered on the basis of the scores. An R implementation of gene set enrichment analysis (fgsea, v.1.20.0) was used to estimate the statistical overrepresentation of lipid-associated gene sets (above) within high-scoring, differentially expressed genes. Lipid-related gene sets with evidence of having an APOE4 effect (P < 0.05) are shown in Extended Data Fig. 2d.
Untargeted lipidomics analysis of post-mortem PFC
The untargeted lipidomics was previously described42. In brief, approximately 50 mg of dorsolateral PFC tissues were added to a 96-well plate with 1 ml PBS and 1.4 mm ceramic beads, and then homogenized for 4.0 m s−1 for 20 s using the Omni homogenizer (OMNI International) at 4 °C. After homogenization, the samples were centrifuged at 4,000 rpm for 10 min and the supernatant was collected and extracted using a combination of methyl tert-butyl ether (MTBE) and methanol using an automated extraction system (Biotage Extrahera, Biotage). The extract was then filtered and dried under nitrogen for lipidomic analysis. Dried lipids were reconstituted in 200 μl 1:1 chloroform:methanol and resolved using a Thermo Accucore C18 column on a Thermo Vanquish Ultimate 3000 UPLC coupled to a Thermo Orbitrap ID-X Tribrid mass spectrometer (Thermo Fisher Scientific). Chromatography was operated at the flow rate of 0.4 ml min−1 at 40 °C during a 15 min gradient. The mobile phase of ultra-performance liquid chromatography (UPLC)-grade solvents consisted of solvent A (10 mM ammonium formate in 60% acetonitrile with 0.1% formic acid) and solvent B (10 mM ammonium formate in 10% acetonitrile and 90% isopropanol with 0.1% formic acid). Data were acquired in both positive and negative mode. Data were acquired in full-scan mode at a resolution of 120,000 for all of the samples. Iterative data-dependent acquisition was collected on pooled samples and National Institute of Standards and Technology (NIST) standard reference material (SRM) 1950 human plasma at a resolution of 15,000 using stepwise collision energy to obtain substructure information of lipid identification. Pooled quality-control samples were run after every ten samples using the same analytical procedure as samples and used for the normalization of data. The raw data were processed using LipidSearch (v.4.2; Thermo Fisher Scientific) and a total 2,200 lipids from both negative and positive modes were identified by full scan with data-dependent MS2. To determine cholesterol ester (ChE) in the samples, we used global profiling of NIST SRM 1950 as a m/z RT library and all ChE identified in this library had an MS/MS confidence as either level B or C. Grade B calls are lipids of which class and some fatty acid chains were identified and grade C calls are lipids of which class or fatty acid was identified. The lipid annotation in the samples was based on m/z and retention time matching to the m/z RT library. We next normalized the lipidomics data to remove systematic variations based on quality-control pooled samples using the Shiny-SERRF R app (https://github.com/slfan2013/Shiny-SERRF) and the normalized data were used for further analysis. Normalized values are reported in Supplementary Table 8. The cohort was subset into 314 individuals (194 individuals with AD pathology, for whom niareagansc was equal to 1 or 2; and 120 individuals without AD pathology, for whom niareagansc was equal to 3 or 4). APOE4 carriers (including APOE3/4 and APOE4/4 genotypes) versus APOE4 non-carriers (that is, APOE3/3 carriers) were compared using Wilcoxon rank-sum tests (stats base package, R v.4.1.2). Only peaks with a signal to noise ratio of >3 and of quality ≥0.6 were considered in the analysis. A total of 4 out of 17 peaks passed this quality-control cut-off (Supplementary Table 8). Underscore_N (for example, ChE(18:1)_1 versus ChE(18:1)_2) refers to the same lipid species, with different retention times. Underscore_c indicates a combination of _1 and _2 peaks. Species without an underscore have a single retention peak.
Untargeted lipidomics of post-mortem corpus callosum
Lipids were extracted using a methyl-tert-butyl ether (MTBE)-based liquid–liquid method. Tissue homogenates were prepared at a concentration of 50 mg (wet weight) per ml in 150 mM NaHCO3 in 7 ml Precellys tubes containing ceramic beads (CK14 for soft tissue). The vials were placed in the Precellys tissue homogenizer and shaken at 7,900 rpm for 3 cycles of 60 s with a 60 s rest between cycles. Then, 60-μl aliquots of the resulting homogenates were removed from each sample and placed in 16 × 100 mm glass tubes for extraction. To these, we added 100 l 150 mM NaHCO3, 100 μl deionized water, 1 ml MTBE/methanol 7:2 (v/v) and 5 μl internal standard mixture (Equisplash, Avanti Polar Lipids, 330731 diluted 1:1 with methanol). The tubes were vortexed and mixed on a tabletop shaker for 15 min at room temperature and then centrifuged at 5 °C for 15 min at 3,000g to separate the phases. The top, organic phase was removed carefully using a Pasteur pipette into new glass tubes, and the organic solvent was removed on a SpeedVac vacuum concentrator. The dried samples were then reconstituted in 200 μl n-butanol/methanol 1:1 (v/v) and placed into autosampler vials for analysis.
Bulk RNA-seq analysis of isogenic iPS-cell-derived oligodendroglia
A total of 700 μl of ethanol and Trizol were added at a 1:1 ratio to each well, and cells were collected into tubes. The procedure was then performed according to the Zymo Direct-Zol Microprep kit instructions. Approximately 500 ng of each sample was submitted for library preparation (Kappa HiFi) and bulk sequencing performed by the BioMicro Center at MIT’s Department of Biology, using the NextSeq Illumina platform. Raw FASTQ data were quasi-mapped to a reference transcriptome derived from the GRCh38 human genome assembly and quantified using Salmon (v.1.3.0). Differential gene expression testing was performed with DESeq2 (v.1.22.1) with median-ratio count normalization, parametric dispersion estimation and additional count normalization by variance-stabilizing transformation. GSVA was used to compute differential pathway activity scores on replicate-level bulk sequencing profiles. A simple linear model, with APOE status as the single predictor, was used to compute effect sizes and confidence intervals. The normalized (FPKM) gene count matrix is reported in Supplementary Table 10. Differentially expressed gene statistics are reported in Supplementary Table 12.
Comparative analysis of iPS cell and post-mortem transcriptomes
Normalized gene expression values (FPKM) for iPS-cell-derived neuronal cells (n = 6), microglial-like cells (n = 8) and astrocytes (n = 6) were obtained from a previous publication (Gene Expression Omnibus: GSE102956)14. To assess whether corresponding cell types in the iPS cell and post-mortem data showed consistent transcriptomic signatures relative to remaining cell types, while accounting for sequencing and batch effects, scaled gene expression values (mean-centred, s.d. = 1) were first independently computed for each dataset and then concatenated into a normalized expression matrix. Pathway activity scores were computed on this concatenated matrix using GSVA. Principal component analysis was performed on the concatenated matrix of gene-by-individual (or replicates for iPS cells). Pairwise distances between individual-level cell type averages from post-mortem tissue and iPS-cell-derived cells were computed in scaled mean-centred gene space.
iPS cell differentiation into oligodendroglia
iPS cells were differentiated into oligodendrocytes as described previously43. In brief, after single-cell disassociation with accutase, iPS cells were plated at nearly 100% confluency. Differentiation was initiated the next day by the addition of 10 μM SB431542 and 100 nM LDN 193189 with 100 nM all-trans RA in DMEM/F12 (1:1) medium supplemented with N2 (defined as day 0 of the induction). The medium was changed every day. At day 8, SAG (1 µM) was added to the medium along with 10 μM SB431542 and 100 nM LDN-193189. At day 12, adherent cells were lifted and seeded in low-attachment plates to promote sphere aggregation. Spheres were cultured in DMEM/F12 (1:1) medium supplemented with N2, RA and SAG. On day 30, spheres were plated into poly-l-ornithine/laminin-coated dishes and cells were allowed to migrate out of the sphere. At this stage, the medium was switched to DMEM/F12 (1:1) medium supplemented with N2, B27, 10 ng ml−1 PDGF-AA, 10 ng ml−1 IGF, 5 ng ml−1 HGF, 10 ng ml−1 NT3, 25 μg ml−1 insulin, 100 ng ml−1 biotin, 1 μM cAMP and 60 ng ml−1 T3. At day 75, cells were dissociated and selected for NG2 by MACS purification. The resulting cells were expanded in DMEM/F12 (1:1) medium supplemented with N2, B27, 10 ng ml−1 PDGF-AA, 10 ng ml−1 β-FGF and 10 ng ml−1 NT3. To promote oligodendrocyte maturation, the medium was switched to DMEM/F12 (1:1) medium supplemented with N2, B27, 20 μg ml−1 ascorbic acid, 10 mM HEPES, 25 μg ml−1 insulin, 100 ng ml−1 biotin, 1 μM cAMP and 100 ng ml−1 T3 for at least 2 weeks.
Drug treatment of iPS-cell-derived oligodendroglia
Cells were plated onto a Millipore eight-chamber glass slide at a density of 250,000 cells per well. Cells were then cultured in Neurobasal medium, supplemented with PDGFR-AA, NT3 and β-FGF. Cells were treated with either atorvastatin (1 μM), simvastatin (1 μM) or cyclodextrin (1 mM) for 2 weeks. Cells were incubated with BODIPY–cholesterol, and then fixed with 4% PFA and counter-stained for WGA-membrane and DAPI.
APOE oligodendroglia lipidomics
Cell pellets from oligodendrocytes were extracted by adding 600 µl methanol, 300 µl H2O and 400 µl chloroform. The samples were vortexed for 10 min and centrifuged for 10 min at 10,000g at 4 °C. The non-polar layer was transferred to a glass vial and dried under a stream of N2 gas and the samples were stored at −80 °C. Dried lipids were resuspended with 200 µl of 1:1:1 methanol:acetonitrile:2-propanol and 10 µl was injected for analysis by MS. The LC–MS system uses an Accela UPLC pump (Thermo Fisher Scientific) and an Exactive orbitrap mass spectrometer (Thermo Fisher Scientific). Chromatography was performed using a Luna C8 reversed-phase column (150 × 2.0 mm, 3 μm particle size, 100 Å pore size, Phenomenex) with a binary gradient (solvent A, 97:3 water:methanol with 10 mM tributylamine and 15 mM acetic acid (pH 4.5); and solvent B, 100% methanol). The gradient ran linearly from 80–99% B from 0 to 20 min, remained at 99% B from 20 to 40 min, ran from 99% B to 80% B to 41 min, and remained steady at 80% B to 50 min to re-equilibrate the column at a flow rate of 200 µl min−1. The autosampler temperature was held at 4 °C, injection volume 10 μl and column temperature 25 °C. The mass spectrometer was operated in negative and positive ionization modes. The electrospray ionization settings were as follows: sheath gas flow rate 30 (arbitrary units), auxiliary gas flow rate 10 (arbitrary units), sweep gas flow rate 5 (arbitrary units), spray voltage 3 kV, capillary temperature 325 °C, capillary voltage −50 V, tube lens voltage 100 V and skimmer voltage −25 V. The MS resolution was set to 100,000 resolving power at m/z 200 and the automatic gain control was set to high dynamic range with a maximum injection time of 100 ms. The scan range was 200–400 m/z in the first 20 min and 300–575 m/z in the subsequent 30 min. Resulting MS data were analysed using El-Maven (https://www.elucidata.io/) and compounds were identified using m/z or using a library of standards with known retention times.
BODIPY–cholesterol stain
Formalin-fixed human post-mortem brain tissue was sectioned using the Leica vibratome at a thickness of 40 μm. Brain sections were incubated overnight at 4 °C in a solution of 1 μg ml−1 BODIPY–cholesterol in PBS with 0.1% Triton X-100 + 10% bovine serum albumin, and primary antibodies. Brain sections were washed four times with PBS-T and stained with Alexa secondary antibodies (1:1,000). The samples were washed four times with PBS after secondary staining. Autofluorescence was quenched using True Black (Sigma-Aldrich) according to the manufacturer’s instructions. The sections were mounted onto slides and imaged using the Zeiss LSM 880 confocal microscope, using the same imaging parameters for each image.
Mouse immunohistochemistry
Mice were anaesthetized by exposure to gaseous isoflurane, and transcardially perfused with ice-cold PBS. The brains were dissected out, one hemisphere was frozen in dry ice, and one hemisphere was drop-fixed in 4% paraformaldehyde. The post-fixed hemisphere was sliced at a thickness of 40 μm using the Leica vibratome. Slices were blocked with a buffer containing 0.3% Triton X-100 and 10% normal donkey serum for 2 h at room temperature, before being incubated overnight with primary antibodies at 4 °C. The slices were then washed with PBS and incubated with secondary antibodies for 2 h at room temperature, washed with PBS again, and mounted. The slides were imaged using a Zeiss LSM 880 microscope with the same parameters for each image, and analysed using the ImageJ or Imaris software. All animal experiments were performed according to the Guide for the Care and Use of Laboratory Animals and were approved by the National Institutes of Health and the Committee on Animal Care at the Massachusetts Institute of Technology.
Mouse models
All experiments were performed according to the Guide for the Care and Use of Laboratory Animals and were approved by the National Institute of Health and the Committee on Animal Care at Massachusetts Institute of Technology. To ensure rigour and reproducibility in data analysis, researchers analysing data were blinded to the experimental conditions. Multiple experimenters were involved in each experiment and commonly switched roles to minimizes possible effects of experimenter bias and ensure that experimental details are consistent, adequately reported and reproducible. APOE-TR mice were obtained from Taconic. Colonies were bred and aged in the MIT vivarium under standard light–dark cycles, temperature and humidity. For all mouse model experiments, APOE-targeted replacement mice, in which the endogenous mouse locus was replaced with human APOE3/3 or human APOE4/4 on the C57BL/6NTac background, were used. Ages ranged from 6 to 9 months. Both male and female mice were included in the experiments and sex was balanced across treatment and genotype conditions. Sample size was determined from previous publications from our laboratory. Power analysis to determine sample size was not performed. Treatment and genotype groups were age-matched as much as possible. Experimenters were not blinded to drug injections or when executing behavioural trials. Data analysis of the TEM images was performed manually under blinded conditions. Data analysis of behavioural and immunohistochemistry data was performed using software and scripts (see the corresponding methods sections).
TEM analysis
Animals were euthanized by exposure to gaseous isoflurane. The head was submerged under EM-grade fixative, and the brain was removed without exposure to air. The corpus callosum was dissected out, and post-fixed in EM-grade fixative for 72 h. Tissue preparation and TEM analysis was performed by the Harvard Medical School Electron Microscopy Core. g-ratios were calculated using Image J software by dividing the axon diameter by the outer diameter of the myelin sheath. The number of lipid droplets per 1 µm2 surface area was also quantified for each mouse. At least four images for each animal were analysed. Each experiment consisted of at least three mice per treatment group. Analysis was performed in a blinded manner.
Western blotting
The protein concentration of each sample was measured using the Bradford protein assay. Volumes corresponding to 50 μg of protein for each sample were loaded into Mini-PROTEAN TGX precast gels (BioRad), and a current of 100 V was applied for 45 min. The gel was transferred to a membrane for 30 min, blocked in 3% milk in TBS-T for 2 h at room temperature and incubated with primary antibodies overnight at 4 °C. Membranes were washed with TBS-T, incubated with an antibody-conjugated horseradish peroxidase (1:2,000) for 2 h at room temperature, and a chemiluminescence activator.
Mouse cyclodextrin treatment
APOE4/4-TR male and female mice (aged 6 months) were injected subcutaneously in the right thigh with 2 g per kg body weight 2-hydroxypropyl-β-cyclodextrin in phosphate-buffered saline (200 μl total volume) or 200 μl PBS twice a week for 8 weeks. For immunohistochemistry and TEM analysis, mice were anaesthetized and subjected to cardiac perfusion of PBS followed by ice-cold 4% paraformaldehyde. Brains were dissected out and post-fixed in 4% PFA overnight at 4 °C.
Novel-object recognition task
Mice were placed into the test arena and allowed to move freely for 10 min. The total distance moved and the time that mice spent in the centre of the arena was tracked using the EthoVision XT software from Noldus. Then, 1 day later, the mice were then placed into the same arena containing two objects and allowed to freely investigate them both for 8 min. After an interval of 4 h, the mice were then placed back into the chamber with one new object and one familiar object. The preference index was calculated by determining how much time an animal spent with its nose interacting with the new object, divided by the total time an animal spent interacting with either object.
Puzzle box task
Mice were placed in a puzzle box and allowed to move freely for 10 min. The total distance moved and time that the mice spent in the centre of the arena were tracked using the EthoVision XT software from Noldus, using the same parameters for each animal. Then, 1 day later, the mice were exposed to the puzzle box task, which consists of an open, brightly-lit arena connected by tunnel to a dark chamber with bedding. On day 0, we habituated the mice by giving them 5 min to freely explore the apparatus. The next day (day 1), the tunnel entrance was blocked with a piece of paper. The mice were then placed in the brightly lit arena, and the latency for the animals to remove the paper and enter the goal box was recorded. The assay was repeated for 4 days, and the time to enter the goal box, or latency, was compared between the first day with the obstacle (day 1) and the fourth day (day 4).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The snRNA-seq data are available at the Synapse AD Knowledge Portal (syn38120890), including corresponding ROSMAP metadata. The data are available under the controlled use conditions set by human privacy regulations. To access the data, a data use agreement is needed. This registration is in place solely to ensure anonymity of the ROSMAP study participants. A data use agreement can be agreed with either RUSH university Medical centre (RUMC) or with SAGE, which maintains Synapse, and can be downloaded from their websites. See our code repository for a detailed list of data availability and download links.
Code availability
Codes, along with detailed instructions on how to reproduce the analyses presented herein, are available on GitHub (https://github.com/djunamay/apoe4myelin).
References
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Corder, E. H. et al. Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer’s disease in late onset families. Science 261, 921–923 (1993).
Strittmatter, W. J. et al. Apolipoprotein E: high-avidity binding to beta-amyloid and increased frequency of type 4 allele in late-onset familial Alzheimer disease. Proc. Natl Acad. Sci. USA 90, 1977–1981 (1993).
Liu, C.-C. et al. ApoE4 accelerates early seeding of amyloid pathology. Neuron 96, 1024–1032 (2017).
Shi, Y. et al. ApoE4 markedly exacerbates tau-mediated neurodegeneration in a mouse model of tauopathy. Nature 549, 523–527 (2017).
Castellano, J. M. et al. Human apoE isoforms differentially regulate brain amyloid-β peptide clearance. Sci. Transl. Med. 3, 89ra57 (2011).
Yamazaki, Y., Zhao, N., Caulfield, T. R., Liu, C.-C. & Bu, G. Apolipoprotein E and Alzheimer disease: pathobiology and targeting strategies. Nat. Rev. Neurol. 15, 501–518 (2019).
Crean, S. et al. Apolipoprotein E ε4 prevalence in Alzheimer’s disease patients varies across global populations: a systematic literature review and meta-analysis. Dement. Geriatr. Cogn. Disord. 31, 20–30 (2011).
Foley, P. Lipids in Alzheimer’s disease: a century-old story. Biochim. Biophys. Acta 1801, 750–753 (2010).
Sienski, G. et al. APOE4 disrupts intracellular lipid homeostasis in human iPSC-derived glia. Sci. Transl. Med. 13, eaaz4564 (2021).
Tcw, J. et al. Cholesterol and matrisome pathways dysregulated in astrocytes and microglia. Cell 185, 2213–2233 (2022).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Xu, Q. et al. Profile and regulation of apolipoprotein E (ApoE) expression in the CNS in mice with targeting of green fluorescent protein gene to the ApoE locus. J. Neurosci. 26, 4985–4994 (2006).
Lin, Y.-T. et al. APOE4 causes widespread molecular and cellular alterations associated with Alzheimer’s disease phenotypes in human iPSC-derived brain cell types. Neuron 98, 1141–1154 (2018).
Bennett, D. A. et al. Religious orders study and rush memory and aging project. J. Alzheimers Dis. 64, S161–S189 (2018).
Wang, D. et al. Comprehensive functional genomic resource and integrative model for the human brain. Science 362, eaat8464 (2018).
Franzén, O., Gan, L.-M. & Björkegren, J. L. M. PanglaoDB: a web server for exploration of mouse and human single-cell RNA sequencing data. Database 2019, baz046 (2019).
Huang, Y.-W. A., Zhou, B., Wernig, M. & Südhof, T. C. ApoE2, ApoE3, and ApoE4 differentially stimulate APP transcription and Aβ secretion. Cell 168, 427–441 (2017).
Zalocusky, K. A. et al. Neuronal ApoE upregulates MHC-I expression to drive selective neurodegeneration in Alzheimer’s disease. Nat. Neurosci. 24, 786–798 (2021).
Thadathil, N. et al. DNA double-strand break accumulation in Alzheimer’s disease: evidence from experimental models and postmortem human brains. Mol. Neurobiol. 58, 118–131 (2021).
Ye, S. et al. Apolipoprotein (apo) E4 enhances amyloid beta peptide production in cultured neuronal cells: apoE structure as a potential therapeutic target. Proc. Natl Acad. Sci. USA 102, 18700–18705 (2005).
Hatters, D. M., Peters-Libeu, C. A. & Weisgraber, K. H. Apolipoprotein E structure: insights into function. Trends Biochem. Sci 31, 445–454 (2006).
Brun, A. & Englund, E. A white matter disorder in dementia of the Alzheimer type: a pathoanatomical study. Ann. Neurol. 19, 253–262 (1986).
Bartzokis, G. Age-related myelin breakdown: a developmental model of cognitive decline and Alzheimer’s disease. Neurobiol. Aging 25, 5–18 (2004).
Jäkel, S. et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 566, 543–547 (2019).
Law, S.-H. et al. An updated review of lysophosphatidylcholine metabolism in human diseases. Int. J. Mol. Sci. 20, 1149 (2019).
Ye, J. et al. ER stress induces cleavage of membrane-bound ATF6 by the same proteases that process SREBPs. Mol. Cell 6, 1355–1364 (2000).
Hapala, I., Marza, E. & Ferreira, T. Is fat so bad? Modulation of endoplasmic reticulum stress by lipid droplet formation. Biol. Cell 103, 271–285 (2011).
Saher, G., Quintes, S. & Nave, K.-A. Cholesterol: a novel regulatory role in myelin formation. Neuroscientist 17, 79–93 (2011).
Ottinger, E. A. et al. Collaborative development of 2-hydroxypropyl-β-cyclodextrin for the treatment of Niemann-Pick type C1 disease. Curr. Top. Med. Chem. 14, 330–339 (2014).
Leger, M. et al. Object recognition test in mice. Nat. Protoc. 8, 2531–2537 (2013).
Chiang, A. C. A. et al. Bexarotene normalizes chemotherapy-induced myelin decompaction and reverses cognitive and sensorimotor deficits in mice. Acta Neuropathol. Commun. 8, 193 (2020).
Dean, D. C. 3rd et al. Association of amyloid pathology with myelin alteration in preclinical Alzheimer disease. JAMA Neurol. 74, 41–49 (2017).
Dean, D. C. 3rd et al. Brain differences in infants at differential genetic risk for late-onset Alzheimer disease: a cross-sectional imaging study. JAMA Neurol. 71, 11–22 (2014).
Remer, J. et al. Longitudinal white matter and cognitive development in pediatric carriers of the apolipoprotein ε4 allele. Neuroimage 222, 117243 (2020).
Gold, B. T., Powell, D. K., Andersen, A. H. & Smith, C. D. Alterations in multiple measures of white matter integrity in normal women at high risk for Alzheimer’s disease. Neuroimage 52, 1487–1494 (2010).
Bennett, D. A. et al. Decision rules guiding the clinical diagnosis of Alzheimer’s disease in two community-based cohort studies compared to standard practice in a clinic-based cohort study. Neuroepidemiology 27, 169–176 (2006).
Mohammadi, S., Davila-Velderrain, J. & Kellis, M. A multiresolution framework to characterize single-cell state landscapes. Nat. Commun. 11, 5399 (2020).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
He, L. et al. NEBULA is a fast negative binomial mixed model for differential or co-expression analysis of large-scale multi-subject single-cell data. Commun. Biol. 4, 629 (2021).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7 (2013).
Kamphorst, J. J., Fan, J., Lu, W., White, E. & Rabinowitz, J. D. Liquid chromatography-high resolution mass spectrometry analysis of fatty acid metabolism. Anal. Chem. 83, 9114–9122 (2011).
Douvaras, P. & Fossati, V. Generation and isolation of oligodendrocyte progenitor cells from human pluripotent stem cells. Nat. Protoc. 10, 1143–1154 (2015).
Acknowledgements
We thank the individuals who donated post-mortem brain samples, and their families, for enabling this research; Y. Zhou, E. McNamara and T. Garvey for administrative support and animal care; and J. M. Bonner for input on the lipidomic analyses. We acknowledge support from the Robert A. and Renee E. Belfer Family, The JPB Foundation, The Carol and Gene Ludwig Family Foundation, the Cure Alzheimer’s Fund and the National Institutes of Health (RF1 AG062377, RF1 AG054012-01, U54HG008097 and 747UG3NS115064). J.W.B. was supported by the NIH grant UG3-NS115064, R01NS114239-01A1, Cure Alzheimer’s Fund; L.A.A. was supported by NIH grant RF1-AG0540124 and the MIT BCS Henry E. Singleton Graduate Student Fellowship; D.v.M. was supported by the MIT BCS Broshy Graduate Student Fellowship and the MIT BCS Halis Graduate Student Fellowship; X.J. was supported by NIH grant U01-NS110453; Y.-T.L. was supported by NIH grant R01-AG058002; W.T.R. was supported by an Alzheimer’s Association Research Fellowship; H.P.C. was supported by NIH grants RF1-AG054012 and RF1-AG062377. ROSMAP is supported by NIA grants P30AG20262, R01AG15819, R01AG17917, U01AG46152, U01AG 61356 and P30AG72975. RF1AG057470, RF1AG051633, K24AG062786 from NIA to I.H. ROSMAP resources can be requested at https://www.radc.rush.edu. Graphic illustrations were generated using BioRender under agreements VI24HP0GQM and NE24HPHF0X.
Author information
Authors and Affiliations
Contributions
J.D.-V. and D.v.M. performed computational analyses of all functional genomic data. J.W.B. and L.A.A. designed and performed the experiments and data analysis. H.M., A.N. and X.J. generated the single-cell data. D.A.B. provided biospecimens and data and reviewed the manuscript. S.M.D. performed lipidomic data generation and analysis. C.-Y.C., K.M.-S., I.H., W.T.R. and E.A.O. generated and analysed human lipidomic data. E.A., M.B. and A.E. assisted with tissue culture. Y.T.-L. and T.K. generated the iPS cell lines used in the study. L.L., M. Kahn, C.B.-D., R.R., L.R’B. and N.L. assisted with experiments and data analysis. The study was conceived and designed by J.W.B., L.A.A., J.D.-V., D.v.M., L.-H.T. and M. Kellis. All of the authors contributed to writing the paper and making the figures.
Corresponding authors
Ethics declarations
Competing interests
The authors J.W.B., L.A.A., J.D.V., D.v.M., A.E., M. Kellis and L.-H.T. have filed a patent application (PCT/US2022/020271) based on the findings.
Peer review
Peer review information
Nature thanks Gonclao Castelo-Branco, Jeremy Miller and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Subject-level metadata and single-cell annotation quality control.
a, Study cohort description by APOE genotype groups: APOE3/3-carriers (grey), APOE3/4-carriers (pink), APOE4/4-carriers (red); balanced according to positive (glow) or negative pathological diagnosis (no glow) Cartoons generated with BioRender. APOE4/4 group comprised 3 male and 5 female subjects, all with an AD diagnosis. AD status is defined based on pathological (high amyloid and tau pathology) and cognitive diagnosis of Alzheimer’s dementia (final consensus cognitive diagnosis, cogdx=4). b, Distribution of pathology variables, PMI, and age at death in cohort. Unadjusted wilcoxon test p-values shown, two-sided (N = 6 per group, except for APOE4/4, where N = 8). Boxplots indicate median, 25th and 75th percentiles. c, Experimental and computational workflow of the single-cell analysis involves single nuclei isolation and sequencing, followed by computational analysis for sub-clustering and cell type annotation. Cartoons generated with BioRender d, Two-dimensional representation of all high-quality cells included in downstream analysis labelled by major cell types. e, Cell-type-specific marker gene expression projected onto two-dimensional representation of cell space. Coloured bars indicate the cell type for which a given gene was considered a marker. See panel i for legend. f, Expanded view of immune and vascular cell types within cell space. g, Enrichment of markers from two independent datasets (columns) within genes with preferential gene expression across annotated cell groups (rows). h, Distribution of inter-subject correlation values by cell type. Boxplots indicate median, 25th and 75th percentiles. i, Pairwise correlations of cell-type-specific individual-level transcriptomic profiles (average expression values across cells of a given type, N = 32 profiles per cell type). j, Median number of cells per subject by cell type. k, Fraction of subjects lacking cells of a given type. l, Individual distributions across cell types. m, Individual cell-type fractions organized by pathological diagnosis and APOE genotype.
Extended Data Fig. 2 APOE-associated and lipid pathway changes in APOE4 and AD.
a, Curation process for APOE-associated pathway database. Brain cell type expression was defined as nonzero detection of a gene in >10% of cells of that cell type. b, Transcriptional activity scores of APOE-associated pathways that show cell-type-specific patterns. c, Individual-level average expression of cholesterol biosynthesis genes from Fig. 2a in oligodendrocytes. d, Overrepresentation of lipid-related pathways (database from Fig. 1c) within genes differentially expressed in APOE4 relative to APOE3 in human post-mortem oligodendrocytes as estimated by a negative binomial mixed model (NBMM). e, Altered lipid-associated pathways (see methods) in APOE3/3 vs APOE3/4-carriers, in no AD background (control) (nominal p-value < 0.05, linear model, unadjusted, N = 6 per group). Pathways of special interest are highlighted in bold. (-) indicates negative regulation. f–k, Pathway activity scores for ‘Cholesterol biosynthesis III (via desmosterol) stratified by APOE genotype and/or AD pathology (p-values, linear model, unadjusted, n = 6 per group). Boxplots indicate median, 25th and 75th percentiles.
Extended Data Fig. 3 Large-scale lipidomic analysis of human prefrontal cortex.
a, Overview of ROSMAP cohorts on which lipidomic analysis was performed. Cartoons generated with BioRender b, Cohort clinicopathological characteristics. NFT = neurofibrillary tangles (p values calculated with Wilcoxon rank-sum test, two-sided). c, A single cholesterol ester (ChE) species passed quality control (see Methods) and showed significant (p<0.05, Wilcoxon rank-sum test, two-sided) differential concentration when comparing APOE3/3 vs APOE3/4 and APOE4/4-carriers stratified by pathology. ChE(18:1) indicates a ChE with carboxylate position 18 and one unsaturated fatty acid bond. d, Distribution of the concentration of ChE species from c) (p-value = 0.046, Wilcoxon rank-sum test, two-sided). Boxplots indicate median, 25th and 75th percentiles.
Extended Data Fig. 4 Lipid droplets in human and mouse brain.
a, Co-localization of immunohistochemistry against lipid-droplet associated protein perilipin-1 (PLIN1) with Bodipy-cholesterol staining in the human prefrontal cortex from APOE3/3 and APOE3/4 individuals (n = 3 imaged per genotype). Cell nuclei stained with Hoechst dye. b, Transmission electron microscopy (TEM) on corpus callosum from six-month-old APOE3-TR and APOE4-TR mice (males, n = 3 per genotype). The number of lipid droplets was quantified in four 1 μm2 areas per image from three images per mouse. Right panel, dots represent mean value per mouse and bar represents mean value for group, p value was calculated using unpaired, two-tailed student’s t-test. c, Representative images of Bodipy-cholesterol staining, with markers for microglia (IBA1), astrocytes (GFAP) and oligodendrocytes (OLIG2) in the prefrontal cortex of APOE4-carriers (n = four individuals). Dotted outline in the OLIG2 panel depicts the 2 μm radius around the nucleus that was quantified for the presence of Bodipy-cholesterol. Scale bar 10 μm. Bodipy-cholesterol staining was quantified for cell type based on localization with cell-type-specific markers. Bars depict means from different biological replicates. P values calculated using unpaired, two-tailed student’s t-test. The dotted outline in the OLIG2 panel depicts the 2 μm radius around the nucleus that was quantified for the presence of Bodipy-cholesterol. d, Co-localization of perilipin-1 (PLIN1) immunoreactivity around OLIG2-positive nuclei in prefrontal cortex from an APOE3/4 individual. e, Quantification of the number of perilipin-1 punctae around representative OLIG2-positive nuclei in APOE3/3 and APOE3/4 individuals (n=4 per genotype). The number of Perilipin-1 punctae was determined using Imaris software. P-value was calculated using unpaired, two-tailed student’s t-test.
Extended Data Fig. 5 Comparison of post-mortem oligodendrocytes and iPSC-derived oligodendroglia.
a, Comparison of the relative expression of myelination genes (MYRF, MOG, PLP1, PLLP, MAG, OPALIN) in iPSC-derived brain cell types and aggregated cell type gene expression profiles from post-mortem human brain single-nucleus data. b, Staining of MBP, MYRF, and MOG in iPSC-derived oligodendroglia. Scale bar 10 μm. c, Representative images of cultures of iPSC-derived oligodendroglia stained for PLP1 and MBP. Dots represent the percentage of nuclei positive for each marker across independent wells subjected to the same conditions (n = 5 biological replicates). Bars represent mean across all wells, error bars represent standard deviation. d, Principal component analysis of relative gene expression for post-mortem human brain cells and iPSC-derived counterparts. e, Transcriptional similarity between post-mortem human cell types and iPSC-derived oligodendroglia, p values calculated with Wilcoxon test, two-sided. Distributions represent distances between each post-mortem cell type (x-axis) and iPSC oligodendroglia in scaled gene space (N = 32 per post-mortem cell type, N = 6-8 for iPSC cell types). f–g, Gene set activity scores by GSVA on scaled expression values using genes shown in a or h, p value calculated by Wilcoxon test, two-sided. Boxplots indicate median, 25th and 75th percentiles. h, Relative expression of cholesterol-associated genes across iPSC-derived oligodendroglia, post-mortem human brain oligodendrocytes, and additional brain cell types. i, Gene expression distribution in iPSC-derived oligodendroglia. j, APOE immunoreactivity in APOE3/3 (n = 4 biological replicates) and APOE4/4 (n = 5 biological replicates) iPSC-derived oligodendroglia. Dots in right panel plots represent mean from independent images, p values were calculated with an unpaired, two-tailed student’s t-test. k, Distribution of gene nonzero detection rates in human post-mortem oligodendrocytes (snRNAseq). The nonzero APOE detection-rate is circa 4%, corresponding to circa 53rd percentile of genes. l, Distribution of expression levels for APOE-non-expressing (n = 53,095) and APOE-expressing (n = 1,882) post-mortem oligodendrocytes (snRNAseq). Groups were identified by K-means clustering on the non-zero detection rate.
Extended Data Fig. 6 Lipidomics on iPSC-derived oligodendroglia.
a, Volcano plots depicting differentially (adjusted p value < 0.05) detected lipid species in APOE4/4 oligodendroglia, compared to isogenic APOE3/3 controls. Each detected lipid species is organized according to lipid class, with cholesteryl esters having the highest number (15) of differentially expressed species.
Extended Data Fig. 7 Cell stress in APOE4 oligodendrocytes.
a, mRNA expression levels of SOAT1 (ACAT1) and CYP46A1 from bulk sequencing of isogenic iPSC-derived APOE3/3 and APOE4/4 oligodendroglia (n = 3 biological replicates per genotype, adjusted p values shown, computed by linear model). b, Perilipin-1 (PLIN1) immunoreactivity in APOE4/4 iPSC-derived oligodendroglia, and PLIN1 co-localization with Bodipy-cholesterol staining in APOE4/4 iPSC-derived oligodendroglia. c, Representative Bodipy staining for lipid droplets (n = 6 biological replicates). Scale bar represents 10 μm. Lipid droplets were quantified in two different isogenic sets of APOE3/3 and APOE4/4 oligodendroglia, that were generated from different individuals. Data points represent biological replicates and bars show means. p values were calculated using an unpaired, two-tailed student’s t-test. d, Percent of Bodipy-cholesterol staining in lysosome. Quantification was performed using ImageJ software with the same threshold setting for all images and conditions. Data points represent the mean of four images from one biological replicate. Bars represent means from four biological replicates. p value was calculated using unpaired, two-tailed student’s t-test. e, (left) Box plot of ATF-6 mediated unfolded protein response gene set activity (computed by GSVA) in human post-mortem oligodendrocytes. unadjusted p-value = 0.016, linear model. Boxplots indicate median, 25th and 75th percentiles. (right) Pathway gene members that are differentially expressed (FDR < 0.05, negative binomial mixed model) f, Representative images of immunohistochemistry against ATF-6 protein in APOE3/3 and APOE4/4 iPSC-derived oligodendrocytes, and quantification of number of cells with nuclear ATF-6 immunoreactivity (n = 5 biological replicates). Dots represent technical replicates, and bars represent mean per genotype. p value was calculated using an unpaired, two-tailed student’s t-test. g, Fold change for unfolded protein response genes in APOE4/4 vs APOE3/3 oligodendroglia (N = 3 per genotype) from panel e (adjusted p-value < 0.05, negative binomial distribution).
Extended Data Fig. 8 Myelin expression in humans and mice.
a, Myelin-associated gene expression changes in APOE3/3 vs APOE3/4 post-mortem oligodendrocytes for individuals with and without AD pathology (logFC<0 indicates down in APOE3/4; Wilcoxon test by wilcoxauc(), adjusted p value < 0.05). b, Immunohistochemistry for Myelin Basic Protein (MBP) in APOE3/3 and APOE3/4 individuals with AD diagnosis (n = 3 per genotype), and quantification. Mean fluorescence intensity quantified using FIJI ImageJ software. Dots represent mean value calculated from four images from one individual, and bars represent mean value from three separate individuals for each genotype. p value was calculated using an unpaired, two-tailed student’s t-test. c, Immunohistochemistry for myelin basic protein (MBP) in hippocampal slices of APOE3/3-TR (n = 7 mice) and APOE4/4-TR (n = 7 mice) at nine months of age, quantified according to mean fluorescence intensity. Quantification was performed using ImageJ. Bars represent mean intensity from all mice for each genotype, and error bars represent standard deviation. p values were calculated using unpaired, two-tailed student’s t-test. Scale bar represent 200 μm. d, Western Blot for Myelin Basic Protein (MBP) in APOE3/3-TR (n = 4) and APOE4/4-TR (n = 4) mouse cortex at six months of age. Each lane is a brain lysate prepared from a different mouse. Total area and intensity of bands normalized to GAPDH was quantified via ImageJ using mean intensity for each band. Bars represent means. p value was calculated using an unpaired, two-tailed student’s t-test. e, TEM on corpus callosum from APOE3/3-TR (n = 3) and APOE4/4-TR (n = 3) knock-in mice at 2 months of age. G-ratio was quantified using ImageJ software as stated in Main Fig. 4d). Scale bar represents 500 μm. Data points representing g-ratios for axons from each genotype, p value was calculated using an unpaired Wilcoxon test.
Extended Data Fig. 9 Myelin expression in iPSC-derived oligodendrocyte cultures.
a, Schematic of iPSC-derived NGN2-induced neuron (iNeuron) and oligodendrocyte encapsulation in Matrigel. Cartoons generated with BioRender b, Representative image of Neurofilament (red), Myelin Basic Protein (green), and Hoechst (blue) immunoreactivity in iPSC-derived neuron and oligodendrocyte co-culture, after two weeks encapsulated. Scale bar represents 50 μm. c, Representative images of (left) Neurofilament (red), MBP (green), and Hoechst (blue), and (right) Neurofilament (red), O4 (green), and Hoechst (blue) immunoreactivity in iPSC-derived neuron and oligodendrocyte co-culture, after six weeks encapsulated. Scale bar represents 50 μm. d, iPSC-derived oligodendroglia were labelled with GFP to visualize cellular localization after six weeks of co-culture with NGN2-induced neurons. Scale bar represents 50 μm. e, Representative image of Neurofilament (red), MBP (green), and Hoechst (blue), and Neurofilament (red), and Hoechst (blue) immunoreactivity in iPSC-derived oligodendroglia and NGN2-induced neuron after six weeks of co-culture. Scale bar represents 50 μm. f, TEM on myelinated axon from iPSC-derived neuron and oligodendrocyte co-culture, suggesting presence of myelin rings. g, Representative images showing Neurofilament (red), MBP (green) immunoreactivity in APOE3/3 parental and APOE4/4 isogenic co-cultures after three weeks encapsulated. Scale bar represents 10 μm. h, Axonal area per co-culture, comparing APOE3/3 (parental), APOE4/4 (isogenic) and APOE-/- (isogenic) co-cultures (n = 3 biological replicates). Axonal area was calculated by measuring the area immunoreactive to neurofilament, and normalized to APOE3/3. Bars represent means, error bars represent standard deviation, and p values were calculated using one-way ANOVA with Bonferroni correction. i, MBP density between APOE3/3 (parental) and APOE4/4 (isogenic) mono-cultures (n = 6 biological replicates) P value represents unpaired, two-tailed student’s t-test. j, Representative images showing Neurofilament (red), and MBP (green) immunoreactivity in APOE3/3 (parental), APOE4/4 (isogenic), APOE4/4 (isogenic) with recombinant human APOE3 protein, and APOE knock-out (isogenic) co-culture conditions (n = 3 biological replicates). The area of neuronal axon (immunoreactive against Neurofilament) positive for MBP was calculated using Imaris, and each experimental condition was compared to the APOE3/3 control condition. P values were calculated using one-way ANOVA with Bonferroni correction.
Extended Data Fig. 10 (2-Hydroxypropyl)-beta-cyclodextrin treatment in APOE4 knock-in mice.
a, Bodipy (neutral lipid) staining in control and (2-Hydroxypropyl)-beta-cyclodextrin (cyclodextrin) treated iPSC-derived APOE4/4 oligodendrocytes (n = 6 biological replicates). The number of lipid droplets was normalized by the total number of cell nuclei for each image. Bars represent the mean number of droplets per cell for each condition, error bars represent standard deviation. P value was calculated using unpaired, two-tailed student’s t-test. b, MBP and Bodipy-cholesterol staining in control (n = 5 mice) and cyclodextrin-treated (n = 4 mice) APOE4/4-TR mouse brain. Scale bar represents 50 μm. Quantification of cholesterol-myelin colocalization, and MBP signal, in cyclodextrin-treated APOE4/4-TR mice. MBP-cholesterol colocalization was quantified using Imaris. MBP staining was quantified using Image J. Bars represent mean, error bars represent standard deviation, and p-value was calculated using an unpaired two-tailed student’s t-test. c, Representative activity traces for open field assay on APOE4/4-TR control- (n = 14 mice) and cyclodextrin-treated (n = 15 mice) used for Novel Object Recognition task. Distance moved (centimeters) and duration in the centre (seconds) were measured and quantified using Noldus EthoVision software, with the same parameters for each animal. Dots represent individual mice, error bars are standard error of the mean, and p values were calculated using an unpaired, two-tailed student’s t-test. d, Representative activity traces for open field assay on APOE4/4-TR control- (n = 10 mice) and cyclodextrin- treated (n = 9 mice) used for the Puzzle Box task. Distance moved (centimeters) and duration in the centre (seconds) were measured and quantified using Noldus EthoVision software, with the same parameters for each animal. Dots represent individual mice, error bars represent standard error of the mean, and p values were calculated using an unpaired, two-tailed student’s t-test.
Supplementary information
Supplementary Information
Table of contents for Supplementary Tables 1–16 and Supplementary Videos 1 and 2.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Blanchard, J.W., Akay, L.A., Davila-Velderrain, J. et al. APOE4 impairs myelination via cholesterol dysregulation in oligodendrocytes. Nature 611, 769–779 (2022). https://doi.org/10.1038/s41586-022-05439-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05439-w
This article is cited by
-
GBA1 inactivation in oligodendrocytes affects myelination and induces neurodegenerative hallmarks and lipid dyshomeostasis in mice
Molecular Neurodegeneration (2024)
-
The associations between peripheral inflammatory and lipid parameters, white matter hyperintensity, and cognitive function in patients with non-disabling ischemic cerebrovascular events
BMC Neurology (2024)
-
Updates on mouse models of Alzheimer’s disease
Molecular Neurodegeneration (2024)
-
Border-associated macrophages in the central nervous system
Journal of Neuroinflammation (2024)
-
Neuropathogenesis-on-chips for neurodegenerative diseases
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.