Abstract
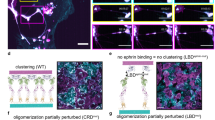
Most human cells require anchorage for survival. Cell–substrate adhesion activates diverse signalling pathways, without which cells undergo anoikis—a form of programmed cell death1. Acquisition of anoikis resistance is a pivotal step in cancer disease progression, as metastasizing cells often lose firm attachment to surrounding tissue2,3. In these poorly attached states, cells adopt rounded morphologies and form small hemispherical plasma membrane protrusions called blebs4,5,6,7,8,9,10,11. Bleb function has been thoroughly investigated in the context of amoeboid migration, but it has been examined far less in other scenarios12. Here we show by three-dimensional imaging and manipulation of cell morphological states that blebbing triggers the formation of plasma membrane-proximal signalling hubs that confer anoikis resistance. Specifically, in melanoma cells, blebbing generates plasma membrane contours that recruit curvature-sensing septin proteins as scaffolds for constitutively active mutant NRAS and effectors. These signalling hubs activate ERK and PI3K—well-established promoters of pro-survival pathways. Inhibition of blebs or septins has little effect on the survival of well-adhered cells, but in detached cells it causes NRAS mislocalization, reduced MAPK and PI3K activity, and ultimately, death. This unveils a morphological requirement for mutant NRAS to operate as an effective oncoprotein. Furthermore, whereas some BRAF-mutated melanoma cells do not rely on this survival pathway in a basal state, inhibition of BRAF and MEK strongly sensitizes them to both bleb and septin inhibition. Moreover, fibroblasts engineered to sustain blebbing acquire the same anoikis resistance as cancer cells even without harbouring oncogenic mutations. Thus, blebs are potent signalling organelles capable of integrating myriad cellular information flows into concerted cellular responses, in this case granting robust anoikis resistance.
Similar content being viewed by others
Main
Attachment-dependent survival is a fundamental aspect of metazoan cell physiology. Points of attachment—either to the extracellular matrix or other cells—generate pro-survival signals using adhesive structures such as focal adhesions13,14 and adherens junctions15. These structures operate as signalling hubs by controlling local concentrations of signalling factors, protein scaffolds and effector proteins. If deprived of attachments, most cells will undergo a form of programmed cell death called anoikis1. Acquisition of molecular strategies that confer anoikis resistance is a critical step in oncogenesis2,3.
Detachment causes most metazoan cell types to become rounded and exhibit dynamic surface blebs4,5,6,7 (Supplementary Video 1). Blebs are hemispherical, pressure-driven cellular protrusions that occur when portions of the cell membrane decouple from the actomyosin cortex8. Although blebbing famously occurs during apoptosis, a second class of ‘dynamic blebs’ are found in healthy cells with intact cortices and are associated with cellular detachment, mitosis and amoeboid motility8,9,10. Detached non-malignant cells maintain dynamic blebbing for only 1–2 h, later undergoing anoikis if attachment is not re-established1,6,11. By contrast, cancer cells, such as those forming melanoma, are highly anoikis resistant and able to indefinitely sustain rounded blebby morphologies in no- and low-attachment environments, often adopting this amoeboid-associated phenotype both in vitro and in vivo—notably at the invasive front of tumours16,17,18,19,20. This increased blebbiness has recently been associated with increased metastatic potential in melanoma and prostate cancer16,21, suggesting that the morphotype might be an indicator of disease aggressiveness.
The correlation between blebby morphologies and metastasis has largely been attributed to the role of blebs in cell motility16,17,18,19, although recent work has begun to suggest that blebbiness might also convey a survival advantage20. This would agree with our previous finding that signalling factors associated with cell survival tend to be concentrated in the vicinity of melanoma blebs, including KRAS and active PI3K16,22. Similar bleb-localized enrichments of ERK1 and ERK2 have also been observed23. Moreover, inhibition of blebbing causes a sharp reduction in local PI3K activity, raising the possibility that blebbing might directly enable this signalling16. Taken together, these pieces of information motivated an investigation of blebbing as a potential contributor to anoikis resistance in melanoma.
Blebbing promotes anoikis resistance
We first inhibited bleb formation using wheat germ agglutinin (WGA), a lectin that reduces plasma membrane deformability by crosslinking components of the glycocalyx8. Using MV3 melanoma cells, we confirmed by morphological motif analysis22 WGA’s effects as a dosage-dependent inhibitor of bleb density on single-cell surfaces (Extended Data Fig. 1a). To investigate putative bleb function as a driver of anchorage-independent cell survival, we designed assays that consistently minimized both cell–substrate and cell–cell attachment over a period of 24 h. We seeded cells at very low density (approximately 100 cells ml−1) in non-adherent culture dishes under constant gentle nutation, which we found necessary to prevent cell aggregation. For controls, we grew cells at identical concentrations in adherent variants of the same culture dishes. Together, these steps generated pairs of ‘detached’ and ‘adhered’ cell populations in paired same-day experiments.
We carried out WGA dose–response survival analysis on three different melanoma cell lines: MV3 (NRAS(Q61R)-driven), A375 (BRAF(V600E)-driven) and M498 (BRAF(V600E)-driven primary cell line). After 24 h of bleb inhibition, detached MV3 cells showed high levels of cell death as measured by ethidium homodimer staining, but little to none when adhered (Fig. 1a). Both adhered and detached M498 cells die after WGA treatment, but detached cells experience significantly more death, both in aggregate and in individual matched experiments (Fig. 1a and Extended Data Fig. 1b). Compared to adherent MV3 cells, adherent M498 cells spread poorly and are easily detached with gentle pipetting (Extended Data Fig. 1c), suggesting poor adhesion even in growth conditions that permit attachment. Accordingly, bleb inhibition induces an increase in cell death also for the adherent population. By contrast, A375 cells were unaffected by WGA treatment, regardless of their attachment state (Fig. 1a). During the design of this viability assay, we observed that the confluency of A375 cells before treatment seemed to influence outcomes, with cells from more confluent culture conditions apparently experiencing more death due to WGA treatment. To examine this systematically, we allowed A375 cells to remain overconfluent for 48 h (changing culture medium daily) before carrying out viability experiments as before. We found that anoikis resistance in these ‘prestressed’ A375 cells was strongly dependent on blebbing, with an effect size even larger than that seen in MV3 cells (Fig. 1a). Having established that A375 cells can be made reliant on blebbing by certain stress conditions, we applied the viability assay also to experiments in which A375 cells were challenged by direct pharmacological stress. Specifically, we measured the effect of bleb inhibition on A375 cells pre-incubated for 48 h in medium containing 10 nM dabrafenib (targeting BRAF(V600E)) and 1 nM trametinib (targeting MEK1 and MEK2), a combination used as a frontline therapy for BRAF-mutated tumours. Drug-treated A375 cells experienced increased cell death after WGA treatment when detached and minimal change in viability when adhered (Fig. 1a).
a, Mean cell death as a function of WGA treatment at different doses for adhered and detached melanoma cells. Cells were grown for 24 h and assayed for cell death using ethidium homodimer staining. All treatment groups were grown and assayed in simultaneous paired experiments, which were carried out three times for each cell line (see Extended Data Fig. 1b for individual experiments). Replicate data were normalized by subtracting negative control values from each treatment group. Error bars represent 95% confidence intervals. All five experimental groups were treated with the same dosages of 0, 2.5, 5, 10 and 20 µg ml−1 WGA. b, Cell surface renderings of representative cells showing the spatial variation of intracellular mean curvature. c, Cell death after bleb inhibition using VitroGel coffins for adhered and detached melanoma cells. Cells were grown for 24 h in either integrin-binding VitroGel RGD (adhered) or non-integrin-binding VitroGel (detached) and assayed for cell death using ethidium homodimer staining. Dots represent individual experiments. See Extended Data Table 1 for individual cell counts.
To ensure that the WGA-mediated disruption of anoikis resistance was due to bleb inhibition and not an off-target effect of WGA, we carried out a complementary experiment using VitroGel, an abiotic hydrogel used in cell culture, to form bleb-restricting ‘coffins’ around individual cells (Fig. 1b). Coffins for ‘detached’ cell growth were made using non-adherent VitroGel, whereas adherent coffins were made using VitroGel RGD, which contained integrin-binding RGD domains that allow for cell adhesion. We carried out these experiments on WGA-sensitive MV3 cells and WGA-insensitive A375 cells. As expected, far more MV3 cells died in non-adherent than in adherent VitroGel coffins, whereas A375 cells showed no difference in viability between the two conditions (Fig. 1c). Hence, dynamic blebbing contributes to anoikis resistance both in NRAS- and in BRAF-mutated melanoma, suggesting that this cell morphological program is a broadly adopted survival strategy in melanoma cells. Moreover, the observation that MAPK pathway inhibition (MAPKi) through dabrafenib and trametinib treatment sensitizes BRAF(V600E)-driven cells to bleb inhibition suggests that bleb formation serves also as an acute defence mechanism against drug attacks.
Blebs recruit septins to cell surface
A notable feature of blebbing versus bleb-inhibited cells is the enrichment of micrometre-scale plasma membrane curvature. From an intracellular viewpoint, curvature is negative (concave) in the bleb proper (reaching values in the region of κ = −1.5 µm−1, in which κ represents the inverse of the radius of curvature) and positive (convex) between the blebs (reaching values in the region of κ = 1.5 µm−1; Fig. 1b). We discovered a marked recruitment of septins near densely packed blebs in MV3 cells expressing mouse SEPT6–GFP (Extended Data Fig. 2a). Indeed, members of the septin cytoskeletal family are the only eukaryotic proteins known to detect the positive micrometre-scale membrane curvature that blebbing creates24,25. The notion that bleb-generated curvature could recruit septins to the plasma membrane after cell detachment is supported by previous work26,27, although we found no reports of an explicit connection between bleb-related surface curvature and septin recruitment. Septins are known to scaffold and regulate factors from diverse signalling pathways28,29,30, including pro-survival pathways in cancer cells such as HIF1 (ref. 31), EGFR32, MET33, JNK34 and HER2 (ref. 35). Therefore, we reasoned that bleb-dependent anoikis resistance could be achieved through recruitment of curvature-sensitive septin proteins to the plasma membrane through bleb-generated curvature, where they promote pro-survival signalling.
To test such an involvement of septins as translators of cell morphology into survival signalling, we carried out mass spectrometry of MV3 cell lysate and found robust expression of several septin proteins (Extended Data Table 2 and Supplementary Table 1). We opted to visualize septin structures in living cells by ectopic expression of mouse SEPT6–GFP26 because this construct has been shown to successfully integrate into the endogenous septin cytoskeleton with little to no effect on septin function26,36. We confirmed proper integration of the fluorescent probe into native septin oligomers by pulldowns of mouse SEPT6–HALO. Mass spectrometry showed that all endogenous septins expressed in MV3 cells were pulled down by SEPT6–HALO, at levels 36 times higher than in control pulldowns with no SEPT6–HALO expression (septins accounted for 0.22% of total protein abundance in the control and 8.1% in SEPT6–HALO pulldown), suggesting that the probe efficiently forms oligomers with endogenous septins (Extended Data Table 2). As previous work has suggested that septins contribute to the retraction of large blebs induced by osmotic shock in T cells26, we measured the blebby surface fraction of MV3 cells with and without SEPT6–GFP expression to be sure that our visualization strategy did not itself alter blebbiness. This analysis showed no difference in blebby surface fraction between the two groups (Extended Data Fig. 2b).
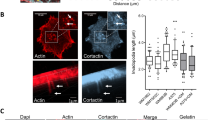
Cell surface renderings of SEPT6–GFP indicated that septin structures are primarily found near blebby cell surface regions, whereas less blebby regions have few to none (Fig. 2a and Supplementary Video 2). To confirm this observation quantitatively, we applied a directional correlation method22, which determines the spatial alignment of cell surface events. Applied to multiple MV3 cells, we confirmed association between bleb occurrence and septin localization (Fig. 2b). Specifically, septins are enriched at positive curvatures above κ = 0.4 µm−1, in agreement with previous reports of high septin affinity for curvature values κ > 0.4 µm−1 (Fig. 2c; refs. 24,25,37). Analysis of control and WGA-treated MV3 cells also confirmed that bleb inhibition reduces mean plasma membrane curvature above κ = 0.4 µm−1 (Fig. 2d). We further examined the localization of septin structures relative to blebs and found them to be enriched at bleb edges, aligned with maximal positive curvature (Fig. 2e). Septin intensities fall steeply towards the bleb centre and more slowly as one moves away from the bleb (Fig. 2e). To ensure that these bleb-associated septin structures are not artefacts of SEPT6–GFP expression, we repeated our analyses on MV3 cells that lacked the construct using immunofluorescence. We found that signal from an anti-SEPT2 antibody was enriched at the cell surface near blebs (Extended Data Fig. 3a–c), localized to the curvy bases of blebs (Extended Data Fig. 3d), and that SEPT6–GFP expression did not alter anti-SEPT2 enrichment near the surface (Extended Data Fig. 3e).
a, Surface renderings of intracellular mean curvature (left) and SEPT6–GFP mean intensity within 1 µm of cell surface (right). b, Directional (Dir.) correlation (cor.) between blebs and SEPT6–GFP in 17 cells (2,025 blebs). Solid orange: cumulative correlative; dashed blue: randomized bleb localization control; dotted black: zero correlation. c, Mean local septin intensity as a function of local intracellular curvature, using the same cells. d, Probability distributions of positive intracellular mean curvature for the cells in Supplementary Fig. 1A; WGA, µg ml−1. e, Mean septin intensity and intracellular mean curvature as a function of distance from bleb edges for cells in b. f, SEPT6–GFP signal in untreated cell, and cells with WGA (50 µg ml−1), VitroGel, H1152 (1 µM; ROCK inhibitor) and NSC668394 (10 µM; ezrin inhibitor). Maximum-intensity projections (MIPs) and 0.16-µm z-slices are shown. Arrowheads, septin accumulation at high curvature. g, Fraction of cortical voxels with septin intensity higher than cytoplasmic mean intensity. WGA, 10 µg ml−1. Dashed lines show quartiles. Basal versus VitroGel was tested with two-sample one-sided t-test pooled variance (P = 0.0130), Shapiro–Wilk (P = 0.257 and 0.310) and two-tailed F-test (P = 0.448). Basal versus WGA was tested with Welch’s t-test (P = 7.13 × 10−6), Shapiro–Wilk (P = 0.453 and 0.794) and two-tailed F-test (P = 0.0114). One asterisk indicates P < 0.05, two indicate P < 0.01, three indicate P < 0.001. h, Data from g as a function of cell surface fraction with intracellular curvature above κ > 0.4 µm−1 (R2 P values = 0.0032, 0.0054 and 0.000027). i, Curvature-sensing mutant SEPT6(ΔAH)–GFP. j, Time-lapse maximum-intensity projections showing SEPT6–GFP. Greyscale: individual time points; pseudo-colour: composite. k, Time-lapse images of SEPT6–GFP during blebbing (0.16 µm z-slice). l, Temporal cross-correlation between SEPT6–GFP intensity and intracellular mean curvature for positive (pos.) and negative (neg.) contigs. The cell was imaged for 4 min; stack acquisition ≈ 0.83 Hz. m, SEPT6–GFP in a cell expressing the SEPT2(33–306) polymerization mutant. n, SEPT6–GFP intensity as a function of distance from bleb edges in cells expressing SEPT2(33–306), overlaying SEPT6–GFP intensity from e. All data are from MV3 cells embedded in soft bovine collagen. Error bands represent 95% confidence interval.
To more thoroughly test the notion that bleb formation as a morphodynamic process causally drives the assembly of septin structures, we perturbed blebbing by a variety of methods: in addition to WGA and VitroGel, we used the ROCK inhibitor H1152 (reduces bleb formation by lowering intracellular pressure) and the ezrin inhibitor NSC668394 (causes large-scale membrane–cortex detachment and thus substitutes single-micrometre-scale, dynamic blebs with large, stable blebs). Every approach had pronounced effects on septin localization, reducing the overall cortical levels with remaining foci confined to high-curvature areas (Fig. 2f). To quantify these observations, we measured cortical septin levels as the surface fraction of cortical voxels (voxels <0.96 µm from the surface) above mean cytoplasmic SEPT6–GFP intensity. Both WGA and VitroGel treatments significantly reduced cortical septin levels relative to control conditions (Fig. 2g). Moreover, all three experimental conditions exhibited a strong linear relationship at the single-cell level between the surface fraction of predicted septin-recruiting curvature (κ ≥ 0.4 µm−1) and cortical septin levels (Fig. 2h). Thus, septin levels at the cortex universally correlate with the presence of septin-recruiting plasma membrane curvature, and experimental conditions that reduce such curvature, such as those inhibiting bleb formation, abrogate septin recruitment. To exclude the possibility that septins localize to bleb edges through a process independent of membrane curvature sensing, we generated mutants of our SEPT6–GFP construct lacking the carboxy-terminal amphipathic helix necessary for curvature sensing by septins24,38 (Extended Data Fig. 4). In contrast to those expressing wild-type SEPT6–GFP, MV3 cells expressing SEPT6(ΔAH)–GFP lacked septin localization to the surface (Fig. 2i and Extended Data Fig. 4). This demonstrates that the observed cortical septin structures depend entirely on the ability of septins to sense the micrometre-scale membrane curvature associated with bleb formation.
Polymerized septins remain at surface
We found gross septin localization to be relatively stable on a timescale of minutes (Fig. 2j), despite bleb lifetimes typically falling below 60 s (ref. 8). This poses the question of how a relatively brief morphodynamic event can determine longer-lived molecular architectures. To address this, we turned to high-speed time-lapse light-sheet microscopy. The resulting data revealed that although ‘pulses’ of septin accumulation can be seen at the curvy bases of blebs, they fade quickly once the bleb is resolved and curvature subsides (Fig. 2k and Supplementary Video 3). To confirm this observation, we computationally identified regions of the surface with high septin intensity that maintained either positive or negative curvature for at least 25 s (the average bleb lifetime, estimated by curvature autocorrelation; Extended Data Fig. 5a). Within these ‘contigs’, cross-correlation analysis between septin intensity and curvature magnitude showed positive correlation in the positively curved dataset, whereas the negatively curved dataset produced the opposite effect (Fig. 2l and Extended Data Fig. 5b). When we subdivided contigs on the basis of septin intensity dynamics, we found that more stable regions were less sensitive to positive curvature than dynamic regions; this is in contrast to the case in negative contigs, which maintained the same septin–curvature relationship regardless of dynamicity. Examining the mean rates of septin intensity gain or loss within individual contigs as a function of dynamicity showed increased rates of change within more dynamic regions (Extended Data Fig. 5c), meaning that in the areas in which septin concentration shifts more frequently, those shifts also occur at faster rates. This suggests the presence of two septin populations at the cell surface: dynamic pools dependent on positive curvature, and more stable pools that are less reliant on specific local curvature profiles.
Cytosolic septins predominantly exist as single hexameric or octameric oligomers that preferentially bind to curved membrane, but quickly detach unless the local concentration becomes high enough to promote inter-oligomer polymerization, which stabilizes membrane binding24,25,39. If the stable septin regions identified by time-lapse imaging were polymerized higher-order structures, it would predict that not only septin curvature sensing, but also septin polymerization when membrane-bound above a critical local concentration, is necessary for the formation of bleb-associated septin structures. To test this, we engineered the SEPT2 mutant SEPT2(33–306), which is capable of integrating into septin oligomers but lacks the amino- and C-terminal domains necessary for inter-oligomeric polymerization40 (Extended Data Fig. 6). When we observed gross septin localization in MV3 cells expressing SEPT2(33–306), we found that septin enrichment at the surface was strongly inhibited in a dominant-negative fashion (Fig. 2m and Extended Data Figs. 2c and 6). Bleb-adjacent septin localization still showed a sharp peak at the curvy bleb edge, indicating a continuation of curvature-sensing septin pulses, but moving away from the bleb the signal decayed rapidly and to lower levels compared to the signal profiles in unperturbed cells (Fig. 2n). Conversely, the signal decay was slowed moving towards the centre of blebs, matching the decay rate in the opposite direction, and the on-bleb signal was significantly increased. This altered bleb-adjacent localization supports a requirement for inter-oligomer polymerization in the formation of curvature-independent stable septin structures and suggests that polymerized septin structures have an increased affinity for non-bleb plasma membrane. This bleb exclusion might be driven by polymerization-dependent crosslinking of the plasma membrane and actomyosin cortex by septins and associated factors, as has been previously observed41.
Our time-lapse data show that most septin pulses endure only as long as the associated blebs, but occasionally a rare de novo formation of stable septin structure can also be observed. In each such instance we noted several pulses occurring in close proximity to one another, resulting in their coalescence into bright long-lived structures. Eventually, their intensities seemed to no longer rely on local curvature profiles (Supplementary Video 4). Taken together, these findings suggest that stable cortical septin structures are formed by iterative bleb-driven curvature events that result in local levels of septin oligomers surpassing the threshold necessary for inter-oligomer polymerization and enabling stabilization through formation of higher-order structures. This would mean that septins are acting as a discrete time integrator of a sustained and spatially persistent dynamic bleb formation process, while also efficiently eliminating the effects of random and isolated morphological events. Thus, septins are ideally suited to translate relevant cell morphological cues into cellular signals.
Anti-anoikic blebbing requires septins
Given the causal relation between bleb formation and septin localization, we sought to determine whether such cortical septin structures would be implicated in the acquisition of bleb-dependent anoikis resistance. Live-cell imaging of poorly adhered cells embedded in soft collagen showed that MV3, M498, prestressed A375 and MAPKi-treated A375 cells possess extensive cortical septin structures spatially associated with blebs (Fig. 3a). By contrast, unperturbed A375 cells, which had shown bleb-independent anoikis resistance (Fig. 1a), exhibited no septin enrichment at the cortex despite robust bleb formation.
a, Mouse SEPT6–GFP probe localization in different melanoma cell lines and conditions. Maximum intensity projections (top) and single z-slices of 0.16 µm thickness (bottom) for representative cells. Cells were embedded in soft bovine collagen. b, Cell death after septin inhibition with 50 µM FCF for adhered and detached melanoma cells. Cells were grown as in Fig. 1b and treated with either FCF or ethanol control for 24 h. Data were normalized by subtracting paired negative control values from each treatment group. Dots represent individual experiments. See Extended Data Table 1 for individual cell counts.
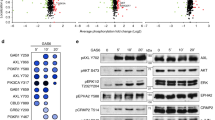
We repeated pulldown mass spectrometry experiments in unperturbed A375 cells and confirmed that the apparent lack of cortical septins was not due to a failure of the fluorescent probe to integrate into native septin structures (Extended Data Table 2). Moreover, SEPT6–GFP pulldown and whole-lysate mass spectrometry datasets showed no major differences in septin expression patterns or oligomer subunit ratios between MV3 and unperturbed A375 cells (Extended Data Table 2). We further carried out western blot analysis of MAPKi dose-dependent expression shifts in SEPT2, SEPT6, SEPT7 and SEPT9. No consistent changes in expression or appearance of bands indicating expression of alternative isoforms could be detected (Extended Data Fig. 7). Hence, the ‘septin activation’ seen in prestressed and MAPKi cells is not driven by changes in septin expression but is post-translationally regulated. Of note, in neither MV3 nor A375 cells did we find SEPT6 to pull down BORG proteins, a family of septin effector proteins known to regulate higher-order polymerization (Supplementary Table 1).
To directly test the conclusion that bleb-associated septin structures regulate survival signalling, we measured anoikis resistance subject to treatment with the septin inhibitor forchlorfenuron (FCF)42. FCF treatment resulted in a near-complete ablation of cortical septin structures without altering blebbiness (Extended Data Fig. 2b) and, as predicted, greatly disrupted anoikis resistance for MV3, M498 and prestressed or MAPKi-challenged A375 cells (Fig. 3b). By contrast, FCF treatment had no effect on unperturbed A375 cells, mirroring bleb inhibition results. To ensure that these results were not due to off-target effects of FCF, we carried out an orthogonal experiment using FCF-sensitive MV3 cells and FCF-insensitive A375 cells, inhibiting septin assembly with the dominant-negative SEPT2(33–306) mutant. Similar to the case for FCF treatment, expression of this construct has no effect on blebbiness (Extended Data Fig. 2b). As with the FCF treatments, genetic inhibition of septins had no effect on A375 cell survival regardless of adhesion state, whereas MV3 cells experienced significant death that worsened with detachment (Extended Data Fig. 8). Taken together, these data suggest that the mechanism allowing blebs to confer anoikis resistance depends on bleb-adjacent cortical septin structures.
Septins scaffold pro-survival signalling
We reasoned that bleb-induced septin structures promote anoikis resistance as a signalling scaffold that amplifies survival signals. To identify signalling candidates we carried out BioID proximity labelling using SEPT6 as bait. We obtained 321 septin-interacting proteins (Supplementary Table 2), among them 3 plasma membrane-localized components of major signal transduction pathways: Notch, CD44 and NRAS. As Notch and CD44 are canonically juxtacrine, we focused on NRAS as a more likely candidate for survival signalling in detached cells. We also found prohibitin43, 14-3-3ζ (ref. 44) and nucleolin45 among the BioID prey, which are all potent effectors of the RAS–RAF–MEK–ERK and RAS–PI3K–AKT pathways. Reexamination of our previous pulldown data revealed that NRAS interacted with SEPT6 in MV3 and prestressed A375, but not in unperturbed A375 (Supplementary Table 1), indicating that the NRAS–septin interaction depends on the assembly of bleb-nucleated septin structures. NRAS was also an especially intriguing candidate as MV3 cells harbour a constitutively active Q61R mutant, leaving open the possibility that suspended NRAS-driven melanoma cells harness their mutational profile through an acute morphological program to empower oncogenic signalling through the RAS–RAF–MEK–ERK and RAS–PI3K–AKT axes.
To support these proteomic results, we confirmed by live-cell imaging that SEPT6–NRAS interaction occurs in rounded, detached cells. Ectopically expressed NRAS–GFP strongly colocalizes with cortical septins (Fig. 4a). Moreover, perturbation of septin structures using FCF or the dominant-negative SEPT2(33–306) mutant significantly reduced enrichment of NRAS near the cell surface (Extended Data Fig. 9a,b). To further investigate how septin inhibition affects NRAS localization, we computed the earth mover’s distance (EMD) as a metric for the spatial heterogeneity of the NRAS signal at the cell surface using homogeneous distributions of the same amount of signal on the same surface as the reference. The results followed our predictions: septin-inhibited cells produced lower EMD values (suggesting diffuse signal distributions) than unperturbed cells (Fig. 4b). Taken together, these data strongly suggest that cortical septin structures bind and scaffold NRAS, affecting its spatial distribution near the surface and increasing its cortical concentration.
a, Spearman correlation between NRAS–septin signal on MV3 cell surfaces and septin–collagen negative control. White data points, cells with correlations not significantly higher than control analysis with one signal scrambled. Signal distributions from median cells are shown on the right. b, EMD between observed NRAS–GFP and homogeneous distribution of equivalent signal for each cell. Control versus FCF: one-sided Mann–Whitney U-test (P = 2.3 × 10−5); normality: Shapiro–Wilk (P = 0.0767 and 0.935). Control versus SEPT2(33–306) Mann–Whitney U-test, one-sided (P = 2.9 × 10−3); normality: Shapiro–Wilk (P = 0.0767 and 0.0393). One asterisk indicates P < 0.05, two indicate P < 0.01, three indicate P < 0.001. c, Effect of septin inhibition on ERK activation levels in adhered and detached MV3 cells. Percentage change in ERK activation between unperturbed and septin-inhibited MV3 cells was measured with ERK-nKTR–GFP. Cells were grown as in Fig. 1b for 12 h. Dots represent individual experiments. d, Effect of detachment on ERK activation levels in unperturbed, septin-inhibited and bleb-inhibited MV3 cells. Percentage change in ERK activation between attached and detached MV3 cells measured with ERK-nKTR–GFP. Cells grown as in Fig. 1b for 12 h. Dots represent individual experiments. e, Spearman correlation between NRAS–AKT-PH and septin–AKT-PH signal on MV3 cell surfaces and septin–collagen negative control. White data points: cells with correlations not significantly higher than control analysis with one signal scrambled. Signal distributions from median cells are shown above. f, Effect of septin or bleb inhibition on PI3K activity in individual MV3 cells. PI3K activity was measured with the AKT-PH–GFP biosensor and is expressed as the fraction of pixels brighter than cytosolic intensity in normalized sum intensity projections. Median cells are on right. Control versus FCF: one-sided Mann–Whitney U-test (P = 1.29 × 10−4); normality: Shapiro–Wilk (P = 0.3126 and 1.99 × 10−4). Control versus SEPT2(33–306): one-sided Mann–Whitney U-test (P = 4.99 × 10−3); normality: Shapiro–Wilk (P = 0.3126 and 2.22 × 10−4). Control versus VitroGel: one-sided Mann–Whitney U-test (P = 1.34 × 10−7); normality: Shapiro–Wilk (P = 0.3126 and 1.04 × 10−5). One asterisk indicates P < 0.05, two indicate P < 0.01, three indicate P < 0.001. See Extended Data Table 1 for individual cell counts.
To determine whether NRAS signalling is important for anoikis resistance in MV3 cells, we overexpressed NRAS(S17N), a dominant-negative mutant known to disrupt NRAS signalling. As for bleb and septin inhibition, perturbation of NRAS had little effect on the survival of adhered cells, but consistently increased death in detached cells (Extended Data Fig. 9c). To determine whether septin organization of NRAS and its effectors affects downstream MAPK signalling, we used the ERK-nKTR nuclear translocation biosensor, which allowed us to monitor the signalling states of individual cells at ultralow cell densities. After 3 h of septin inhibition with FCF, we found adhered cells to have no difference in ERK signalling compared to controls, whereas ERK activation was significantly reduced in detached cells (Fig. 4c). Similarly, we measured the effect of 3 h of detachment versus adhesion on unperturbed MV3 cells, MV3 cells expressing the dominant-negative SEPT2(33–306) mutant and MV3 cells grown in Vitrogel coffins. Detachment of uninhibited cells caused little change in ERK activation, whereas septin and bleb inhibition in detached cells markedly decreased ERK activation (Fig. 4d). Thus, both septins and blebs are necessary for maintaining ERK signalling in detached, but not in adhered, MV3 melanoma cells.
Our previous work has shown that bleb inhibition alters PI3K activity16. As PI3K can be activated by NRAS, we tested whether septin-mediated cortical scaffolding of NRAS(Q61R) would drive a second survival pathway through PI3K signalling. Using the PI3K activity biosensor AKT-PH–GFP33, we found that in poorly adhered, rounded MV3 cells high PI3K activity colocalizes with both septins and NRAS (Fig. 4e). We then carried out septin inhibition, through either FCF treatment or expression of SEPT2(33–306), and measured the resulting effect on PI3K signalling. In both cases, PI3K activity fell significantly (Fig. 4f). Similarly, bleb inhibition with VitroGel coffins greatly depleted PI3K activity, in agreement with our previously published results showing the same effect following bleb inhibition by WGA16 (Fig. 4f). Together, these data show that bleb and septin inhibition specifically reduces both ERK and PI3K activation levels in detached MV3 melanoma cells, supporting the idea that bleb-nucleated cortical septin structures promote anoikis resistance through the scaffolding and upregulation of NRAS–RAF–MEK–ERK and NRAS–PI3K–AKT signalling.
Blebbing fibroblasts can resist anoikis
Most metazoan cells will round and begin blebbing after detachment from substrate4,5,6,7 (Supplementary Video 1). Over the course of 1–2 h, this blebbing will subside, as excess membrane is endocytosed away from the cell surface6,11. Many cancer cells, however, indefinitely sustain blebbing in low- and no-attachment conditions, an amoeboid phenotype seen at the invasive front of tumours in vivo16,17,18,19,20. Following the discovery of blebbing as a driver of survival signalling in melanoma cells, we wondered whether anoikis resistance can be conferred to non-malignant mammalian cells simply by prolonging post-detachment blebbing.
We first determined whether detached non-malignant cells form bleb-associated septin structures during the 1–2-h window before bleb attenuation. Both mouse embryonic fibroblast (MEF) cells and human epithelial kidney 293 cells expressing SEPT6–GFP exhibited bright bleb-associated cortical septin structures when imaged before bleb attenuation. After blebbing ceased, cortical septins were absent (Fig. 5a and Extended Data Fig. 10a). As attenuation of blebbing is primarily driven by dynamin-dependent clathrin-mediated endocytosis11, we analysed blebbing in MEF cells expressing the dominant-negative dynamin mutant DYN2(K44A), expecting that this mild impairment of clathrin-mediated endocytosis efficiency would prolong post-detachment blebbing. Indeed, about 80% of DYN2(K44A)-expressing MEFs maintained blebbiness 3 h post-detachment, whereas for control MEFs the fraction of blebbing cells fell below 10% over the same time period (Fig. 5b).
a, Recently detached MEF cells expressing SEPT6–GFP, imaged either before or after bleb attenuation. Maximum-intensity projections (above) and single z-slices of 0.16 µm thickness (below) are shown for representative cells. Cells were embedded in soft bovine collagen. b, Fraction of MEF cells showing blebby morphologies 90, 120 and 180 min after detachment from substrate. Orange data points indicate control MEF cells and green data points indicate MEFs expressing DYN2(K44A) in paired, same-day experiments (solid or dashed lines indicate paired data). Representative cells after 180 min of detachment are shown to the right. c, Additional caspase activation after 4 h in detached MEF cells compared to paired adhered cells. Orange data points indicate control MEF cells and green data points indicate MEFs expressing DYN2(K44A). Red data points indicate DYN2(K44A)-expressing cells treated with 10 µg ml−1 WGA and purple indicates DYN2(K44A)-expressing cells treated with 50 µM FCF. Caspase activity was measured using CellEvent caspase-3/7 biosensor. See Extended Data Table 1 for individual cell counts.
To determine the effects of prolonged blebbiness on anoikis resistance, we measured caspase activity after 4 h of detachment using the CellEvent biosensor. This indicator of viability was chosen because DYN2(K44A) expression has been observed to reduce cell viability after ≈20 h of expression (M. Mettlen, personal communication), making it incompatible with the overnight viability assays used in other experiments. To ensure that this assay is able to detect bleb-dependent changes in anoikis resistance, we measured the effect of bleb inhibition on caspase activity in adhered and detached MV3 cells grown for 4 h. In agreement with our overnight ethidium homodimer viability assays (Fig. 1a), we found a consistent and specific increase of caspase activation in detached bleb-inhibited MV3 cells (Extended Data Fig. 10b).
We then repeated this experiment on MEF cells and found that the blebbing DYN2(K44A)-expressing cells were far more resistant to anoikis than matched same-day controls (Fig. 5c and Extended Data Fig. 10c). To ensure that the difference in caspase activity was not due to an off-target effect of dynamin inhibition, we treated cells with either the bleb inhibitor WGA or the septin inhibitor FCF. In both cases, we found that caspase activation was returned to levels seen in matched non-blebbing control cells, meaning that the anoikis resistance acquired by DYN2(K44A) depends on both bleb formation and septins (Fig. 5c and Extended Data Fig. 10c). These data show that cancer cells derived from human melanomas and non-malignant fibroblasts derived from mouse embryos all seem to use the same bleb- and septin-dependent anoikis-resistance strategy.
Discussion
Bleb formation has long been recognized as a morphological program associated with metastatic melanoma and other aggressive cancers16,17,18,19,20. Although this program has been largely interpreted as a means of amoeboid migration, our study now shows that dynamic blebbing prompts the construction of septin signalling hubs that substitute for the loss of anchorage-dependent signalling activity caused by substrate detachment. Once constructed, such signalling would provide improved survival for cancer cells travelling through low-adhesion environments such as blood, lymph and stroma. Our experiments with MEFs demonstrate that bleb-based signalling hubs and the anoikis resistance they confer are not oddities specific to melanoma, but rather represent a distinct detachment-activated signalling pathway that is conserved at least across the evolutionary divide between mice and humans. Although it is unknown what other cell types might use this bleb–septin signalling paradigm, the widely observed tendency of metazoan cells to bleb immediately following detachment leads to the conclusion that this strategy might be more broadly distributed and ancient than our current results can unveil.
Septins are highly versatile protein scaffolds that organize and regulate diverse signalling pathways across multiple kingdoms of life28,29,30. Although our data specifically support the hypothesis that bleb-associated septins drive anoikis resistance in melanoma cells through scaffolding of NRAS and downstream signals, we see no reason to exclude other pathways following the same paradigm. Indeed, according to our proteomic analyses of MV3 cells, septins interact with both CD44 and Notch, which—despite a canonical involvement in juxtacrine signalling—have both been demonstrated to transduce signal in the absence of activation by neighbouring cells or extracellular matrix46,47. Similarly, dynamic blebbing does not occur only in mammalian cells challenged with detachment, but has also been observed in stem cells48,49,50, migratory cells9,10,16 and cells undergoing mitosis10,51. With this in mind, we conjecture that this bleb–septin signalling modality is not exclusively RAS targeting or survival oriented, but rather reflects the diversity of cell types and behaviours with which it is associated.
The oncogenic alteration NRAS(Q61R) is classically thought to drive constitutive upregulation of MAPK signalling. Our results show that this activating alteration is not always sufficient to maintain downstream signalling at levels that guarantee cell viability. Previous work has shown that wild-type and mutant RAS function is regulated through the formation of nanoclusters, small regions of extremely high RAS protein density52. Our experiments now show that septins serve to modulate local NRAS concentration, thereby affecting downstream signal amplitude. Septin scaffolding is also likely to amplify downstream signals by influencing NRAS interactions with its effectors, such as those found in our proteomic studies. In either case, this finding highlights the functional gap between the genomic condition of possessing an oncogene, and the cellular condition of experiencing penetrant oncogenic signalling. Moreover, it suggests the consideration of septins as a potential therapeutic target for NRAS-mutant melanoma, a disease associated with poor prognoses and limited therapeutic options at present owing to the lack of targeted drugs53. FCF is a septin inhibitor that has been used as an experimental tool in vitro for over a decade42, with recent studies showing it is effective and well tolerated in mice54. Furthermore, FCF has been in use as a US government-approved agricultural chemical since 2004, and studies by the Environmental Protection Agency suggest that it possesses low systemic toxicity in humans55, making it (and its chemical derivatives) promising therapeutic candidates.
Our results also demonstrate the potential of such therapeutic strategies against BRAF-mutant melanoma—either as primary therapies as suggested by our M498 results, or adjuvant therapies alongside inhibitors of BRAF(V600E), MEK and other components of the MAPK pathway as suggested by our A375 results. Our finding of dynamic blebbing conferring resistance to targeted therapies in melanoma agrees with work that links ROCK activation, which directly drives bleb formation by promoting actomyosin contraction, to melanoma therapeutic resistance in vitro, in mouse models and in patients20. Furthermore, treatment of melanoma cells with MAPKi therapy sensitizes them to blockade of amoeboid morphologies through ROCK inhibition, greatly increasing cell death compared to MAPKi alone. This closely matches our finding that A375 is sensitized to both bleb and septin inhibition by MAPKi, and that co-inhibition of either blebbing or septins produces significantly more cell death than MAPKi alone. Non-genetic adaptive resistance to MAPKi is achieved through the relaxation of negative feedback on NRAS by ERK56, with RAF–MEK–ERK signalling remaining low owing to sustained MAPKi therapy while survival is maintained by increased NRAS activity feeding into the PI3K pathway. We propose that disruption of bleb formation and/or cortical septin assembly in MAPKi-resistant BRAF(V600E) cells renders the signal produced by the less basally active wild-type NRAS insufficient for cell viability. Support for this proposal is provided by our SEPT6 pulldown experiments, which show NRAS–septin interactions in ‘septin-activated’ MV3 and prestressed A375 cells, but not in ‘septin-inactive’ basal A375 cells that lack such structures. The nature of septin activation in prestressed and drug-challenged A375 cells remains mysterious. Our data show no changes in septin expression between conditions, indicating post-translational regulation of septins at play. Future work will be necessary to understand the mechanism driving this disease-relevant regulation of the septin cytoskeleton.
In the bleb–septin–NRAS signalling pathway described here, information flow is transduced from chemical (morphogenic signalling), to spatial (blebbiness), and back to chemical (survival signalling), with septins acting as a spatiochemical translator and time integrator of pulsatile spatial information (Extended Data Fig. 11). Although control of morphology is often thought of as a terminal objective of cell signalling, consideration of cellular morphodynamics as an upstream input to signal transduction57 and metabolic regulation58 defines an emerging paradigm in cancer regulation. Such morphological cues would act not only as potent drivers of signalling, but also as integrators of cell-autonomous and cell-external information flows. For example, dynamic blebbing is typified by low Rac1 activity, high Rho and ROCK activity, low ERM engagement, high actomyosin contraction, low glycocalyx density and low attachment12,59. All of this information is combined into a state that is interpretable as a single signal transduction node by septins, which can then be activated to form versatile signalling hubs capable of regulating myriad signalling pathways.
Methods
Cell culture and reagents
MV3 cells60 were a gift from Peter Friedl (MD Anderson Cancer Center, Houston TX). A375 cells61 (ATCC CRL1619) were acquired from ATCC. M498 cells62 were acquired from Sean Morrison (UT Southwestern, Dallas, TX). MEF (3T3) cells were acquired from BD Biosciences, Clontech (no. C3018-1). Cell lines were authenticated using GenePrint 10 System from Promega. All cell lines were tested for mycoplasma contamination using Genlantis MycoScope PCR Detection Kit (Fisher). MV3, A375, M498 and MEF cells were cultured in DMEM (Gibco) supplemented with 10% fetal bovine serum (FBS; ThermoFisher) at 37 °C and 5% CO2. We carried out targeted deep sequencing of 1,385 cancer-related genes showing sequence and copy number variations in MV3, A375 and M498, the results of which are publicly available63. ‘Prestressed A375’ cells were generated by allowing A375 cells to grow to confluency and then for an additional 48 h without passaging, with medium changed daily. ‘MAPKi A375’ cells were generated by treating A375 cells with a combination of 10 nM dabrafenib and 1 nm trametinib (alternative concentrations used where noted in the text) for 48 h.
Inhibitors
WGA was purchased from Sigma (product no. L9640). FCF was purchased from Sigma (product no. C2791). VitroGel64 and VitroGel RGD were purchased from TheWell Bioscience (sku nos. TWG001 and TWG002). H1152 was purchased from Tocris (catalogue no. 2414). NSC6683394 was purchased from Sigma (product no. 341216). Dabrafenib (GSK2118436) and trametinib (GSK1120212) were purchased from Selleckchem. Transient expression of dominant-negative Dyn2(K44A) (refs. 65,66) was achieved through adenovirus transduction67 as previously described68 with cells analysed 16–18 h after transduction.
Recombinant DNA constructs
Mouse SEPT6–GFP construct was purchased from Addgene (Addgene plasmid no. 38296) and was cloned into the pLVX-IRES-puro vector (Clontech). The GFP–AKT-PH construct was obtained from the laboratory of Jason Haugh (North Carolina State University, Raleigh NC) and cloned into the pLVX-IRES-puro vector (Clontech). The GFP–tractin construct was a gift from Dyche Mullins (Addgene plasmid no. 58473) and was cloned into the pLVX-IRES-puro vector (Clontech). The BioID2 (ref. 69) construct was obtained from Addgene (Addgene plasmid no. 74223) and cloned onto the N terminus of SEPT6 from SEPT6-GFP-pLVX-IRES-puro, replacing eGFP but maintaining the same 22-amino-acid linker. SEPT6–HALO was made by cloning the HALO tag from pFN21K (Promega catalogue no. G2821) onto the N terminus of SEPT6 from SEPT6-GFP-pLVX-IRES-puro, replacing eGFP but maintaining the same 22-amino-acid linker. ERK-nKTR–GFP70 was purchased from Addgene (Addgene plasmid no. 59150). C-terminally mGFP-tagged human NRAS in pLenti-C-mGFP was purchased from OriGene (OriGene catalogue no. RC202681L2). C-terminal mGFP was removed and eGFP tag was cloned onto the N terminus after aberrant localization was observed following expression in MV3 cells (presumably due to steric inhibition of C-terminal palmitoylation and farnesylation domains by the C-terminal mGFP tag). NRAS(S17N) (ref. 71) mutant was generated by cloning the S17N alteration into untagged NRAS-pLenti construct using HiFi assembly. pBOB-Septin2-GFP was purchased from Addgene (Addgene plasmid no. 118734), amino acids 1–32 and 307–361 were removed through PCR to create SEPT2(33–306), and the construct was cloned into pLVX-IRES-puro vector (Clontech). SEPT6(ΔAH)–GFP was generated by removing amino acids 355–372 of SEPT6 from SEPT6-GFP-pLVX-IRES-puro through PCR. H2B–mCherry was obtained from Addgene (Addgene no. 89766). Cells expressing lentiviral vectors were created by following the manufacturer’s instructions for virus preparation and cell infection (Clontech). Cells were selected for expression by treatment with puromycin or G418, or by using fluorescence-activated cell sorting.
Detached and adhered cell culture
Cells were grown to 70–80% confluency, trypsinized for 3 min, resuspended at approximately 100 cells ml−1 (measured by eye using light microscopy) in DMEM (Gibco) supplemented with 10% FBS (ThermoFisher) and detached from one another by repeated pipetting. Drug treatments, solvent-only controls or CellEvent caspase activity sensor was added to suspensions if appropriate to the experiment, which were then immediately transferred to uncoated (for detached cells) or ibiTreated (for adhered cells) 8-well ibidi µ-slides (ibidi catalogue no. 80821 and 80826) at 200 µl per well. Both slides were stored at 37 °C and 5% CO2 for 24 h for viability assays, 3 h for ERK activity assays or 4 h for caspase activation assays. Uncoated slides (detached) were nutated at 20 r.p.m. during this time. At the end of this period, detached cells were examined by microscopy to confirm that cells did not aggregate—if aggregates were found, the experiment was discarded and repeated. Results from protocol optimization show that this was most commonly caused by wells being seeded at too high a cell density. For experiments using MAPKi A375 cells, treatment with dabrafenib and trametinib was continued over the course of the experiment.
Cells embedded in three-dimensional collagen
Collagen gels were created by mixing bovine collagen I (Advanced Biomatrix 5005 and 5026) with concentrated phosphate-buffered saline (PBS) and water for a final concentration of 2 mg ml−1 collagen. This collagen solution was then adjusted to pH 7 with 1 N NaOH and mixed with cells immediately before incubation at 37 °C to induce collagen polymerization. Cells were suspended using trypsin–EDTA (Gibco), centrifuged to remove medium, and then mixed with collagen immediately before incubation at 37 °C to initiate collagen polymerization. To image collagen fibres, a small amount of collagen was conjugated directly to AlexaFluor 568 dye and mixed with the collagen sample immediately before polymerization. Poorly adhered cells were enriched in samples by visualizing immediately after collagen polymerization, and visually selecting for those exhibiting rounded, blebby morphologies.
Cells grown in VitroGel coffins
VitroGel coffins were prepared by embedding MV3 or A375 cells in VitroGel 3D or RGD at 1:1 dilution according to the manufacturer’s instructions (TheWell Biosciences, sku no. TWG001). Briefly, cells were grown to 70–80% confluency, trypsinized and diluted to approximately 500 cells ml−1 in DMEM (Gibco) supplemented with 10% FBS (ThermoFisher) and detached from one another by repeated pipetting. A 1:1 solution of VitroGel and VitroGel Dilution Solution was made, and added to cell suspension at a ratio of 4:1 (VitroGel/cells) using gentle pipetting until well mixed (with care taken not to form bubbles). Cell–VitroGel solution was transferred to an 8-well uncoated ibidi µ-slide (ibidi catalogue no. 80821) at 100 µl per well, spread over the bottom of wells with a pipette tip and allowed to polymerize at room temperature for 15 min. The remainder of the well was then filled with 10% FBS DMEM, and the slide was stored at 37 °C and 5% CO2 for 24 h for viability assays, 3 h for ERK activity assays or 1 h for visualization of septin and PI3K activity.
Two-dimensional immunofluorescence
Wild-type MV3 or MV3 cells expressing SEPT6–GFP were seeded in MatTek glass-bottom coverslip dishes (P35G-1.5-14-C) at 50,000 cells and incubated overnight. The following day, the cells were washed three times with 1× PBS and fixed with 37 °C prewarmed 4% PFA dissolved in cytoskeletal buffer (10 mM PIPES, 100 mM NaCl, 300 mM sucrose, 1 mM EGTA, 1 mM MgCl2, 1 mM dithiothreitol and protease inhibitor cocktail). The cells were incubated with PFA at 37 °C for 20 min. The MV3-Sept6–GFP cells were then imaged on the Nikon Eclipse Ti widefield epifluorescence microscope with a 100× Plan APO oil-immersion lens using NIS-Elements 4.30.02 (Build 1053). For immunofluorescence staining of wild-type unlabelled cells, the cells were then permeabilized with 0.5% Triton X-100 for 20 min and then blocked with 5% bovine serum albumin (BSA) in 1× PBS for 1 h. Following blocking, the cells were then treated with rabbit anti-SEPT2 primary antibody (Sigma, HPA018481) at a 1:1,000 dilution overnight at 4 °C. The cells were then washed with 1× PBS for 10 min for a total of three times and then treated with donkey anti-rabbit-488 secondary antibody (Thermo, A-21206) at a final dilution of 1:5,000 for 1 h. The cells were then washed with 1× PBS for 10 min for a total of three times and then imaged on the Nikon Eclipse Ti widefield epifluorescence microscope.
Three-dimensional immunofluorescence
MV3 cells were collected from 10-cm dishes and seeded on Corning Ultra-Low Attachment culture dishes (Sigma, CLS3262) for about 2 h for the cells to acquire their blebby architecture. The cells were then centrifuged at 1,000 r.p.m. for 5 min. The pellet was gently washed once by resuspending in 1× PBS to remove any medium and centrifuged. The pellet was then resuspended and fixed in 100 µl of prewarmed 37 °C, 4% PFA dissolved in cytoskeletal buffer (10 mM PIPES, 100 mM NaCl, 300 mM sucrose, 1 mM EGTA, 1 mM MgCl2, 1 mM dithiothreitol and protease inhibitor cocktail) for 20 min at 37 °C. The cells were then washed with PBS, and centrifuged. To permeabilize the cells, the pellet was resuspended in saponin for 20 min. The cells were then centrifuged, and the pellet was blocked by resuspending in 5% BSA for 1 h with gentle rocking for uniform blocking. The cell suspension was then centrifuged and resuspended in rabbit anti-SEPT2 primary antibody (Millipore Sigma, HPA018481) at a concentration of 1:1,000 overnight at 4 °C. The following day, the cells were centrifuged, and washed once in 10 ml PBS. The cells were then centrifuged and resuspended in donkey anti-rabbit-488 secondary antibody (Thermo, A-21206) at a final dilution of 1:5,000 for 1 h. The cells were then centrifuged and washed twice with 10 ml 1× PBS by centrifugation. The pellet was then resuspended in AF568–phalloidin (Invitrogen, A12380) for 1 h, washed, DAPI-stained for 5 min, washed and embedded in soft bovine collagen as described above.
Viability, ERK activity and caspase activity cell counting assays
Viability, ERK activity and caspase activity assays were analysed using live-cell fluorescence and phase-contrast microscopy, carried out on a Nikon Ti microscope equipped with an environmental chamber held at 37 °C and 5% CO2 at ×20 magnification. For viability assays, cells were stained with ethidium homodimer 1 (Invitrogen catalogue no. E1169) at 4 µM and Hoechst (ThermoFisher catalogue no. H3570) at 10 µg ml−1 for 15 min before imaging. Live and dead cells (as identified by cellular ethidium signal) were counted using the Cell Counter ImageJ plugin. For ERK activity assays, live cells carrying ERK-nKTR–GFP and H2B–mCherry were imaged after 3 h of experimental conditions and cells with high and low ERK activity were counted with the Cell Counter ImageJ plugin. Cells in which nuclear GFP signals were by eye higher than cytoplasmic signal (‘brighter’ nucleus visible within ‘darker’ cytoplasm) were labelled ERK low, whereas cytoplasmic signals equal to or higher than nuclear signal (nucleus indistinguishable from cytoplasm or ‘darker’ nucleus within ‘brighter’ cytoplasm) were labelled ERK high. To control for unconscious bias in making this determination, the analyst was blinded to the identity of all ERK experimental groups. For caspase activity assays, cells were treated with CellEvent caspase-3/7 green detection reagent (ThermoFisher catalogue no. C10423) according to the manufacturer’s instructions at a final concentration of 8 µM. Cells were treated following introduction to adhered or attached culture conditions, as described above. After 4 h, cells were treated with Hoechst (ThermoFisher catalogue no. H3570) at 10 µg ml−1 for 15 min and imaged. Cells positive and negative for caspase activity (as identified by perceptible cellular CellEvent signal) were counted using the Cell Counter ImageJ plugin.
Proteomics
MV3 cells, MV3 cells expressing SEPT6–HALO, A375 cells and A375 cells expressing SEPT–Halo were each plated in 150-mm dishes at approximately 7.5 × 106 cells per dish and grown to 70–80% confluency. Pulldowns were carried out as per the manufacturer’s instructions (Promega, G6504). Cells were washed and collected in ice-cold PBS. Cells were pelleted at 2,000 RCF for 10 min at 4 °C. Pellets were stored at −80 °C for 72 h. Pellets were then thawed at room temperature, lysed in mammalian lysis buffer supplemented with protease inhibitor cocktail (Promega). The lysate was homogenized with a 27G syringe and centrifuged at 14,000 RCF for 5 min at 4 °C. The supernatant was diluted in TBS and incubated with pre-equilibrated HaloLink resin (Promega) at room temperature with rotation for 15 min. Resin was then washed four times with resin wash buffer (Promega). Complexed proteins were eluted in SDS elution buffer (Promega) for 30 min at room temperature. Eluted samples and whole-cell-lysate controls were loaded and run on a 10% Mini-PROTEAN TGX protein gel (Bio-Rad), visualized with AcquaStain (Bulldog Bio), excised and analysed with an Orbitrap Fusion Lumos using reverse-phase liquid chromatography with tandem mass spectrometry. Mass spectrometry data were analysed using Proteome Discoverer 2.2 and searched using the human protein database from Uniprot.
For BioID proximity labelling, approximately 4 × 107 cells were incubated in DMEM (Gibco) supplemented with 10% FBS (ThermoFisher) and 50 µm biotin for 16 h at 37 °C and 5% CO2. Cells were washed twice in 1× PBS and lysed with 1:1 dilution of 2xJS buffer (100 mM HEPES pH 7.5, 300 mM NaCl, 10 mM EGTA, 3 mM MgCl2, 2% glycerol, 2% Triton X-100) containing HALT phosphatase–protease cocktail (ThermoFisher no. 23225). Cells were collected using a cell scraper, Triton X-100 was added to 2%, and the resulting mixture was put on ice and sonicated. An equal volume of chilled lysis buffer was added and the mixture was sonicated again before centrifugation at 16,500 RCF for 10 min. The supernatant was collected and incubated overnight with Dynabeads (ThermoFisher no. 65602) at 4 °C. Beads were magnetically collected and supernatant was removed. Beads were washed four times with 50 nM Tris-Cl, pH 7.4 with 8 M urea and supernatant was removed completely. Beads were resuspended in Laemmli buffer and biotinylated proteins were eluted by boiling for 5 min. Supernatant was loaded and run on a 10% Mini-PROTEAN TGX protein gel (Bio-Rad), visualized with AcquaStain (Bulldog Bio), excised and analysed with an Orbitrap Fusion Lumos using reverse-phase LC–MS/MS. MS data were analysed using Proteome Discoverer 2.2 and searched using the human protein database from Uniprot.
Western blots
Cells were lysed in RIPA buffer (50 mM Tris HCl, 150 mM NaCl, 0.5% (w/v) sodium deoxycholate, 1.0 mM EDTA, 0.1% (w/v) SDS, 1.0% (v/v) NP-40 and 0.01% (w/v) sodium azide; pH 7.4). Proteins were run on precast gels (Bio-Rad no. 4568123) and transferred onto PVDF membranes. After transfer, the membranes were rinsed in TBS buffer, air dried at room temperature for 20–30 min, and rewet in TBS buffer supplemented with 0.5% Tween-20 (TBST). The membranes were then blocked for 30 min in 5% BSA for septins or in 5% non-fat dry milk for vinculin. The blocking BSA or milk was dissolved in TBST. Membranes were incubated rocking in blocking solutions with primary antibodies overnight at 4 °C, and the secondary antibodies were rocked at room temperature for 1 h. We used the following primary antibodies: SEPT2 (Sigma no. HPA018481, 1:1,000), SEPT6 (Sigma no. HPA005665, 1:1,000), SEPT7 (Sigma no. HPA023309, 1:1,000), SEPT9 (Sigma no. HPA042564, 1:1,000) and vinculin (Santa Cruz Biotechnology no. sc-25336, 1:1,000). The secondary antibodies used were goat anti-mouse (Invitrogen no. 31430, 1:20,000) and goat anti-rabbit (Invitrogen no. G21234, 1:20,000). Proteins were detected using enhanced chemiluminescence reagents (Thermo Scientific no. 34095) and imaged on the PVDF membrane (Thermo Scientific no. 88518) using the G:Box imager (SYNGENE) and GeneSys software. Protein band densitometry was carried out in ImageJ.
Three-dimensional light-sheet imaging
Three-dimensional (3D) samples were imaged using two variants of axially swept light-sheet microscopy72,73, the first of which provides sub-400 nm isotropic raw resolution, and the second near-isotropic at about 400 × 400 × 450 nm, uniformly maintained throughout large fields of view of about 100 × 100 × 100 µm. The first variant is equipped with 40×, NA 0.8 Nikon illumination and detection objectives, and the second is equipped with an NA 0.67 Special Optics illumination objective and a 25×, NA 1.1 Nikon detection objective. For very fast imaging, for which aberration-free remote focusing of the illumination light sheet becomes rate-limiting (faster than approximately 0.1 Hz full-volume acquisition, depending on cell size), these microscopes could also be operated in traditional light-sheet microscopy mode. Here, the numerical aperture of the illumination beam was reduced to cover a field of view of about 20 μm, and imaging was carried out by scanning the illumination light sheet synchronously in the Z direction with the piezo-mounted detection objective.
Samples were imaged in phenol red-free DMEM containing 25 mM HEPES (ThermoFisher) with 10% FBS and antibiotic–antimycotic (Gibco), held at 37 °C during imaging. Images were collected using sCMOS cameras (Orca Flash4.0 v2, Hamamatsu), and the microscopes were operated using custom LabView software. All software was developed using a 64-bit version of LabView 2016 equipped with the LabView Run-Time Engine, Vision Development Module, Vision Run-Time Module and all appropriate device drivers, including NI-RIO drivers (National Instruments). The software communicated with the camera through the DCAM-API for the Active Silicon Firebird frame-grabber and delivered a series of deterministic TTL triggers with a field-programmable gate array (PCIe 7852R, National Instruments). These triggers included analog outputs for control of mirror galvanometers, piezoelectric actuators, laser modulation and blanking, camera fire and external trigger. All images were saved in the OME-TIFF format. The microscope control software is freely available to academic and nonprofit institutions on completion of a material transfer agreement with the University of Texas Southwestern Medical Center.
3D cell image analysis
Cell morphology and septin localization were analysed principally using u-shape3D22, with all exceptions (Figs. 2g,h,l and 4a,e,f and Extended Data Figs. 5a,b and 9a,b) described below. Briefly, 3D images were first deconvolved using either a Richardson–Lucy or Wiener algorithm with an experimentally measured point spread function. Next, cells were segmented from the image background using the twoLevel mode of u-shape3D, which combines a straightforward Otsu threshold74 of the image to detect the outer cell surface with a blurred version of the image to segment the inside of the cell. The volume segmented from the blurred image was morphologically eroded to ensure the fidelity of the overall segmentation. Cell surfaces were then represented as triangle meshes, and the mean surface curvature at every triangle was calculated as previously described22,75. To remove irregularities, the curvature is next smoothed in real space with a median filter of 1 pixel to remove infinities and then slightly diffused along the mesh.
Septin and NRAS localization was measured from fluorescence images by extending a sphere of either 1 µm (for septins) or 2 µm (for NRAS) about each mesh triangle and mapping to the triangle the mean intensity of intracellular signal within that sphere. Unlike in previous studies22,75, the image intensity was not depth normalized before analysis, but was instead measured from the raw, undeconvolved image.
Blebs were also detected as previously described22. Machine learning models trained using images labelled by three separate expert annotators were combined through voting to classify blebs. Distances from bleb edges were then calculated as the geodesic distance from each triangle to the nearest bleb edge. The blebby surface fraction was also defined as the percentage of total mesh triangles classified as ‘on bleb’. Bleb and septin directional correlations were calculated using spherical statistics22, in particular by fitting spherical normal distributions to distributions defined at each mesh triangle.
Cortical septin levels were quantified by measuring the mean cytoplasmic intensity of each cell using hand-drawn regions of interest that excluded nuclei (carried out with the FIJI ImageJ package), and then calculating for each cell its fraction of cortical voxels (intracellular voxels within 0.96 µm of the u-Shape3D-derived surface) that were higher than that cell’s mean cytoplasmic intensity (carried out using a MATLAB script). NRAS enrichment at the surface was quantified by first calculating the total amount of intracellular signal expected to be within 0.96 µm of the u-Shape3D-derived surface if the cell’s signal was homogeneously distributed between all voxels, and then calculating the percentage change between this value and the observed value for each cell (carried out using a MATLAB script). To quantify PI3K biosensor fluorescence signal, cells were segmented with u-shape3D and the intracellular signal was summed across the z axis to yield a sum projection image (carried out with the FIJI ImageJ package). To account for differential biosensor expression levels, projections were normalized by adjusting brightness until the mean cytoplasmic signals of all images were approximately the same. PI3K activity was then quantified as the fraction of total pixels with intensities brighter than a threshold value (the approximate upper range of cytoplasmic signal), which was held constant across cells (carried out with the FIJI ImageJ package).
Colocalization of fluorescent signal distributions across 3D cell surfaces was quantified by calculating the Spearman’s rank correlation coefficients between signals on a cell-by-cell basis. To approximate the significance of these correlations, a variation of Costes’ randomization76,77 was used, in which one distribution was randomized and the Spearman coefficient was calculated again. This process was repeated 1,000 times, noting the final fraction of randomized Spearman coefficients greater than that calculated for the data as observed. This fraction is referred to as a P value (not to be confused with the P value output by statistical tests of significance). As the sampling of the local intensity of a surface signal through overlapping spheres (as done with u-Shape3D) produces a smoothing effect that leads to spatial autocorrelation, signal distributions were downsampled to below the level of spatial correlation to ensure independence of data points. This was accomplished by selecting 200 random equidistant points on a cell’s surface78 before Costes’ randomization was carried out on these points. To ensure that this random selection did not bias the analysis, the process was repeated 10 times, with the resulting P values averaged to a single mean P value. If the mean P value was less than 0.05, the colocalization of the tested signals was deemed significant.
To quantify NRAS distributions, EMD79,80 was measured on discrete surfaces80. For a single cell, the measured intensity of NRAS–GFP signal at the surface was modified in two steps to compensate for cell-to-cell variation in fluorescence intensity. As we were interested only in bright areas representing high NRAS density, a surface background defined as the median of surface intensity was subtracted from the measured surface intensity, and the resulting values were normalized to the mean cytoplasmic intensity derived manually as described above. Resulting negative intensity values along the triangle mesh surface were set to zero. For each cell, the EMD measured the distance between the modified NRAS signal distribution on each cell surface and homogeneous distribution of the same amount of signal on the same surface (that is, the larger the EMD the more spatially organized is the tested signal). These operations were carried out using a MATLAB script.
Analysis of surface curvature and septin time series
The goal of these time series analyses was to test the central hypotheses that septin intensity co-fluctuates with curvature and that increases in septin intensity on stable structures or associated with de novo assembly are significantly enriched on positive intracellular curvature. Substantial preprocessing was required to bring volumetric time-lapse datasets of SEPT6–GFP fluorescence into a format enabling such analyses. The processing steps fell into two categories, as detailed below: conversion of raw data into analytically tractable mesh datasets (including surface segmentation, registration, curvature measurement, intensity measurement, bleach correction and smoothing); and identifying portions of these datasets suitable for statistical inference of the coupling between curvature and septin accumulation (including gating curvature, intensity and intensity change data, and isolating contiguous regions of the surface that exhibit the desired characteristics for significant amounts of time). Then, using the prepared data, we assessed the coupling of septin intensity and curvature fluctuations by cross-correlation on contiguous regions of negative and positive intracellular curvature, measured signal dynamicity within these regions, and determined the relationship between septin signal dynamicity and the magnitude of septin intensity changes within these regions.
Registration
To extract time series, the segmented mesh from the first frame (3D vertex coordinates, \({{\bf{x}}}_{0}{\boldsymbol{=}}\left({x}_{0},{y}_{0},{z}_{0}\right)\) and face connectivity) must be consistently tracked over time. To achieve this, we first removed whole-cell motion with rigid-body registration. We then applied non-rigid diffeomorphic registration81 on the pre-registered video to infer the individual voxel-wise geometrical \((\Delta {x}_{t},\Delta {y}_{t}\,,\Delta {z}_{t})\) translation vector of all individual frames at time \(t\) relative to the first frame. The vertex coordinates of the tracked mesh at each individual time point \({t}\) are then given by vector summation, \(({x}_{t},{y}_{t}\,,{z}_{t})=({x}_{0},{y}_{0},{z}_{0})\,+\) \((\Delta {x}_{t}({{\bf{x}}}_{0}),\Delta {y}_{t}({{\bf{x}}}_{0}),\Delta {z}_{t}({{\bf{x}}}_{0}))\)relative to the vertex coordinates of the first frame mesh.
Curvature and intensity measurement
The continuous mean curvature at each vertex position is estimated by quadric fitting82 of the vertex coordinates in a five-vertex-ring neighbourhood (about 1 μm radius). The corresponding septin intensity was calculated by extending a trajectory to an absolute depth of 1 μm along the steepest gradient of the distance transform to the mesh surface, and assigning the 95th percentile of intensity sampled along that trajectory to the originating vertex to capture the systematically brightest accumulation of septin signal in the cortical shell.
Bleach correction
The raw septin intensity experiences decay from bleaching. We simultaneously normalized and corrected the intensity by computing the normalized septin intensity as the raw intensity divided by the mean septin intensity in the whole-cell volume at each time point. This normalized septin intensity was used for all subsequent analyses. The instantaneous change in the normalized septin intensity \((\Delta {I}_{\text{sept}})\) was computed by finite differences between consecutive time points at corresponding vertex positions.
Smoothing
The raw extracted time series are stochastic. Smoothing is required to identify temporally continuous regions of negative and positive curvature from the time series. Individual vertex time series were temporally smoothed using a moving average with the window size, \({w}_{\text{autocorr}}\), set by the lag of inflection point in the mean temporal autocorrelation curve (Extended Data Fig. 5a). For \(\Delta {I}_{\text{sept}}\), a smoothed time series was defined by linear regression of all values \({I}_{\text{sept}}(t)\) in the closed time interval \(\left[t-\frac{{w}_{\text{autocorr}}}{2},t+\frac{{w}_{\text{autocorr}}}{2}\right]\). All time series were then smoothed spatially using Laplacian smoothing with the number of smoothing iterations inferred by the lag position of the inflection point in the decay of the mean spatial autocorrelation function, which in our data was approximately 1 μm, or a five-ring neighbourhood. Finally, the vertex time series were converted to mesh face time series using barycentric interpolation.
Curvature gating
Using the mean value of individual time series, we derived unbiased thresholds for determining whether a face had a positive, flat or negative curvature using three-class Otsu thresholding.
Intensity gating
As for the curvature gating, we applied three-class Otsu thresholding to the mean intensity value of all smoothed mesh face septin time series \({I}_{\text{sept}}\), to identify depleted, background and enriched septin intensity values. For \(\Delta {I}_{\text{sept}}\), three-class Otsu thresholding was applied to the data between 0 and 10% and 90 and 100% of each time series, respectively, to identify time points of decreasing and increasing intensity. Three-class thresholding was used to identify nonsignificant, unsure and significant decreasing and increasing intensities.
Identifying contiguous surface regions (contigs) exhibiting significant intensity change
The above thresholds enable us to functionally annotate the mesh face time series at every time point and identify only the subset of faces with significantly fluctuating septin. We do this by counting the number of significant septin intensity increases and decreases and applying binary Otsu thresholding. For this face subset, we extract temporally continuous periods of positive and negative curvature using the criteria established above. These periods are referred to as contigs. Only contigs greater than wautocorr were then used to compute the normalized cross-correlation curve between the raw septin and curvature time series in the contig.
Time series analysis
The mean cross-correlation curve and 95% confidence interval for contigs of positive and negative intracellular curvature were computed to test the extent of co-fluctuation between septin and curvature on faces of negative and positive curvature (Fig. 2l and Extended Data Fig 5b). We further bipartitioned the contigs of positive and negative intracellular curvature by a score of dynamicity, which describes the total number of septin increase and decrease events divided by the duration of the contig (\({T}_{\text{contig}}\)); that is, \(\text{dynamicity}\,=\,\frac{\#\text{increase}+\#\text{decrease}}{{T}_{\text{contig}}}\). Binary Otsu thresholding was applied to distinguish contigs with low versus high dynamicity. To test how dynamicity relates to \(\Delta {I}_{\text{sept}}\) on negative and positive curvature faces, we computed the continuous relationship between the mean absolute increasing and decreasing \(\Delta {I}_{\text{Sept}}\) versus dynamicity (Extended Data Fig. 5c) in contigs using kernel density analysis. Gaussian kernel density with a bandwidth set by Scott’s rule was used to derive the joint density distribution of \(\Delta {I}_{\text{Sept}}\) and dynamicity (that is, \(p\left(X,Y\right)\), with X representing dynamicity and Y representing \(\Delta {I}_{\text{Sept}}\) over the closed intervals \(X\in \left[0,1\right]\) and \(Y\in [0,0.1]\)). The continuous relationship is then given by the marginal expectation, with capital letters denoting the random variable and \({\mathbb{E}}[\cdot ]\) representing the expectation operator, \({\mathbb{E}}[Y|X=x]=\int Y\,p(Y|X=x){\rm{d}}Y=\int Y\frac{p(X,Y)}{p(X)}{\rm{d}}Y=\frac{\int X\,p(X,Y){\rm{d}}Y}{\int p(X,Y){\rm{d}}Y}\) with standard deviation equivalently defined as the square root of the variance, \({\mathbb{E}}[{(Y-\bar{Y})}^{2}|X=x]{\mathbb{=}}{\mathbb{E}}[{Y}^{2}|X=x]{\mathbb{-}}{\mathbb{E}}{[Y|X=x]}^{2}\). The evaluation of the integrals uses 100 bins for both dynamicity and \(\Delta {I}_{\text{sept}}\).
Visualization and statistics
3D surfaces were rendered by ChimeraX83. Coloured triangle meshes representing the cell surface were imported into ChimeraX from u-shape3D as Collada dae files, as previously described22. The FIJI ImageJ package (https://imagej.net/Fiji) was used to prepare maximum-intensity projection and slice images. High-speed 3D light-sheet imaging videos were prepared using Arivis4D. Gardner–Altman paired mean difference estimation plots were generated using DABEST software suite84. All other figures were prepared using the Seaborn85 and Matplotlib86 Python visualization libraries, and the pandas87,88 Python data analysis library. Figures were assembled using the InkScape (https://inkscape.org/) vector graphics editor. Significance tests were carried out using two-sample t-tests with pooled variance, Welch’s t-tests or Mann–Whitney U-tests, depending on whether datasets had normal distributions (as measured by Shapiro–Wilk tests) and equal variance (as measured by F-tests). Statistical calculations were carried out using R for Windows, 4.0.5 (https://www.r-project.org/). Error bars in figures show 95% confidence intervals. Number of cells and/or number of different experiments analysed are given in the figure legends and in Extended Data Table 1.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The proteomic datasets generated during this study are available in the MassIVE repository at https://doi.org/10.25345/C5HQ3S35T (corresponding to Supplementary Table 1) and https://doi.org/10.25345/C5NG4GW9J (corresponding to Supplementary Table 2). All additional data that support the findings of this study are available in a Zenodo repository (https://doi.org/10.5281/zenodo.7416301). Source data are provided with this paper.
Code availability
All code, algorithms and software central to the findings of this study are available from the corresponding author upon reasonable request. Most can be found at https://github.com/DanuserLab, with code specific to this paper at https://github.com/DanuserLab/Weems-Septin-Paper. This code is also available in a Zenodo repository (https://doi.org/10.5281/zenodo.7435279).
References
Frisch, S. M. & Francis, H. Disruption of epithelial cell-matrix interactions induces apoptosis. J. Cell Biol. 124, 619–626 (1994).
Shin, S. I., Freedman, V. H., Risser, R. & Pollack, R. Tumorigenicity of virus transformed cells in nude mice is correlated specifically with anchorage independent growth in vitro. Proc. Natl Acad. Sci. USA 72, 4435–4439 (1975).
Paoli, P., Giannoni, E. & Chiarugi, P. Anoikis molecular pathways and its role in cancer progression. Biochim. Biophys. Acta Mol. Cell Res. 1833, 3481–3498 (2013).
Zollinger, H. U. Cytologic studies with the phase microscope; the formation of blisters on cells in suspension, photocytosis, with observations on the nature of the cellular membrane. Am. J. Pathol. 24, 545–567 (1948).
Erickson, C. A. & Trinkaus, J. P. Microvilli and blebs as sources of reserve surface membrane during cell spreading. Exp. Cell. Res. 99, 375–384 (1976).
Garnett, H. M. A scanning electron microscope study of the sequential changes in morphology occurring in human fibroblasts placed in suspension culture. Cytobios 27, 7–18 (1980).
Rovensky, Y. A. & Vasiliev, J. M. Surface topography of suspended tissue cells. Int. Rev. Cytol. 90, 273–307 (1984).
Charras, G. T., Coughlin, M., Mitchison, T. J. & Mahadevan, L. Life and times of a cellular bleb. Biophys. J. 94, 1836–1853 (2008).
Charras, G. & Paluch, E. Blebs lead the way: how to migrate without lamellipodia. Nat. Rev. Mol. Cell Biol. 9, 730–736 (2008).
Charras, G. T. A short history of blebbing. J. Microsc. 231, 466–478 (2008).
Norman, L. L., Bruges, J., Sengupta, K., Sens, P. & Aranda-Espinoza, H. Cell blebbing and membrane area homeostasis in spreading and retracting cells. Biophys. J. 99, 1726–1733 (2010).
Paluch, E. K. & Raz, E. The role and regulation of blebs in cell migration. Curr. Opin. Cell Biol. 25, 582–590 (2013).
Kanchanawong, P. et al. Nanoscale architecture of integrin-based cell adhesions. Nature 468, 580–584 (2010).
Frisch, S. M., Vuori, K., Ruoslahti, E. & Chan-Hui, P. Y. Control of adhesion-dependent cell survival by focal adhesion kinase. J. Cell Biol. 134, 793–799 (1996).
Hofmann, C. et al. Cell-cell contacts prevent anoikis in primary human colonic epithelial cells. Gastroenterology 132, 587–600 (2007).
Driscoll, M. K. et al. Proteolysis-free cell migration through crowded environments via mechanical worrying. Preprint at bioRxiv https://doi.org/10.1101/2020.11.09.372912 (2022).
Guzman, A., Avard, R. C., Devanny, A. J., Kweon, O. S. & Kaufman, L. J. Delineating the role of membrane blebs in a hybrid mode of cancer cell invasion in three-dimensional environments. J. Cell Sci. 133, jcs236778 (2020).
Cantelli, G. et al. TGF-β-induced transcription sustains amoeboid melanoma migration and dissemination. Curr. Biol. 25, 2899–2914 (2015).
Tozluoǧlu, M. et al. Matrix geometry determines optimal cancer cell migration strategy and modulates response to interventions. Nat. Cell Biol. 15, 751–762 (2013).
Orgaz, J. L. et al. Myosin II reactivation and cytoskeletal remodeling as a hallmark and a vulnerability in melanoma therapy resistance. Cancer Cell 37, 85–103 (2020).
Khan, Z. S., Santos, J. M., Vaz, N. G. & Hussain, F. Enhanced blebbing as a marker for metastatic prostate cancer. Biomicrofluidics 13, 034110 (2019).
Driscoll, M. K. et al. Robust and automated detection of subcellular morphological motifs in 3D microscopy images. Nat. Methods 16, 1037–1044 (2019).
Moes, M. J. A., Bijvelt, J. J. & Boonstra, J. Attachment of HeLa cells during early G1 phase. Histochem. Cell Biol. 136, 399–411 (2011).
Cannon, K. S., Woods, B. L., Crutchley, J. M. & Gladfelter, A. S. An amphipathic helix enables septins to sense micrometer-scale membrane curvature. J. Cell Biol. 218, 1128–1137 (2019).
Bridges, A. A., Jentzsch, M. S., Oakes, P. W., Occhipinti, P. & Gladfelter, A. S. Micron-scale plasma membrane curvature is recognized by the septin cytoskeleton. J. Cell Biol. 213, 23–32 (2016).
Gilden, J. K., Peck, S., Chen, Y. C. M. & Krummel, M. F. The septin cytoskeleton facilitates membrane retraction during motility and blebbing. J. Cell Biol. 196, 103–114 (2012).
Østevold, K. et al. Septin remodeling is essential for the formation of cell membrane protrusions (microtentacles) in detached tumor cells. Oncotarget 8, 76686–76698 (2017).
Oh, Y. & Bi, E. Septin structure and function in yeast and beyond. Trends Cell Biol. 21, 141–148 (2011).
Dolat, L., Hu, Q. & Spiliotis, E. T. Septin functions in organ system physiology and pathology. Biol. Chem. 395, 123–141 (2014).
Neubauer, K. & Zieger, B. The mammalian septin interactome. Front. Cell Dev. Biol. 5, 3 (2017).
Amir, S., Wang, R., Simons, J. W. & Mabjeesh, N. J. SEPT9_v1 up-regulates hypoxia-inducible factor 1 by preventing its RACK1-mediated degradation. J. Biol. Chem. 284, 11142–11151 (2009).
Diesenberg, K., Beerbaum, M., Fink, U., Schmieder, P. & Krauss, M. SEPT9 negatively regulates ubiquitin-dependent downregulation of EGFR. J. Cell Sci. 128, 397–407 (2015).
Mostowy, S. et al. Septin 11 restricts InlB-mediated invasion by Listeria. J. Biol. Chem. 284, 11613–11621 (2009).
Gonzalez, M. E., Makarova, O., Peterson, E. A., Privette, L. M. & Petty, E. M. Up-regulation of SEPT9_v1 stabilizes c-Jun-N-terminal kinase and contributes to its pro-proliferative activity in mammary epithelial cells. Cell. Signal. 21, 477–487 (2009).
Marcus, E. A. et al. Septin oligomerization regulates persistent expression of ErbB2/HER2 in gastric cancer cells. Biochem. J 473, 1703–1718 (2016).
Sirianni, A. et al. Mitochondria mediate septin cage assembly to promote autophagy of Shigella. EMBO Rep. 17, 1029–1043 (2016).
Beber, A. et al. Membrane reshaping by micrometric curvature sensitive septin filaments. Nat. Commun. 10, 420 (2019).
Lobato-Márquez, D. et al. Mechanistic insight into bacterial entrapment by septin cage reconstitution. Nat. Commun. 12, 4511 (2021).
Bridges, A. A. et al. Septin assemblies form by diffusion-driven annealing on membranes. Proc. Natl Acad. Sci. USA 111, 2146–2151 (2014).
Sirajuddin, M., Farkasovsky, M., Zent, E. & Wittinghofer, A. GTP-induced conformational changes in septins and implications for function. Proc. Natl Acad. Sci. USA 106, 16592–16597 (2009).
Silva Martins, C. et al. Human septins organize as octamer-based filaments and mediate actin-membrane anchoring in cells. J. Cell Biol. 3, 222, e202203016 (2022).
Hu, Q., Nelson, W. J. & Spiliotis, E. T. Forchlorfenuron alters mammalian septin assembly, organization, and dynamics. J. Biol. Chem. 283, 29563–29571 (2008).
Mishra, S., Ande, S. R. & Nyomba, B. L. G. The role of prohibitin in cell signaling. FEBS J. 277, 3937–3946 (2010).
Freeman, A. K. & Morrison, D. K. 14-3-3 proteins: diverse functions in cell proliferation and cancer progression. Semin. Cell Dev. Biol. 22, 681–687 (2011).
Farin, K. et al. Oncogenic synergism between ErbB1, nucleolin, and mutant ras. Cancer Res. 71, 2140–2151 (2011).
Meurette, O. et al. Notch activation induces Akt signaling via an autocrine loop to prevent apoptosis in breast epithelial cells. Cancer Res. 69, 5015–5022 (2009).
Teramoto, H. et al. Autocrine activation of an osteopontin-CD44-Rac pathway enhances invasion and transformation by H-RasV12. Oncogene 24, 489–501 (2005).
De Belly, H. et al. Membrane tension gates ERK-mediated regulation of pluripotent cell fate. Cell Stem Cell 28, 273–284 (2020).
Bergert, M. et al. Cell surface mechanics gate embryonic stem cell differentiation. Cell Stem Cell 28, 209–216 (2020).
Otto, A., Collins-Hooper, H., Patel, A., Dash, P. R. & Patel, K. Adult skeletal muscle stem cell migration is mediated by a blebbing/amoeboid mechanism. Rejuvenation Res. 14, 249–260 (2011).
Sedzinski, J. et al. Polar actomyosin contractility destabilizes the position of the cytokinetic furrow. Nature 476, 462–468 (2011).
Zhou, Y. & Hancock, J. F. Ras nanoclusters: versatile lipid-based signaling platforms. Biochim. Biophys. Acta Mol. Cell Res. 1853, 841–849 (2015).
Muñoz-Couselo, E., Adelantado, E. Z., Ortiz, C., García, J. S. & Perez-Garcia, J. NRAS-mutant melanoma: current challenges and future prospect. OncoTargets Ther. 10, 3941–3947 (2017).
Blum, W. et al. The phytohormone forchlorfenuron decreases viability and proliferation of malignant mesothelioma cells in vitro and in vivo. Oncotarget 10, 6944–6956 (2019).
Pesticides - Fact Sheet for Forchlorfenuron (US EPA, 2004).
Lito, P. et al. Relief of profound feedback inhibition of mitogenic signaling by RAF inhibitors attenuates their activity in BRAFV600E melanomas. Cancer Cell 22, 668–682 (2012).
Mohan, A. S. et al. Enhanced dendritic actin network formation in extended lamellipodia drives proliferation in growth-challenged Rac1P29S melanoma cells. Dev. Cell 49, 444–460 (2019).
Park, J. S. et al. Mechanical regulation of glycolysis via cytoskeleton architecture. Nature 578, 621–626 (2020).
Shurer, C. R. et al. Physical principles of membrane shape regulation by the glycocalyx. Cell 177, 1757–1770 (2019).
van Muijen, G. N. P. et al. Establishment and characterization of a human melanoma cell line (MV3) which is highly metastatic in nude mice. Int. J. Cancer 48, 85–91 (2007).
Giard, D. J. et al. In vitro cultivation of human tumors: establishment of cell lines derived from a series of solid tumors. JNCI J. Natl Cancer Inst. 51, 1417–1423 (1973).
Quintana, E. et al. Human melanoma metastasis in NSG mice correlates with clinical outcome in patients. Sci. Transl. Med. 4, 159ra149 (2012).
Zaritsky, A. et al. Interpretable deep learning uncovers cellular properties in label-free live cell images that are predictive of highly metastatic melanoma. Cell Syst. 12, 733–747 (2021).
Huang, J. 3D cell culture on VitroGel system. J. Cytol. Tissue Biol. https://doi.org/10.24966/CTB-9107/S1001 (2019).
Damke, H., Baba, T., Warnock, D. E. & Schmid, S. L. Induction of mutant dynamin specifically blocks endocytic coated vesicle formation. J. Cell Biol. 127, 915–934 (1994).
Altschuler, Y. et al. Redundant and distinct functions for dynamin-1 and dynamin-2 isoforms. J. Cell Biol. 143, 1871–1881 (1998).
Hardy, S., Kitamura, M., Harris-Stansil, T., Dai, Y. & Phipps, M. L. Construction of adenovirus vectors through Cre-lox recombination. J. Virol. 71, 1842–1849 (1997).
Song, B. D., Yarar, D. & Schmid, S. L. An assembly-incompetent mutant establishes a requirement for dynamin self-assembly in clathrin-mediated endocytosis in vivo. Mol. Biol. Cell 15, 2243–2252 (2004).
Kim, D. I. et al. An improved smaller biotin ligase for BioID proximity labeling. Mol. Biol. Cell 27, 1188–1196 (2016).
de la Cova, C., Townley, R., Regot, S. & Greenwald, I. A real-time biosensor for ERK activity reveals signaling dynamics during C. elegans cell fate specification. Dev. Cell 42, 542–553 (2017).
Nassar, N., Singh, K. & Garcia-Diaz, M. Structure of the dominant negative S17N mutant of Ras. Biochemistry 49, 1970–1974 (2010).
Dean, K. M., Roudot, P., Welf, E. S., Danuser, G. & Fiolka, R. Deconvolution-free subcellular imaging with axially swept light sheet microscopy. Biophys. J. 108, 2807–2815 (2015).
Dean, K. M. et al. Diagonally scanned light-sheet microscopy for fast volumetric imaging of adherent cells. Biophys. J. 110, 1456–1465 (2016).
Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. SMC-9, 62–66 (1979).
Elliott, H. et al. Myosin II controls cellular branching morphogenesis and migration in three dimensions by minimizing cell-surface curvature. Nat. Cell Biol. 17, 137–147 (2015).
Costes, S. V. et al. Automatic and quantitative measurement of protein-protein colocalization in live cells. Biophys. J. 86, 3993–4003 (2004).
Cordelieres, F. P. & Bolte, S. Experimenters’ guide to colocalization studies: finding a way through indicators and quantifiers, in practice. Methods Cell. Biol. 123, 978 (2014).
Jacobson, A. gptoolbox: Geometry processing toolbox (2021).
Monge, G. Mémoire sur la théorie des déblais et des remblais. Mem. Math. Phys. Acad. Royale Sci. 666–704 (1781).
Solomon, J., Rustamov, R., Guibas, L. & Butscher, A. Earth mover’s distances on discrete surfaces. ACM Trans. Graphics 33, 1–12 (2014).
Vercauteren, T., Pennec, X., Perchant, A. & Ayache, N. Non-parametric diffeomorphic image registration with the demons algorithm. In Proc. 10th International Conference on Medical Image Computing and Computer-Assisted Intervention (eds Ayache, N., Ourselin, S. & Maeder, A.) 319–326 (Springer, 2007).
Panozzo, D., Puppo, E. & Rocca, L. Efficient multi-scale curvature and crease estimation. In Proc. 2nd International Workshop on Computer Graphics, Computer Vision and Mathematics 9–16 (GraVisMa, 2010).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. https://doi.org/10.1002/pro.3235 (2018).
Ho, J., Tumkaya, T., Aryal, S., Choi, H. & Claridge-Chang, A. Moving beyond P values: data analysis with estimation graphics. Nat. Methods 16, 565–566 (2019).
Waskom, M. seaborn: statistical data visualization. J. Open Source Softw. 6, 3021 (2021).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Reback, J. et al. pandas-dev/pandas: Pandas 1.2.4. https://doi.org/10.5281/ZENODO.4681666 (2021).
McKinney, W. Data structures for statistical computing in Python. In Proc. 9th Python in Science Conference (eds van der Walt, S. & Millman, J.) 56–61 (SciPy, 2010).
Acknowledgements
We thank J. Huang of TheWell Bioscience for providing information about the physical properties of VitroGel, D. Segal for discussions about septin inhibition, X. Jiang for conversations about optimal transport, P. Roudot for conversations on 3D image analysis, T. Isogai for advice on proximity proteomics, B. Nanes for assistance in data blinding and M. McMurray for feedback on the manuscript. We thank the UT Southwestern BioHPC team for providing necessary computing infrastructure. Funding for this work in the Danuser lab has been provided through grants R35 GM136428 (NIH) and I-1840-20200401 (Welch Foundation), and in the Fiolka lab through R33 CA235254 (NIH) and R35 GM133522 (NIH). M.D. was supported by the fellowship K99 GM123221 (NIH). A.D.W.s is a fellow of the Jane Coffin Childs Memorial Fund.
Author information
Authors and Affiliations
Contributions
Study conceived by A.D.W. and developed by A.D.W., E.S.W., M.K.D. and G.D. Experimental design by A.D.W., E.S.W., M.K.D., G.M.G., V.S.M. and G.D. Experiments carried out by A.D.W., E.S.W., B.G.W., G.M.G., V.S.M. and D.R. Data analysed by A.D.W., M.K.D., H.M.-F. and F.Z. Cloning and cell line creation carried out by J.C. Statistical analysis carried out by A.D.W., M.K.D. and F.Z. Data visualization and figure preparation carried out by A.D.W. Figure layouts designed by A.D.W. and G.D. Software designed by M.K.D. (u-shape3D), H.M.-F. (EMD analysis) and F.Z. (time series analysis). B.-J.C., K.M.D. and R.F. designed, built and maintained 3D light-sheet microscopes. A.D.W. and G.D. wrote the paper, which was edited and approved by all authors. Experiments in Fig. 1a carried out by A.D.W. and E.S.W. and analysed by A.D.W. All other experiments in Fig. 1 carried out and analysed by A.D.W. Experiments in Fig. 2d carried out by E.S.W. and analysed by M.K.D. Experiment in Fig. 2j carried out by A.D.W. and E.S.W. Analysis in Fig. 2l carried out by F.Z. All other experiments and analysis in Fig. 2 carried out by A.D.W. Experiments in Fig. 3b carried out by A.D.W. and E.S.W. and analysed by A.D.W. All other experiments in Fig. 3 carried out by A.D.W. Analysis in Fig. 4b carried out by H.M.-F. All other experiments and analysis in Fig. 4 carried out by A.D.W. All experiments and analysis in Fig. 5 carried out by A.D.W. Experiments in Extended Data Fig. 1A carried out by E.S.W. and analysed by M.K.D. Experiments in Extended Data Fig. 1B–D carried out by A.D.W. and E.S.W. All other analysis in Extended Data Fig. 1 carried out by A.D.W. Experiments and analysis in Extended Data Fig. 2 carried out by A.D.W. Experiments in Extended Data Fig. 3 carried out by V.S.M., D.R. and A.D.W. and analysed by A.D.W. Experiments and analysis in Extended Data Fig. 4 carried out by A.D.W. Experiments in Extended Data Fig. 5 carried out by A.D.W. and analysed by F.Z. Experiments and analysis in Extended Data Fig. 6 carried out by A.D.W. Experiments in Extended Data Fig. 7 carried out by G.M.G. and analysed by G.M.G. and A.D.W. Experiments and analysis in Extended Data Figs. 8–10 carried out by A.D.W. Experiments in Extended Data Table 2 carried out by B.G.W and analysed by A.D.W. Experiments and analysis in Supplementary Videos 1–4 carried out by A.D.W.
Corresponding authors
Ethics declarations
Competing interests
The Board of Regents of the University of Texas System has filed for a patent (US patent 63/334,029, filed 22 April 2022) covering therapeutic targeting of septins and blebbing, which was based in part on data from this article. A.D.W., G.D., E.S.W. and M.K.D. are named as inventors. The authors declare no other competing interests.
Peer review
Peer review information
Nature thanks Philippe Bastiaens, Erez Raz, Elias Spiliotis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Replicates of bleb inhibition viability experiments.
(A) Fraction of cell surface comprised of blebs for MV3 cells treated with different concentrations of WGA. Dashed lines separate quartiles. Dots represent individual cells. (B) Individual bleb inhibition viability experiments, as summarized in Fig. 1a. Each pairing of adhered/detached cell experiments was performed on the same day and seeded with cells from the same diluted cell suspensions. (C) Morphology of cells in “adhered” conditions, related to Fig. 1a. Cells grown without perturbation for 24 h on adherent chamber slides. Representative of either 688 (MV3), 405 (M498), or 576 (A375) cells observed. (D) Paired mean differences between control and experimental condition shown in Gardner-Altman estimation plots for the data presented in Figs. 1d, 3b, 4d, 4e, and 4f. Gray distributions represent 5000 bootstrapped samples, black bars represent 95% confidence intervals, black dots represent mean difference.
Extended Data Fig. 2 SEPT6-GFP localization and the effects of septin inhibition.
(A) Mouse SEPT6-GFP probe localization in MV3 cells embedded in soft bovine collagen. Maximum intensity projections (left) and single z-slices of 0.16 micron thickness (right) shown for representative cells in two columns. (B) Maximum intensity projections of mouse SEPT6-GFP probe localization in MV3 cells with and without inhibition by FCF or SEPT2(33–306) expression. Cells embedded in soft bovine collagen. (Below) Fraction of cell surface comprised of blebs for MV3 cells with and without SEPT6-GFP expression, FCF treatment, and SEPT2(33–306) expression. Dashed lines separate quartiles. Dots represent individual cells. (C) Mouse SEPT6-GFP probe localization in MV3 cells embedded in soft bovine collagen. Maximum intensity projections and single z-slices of 0.16 micron thickness shown for representative cells in two columns. (Above) Cells expressing the SEPT2(33–306) mutant. (Below) Cells expressing cytosolic mCherry (not shown) using the same pLVX-IRES-Hyg vector as the SEPT2(33–306) construct. Included as a negative control demonstrating that the septin mislocalization in SEPT2(33–306)-expressing cells is not due to effects arising from the protein expression construct.
Extended Data Fig. 3 Endogenous septin localization via immunofluorescence.
(A) MV3 cells adhered to fibronectin-coated glass slides showing either the mouse SEPT6-GFP probe or anti-SEPT2 immunofluorescence localization. While the well-established septin localization to actin stress fibers (https://doi.org/10.1016/S1534-5807(02)00366-0) is apparent in both samples, the anti-SEPT2 signal is irregular, punctate, and possesses low signal-to-noise ratio (SNR) compared to SEPT6-GFP. Scale bars are 10 µm. (B) Anti-SEPT2 signal in a rounded blebbing MV3 cell embedded in soft bovine collagen, with maximum intensity projections (top) and single z-slices of 0.16 micron thickness (below). Phalloidin signal from the same cell shown on right to visualize the cell surface. Anti-SEPT2 signal possesses low SNR, as seen in adhered cells. Despite this, the signal is enriched at the cell surface in blebby regions of the cell just as seen in cells expressing the SEPT6-GFP probe (see Extended Data Fig. 2a). (C) Cell surface distribution of anti-SEPT2 signal for the cell shown in Extended Data Fig. 3b. (D) Local anti-SEPT2 or SEPT6-GFP intensity and intracellular mean curvature as a function of distance from bleb edges. Mean intensity is comprised only of the brightest 50% of the cell surface to account for punctate and discontinuous immunofluorescent signal. (E) Fraction of cortical voxels (within 0.96 µm of surface) with anti-SEPT2 intensity higher than cytoplasmic mean intensity in MV3 cells with and without expression of SEPT6-GFP. Dashed lines separate quartiles and dots represent individual cells.
Extended Data Fig. 4 A mutation that disrupts curvature sensing blocks septin localization to the cell surface.
Top, cartoon illustrating the mechanism underlying the altered function of the SEPT6(ΔAH)-GFP mutant. Bottom left, mean local SEPT6(ΔAH)-GFP intensity and intracellular mean curvature as a function of distance from bleb edges, calculated from 630 blebs across 5 MV3 cells. Error bands indicate 95% confidence intervals. Bottom right, difference in mean local septin intensity compared to local maxima as a function of distance from bleb edge for both SEPT6-GFP (as seen in Fig. 2e) and SEPT6(ΔAH)-GFP.
Extended Data Fig. 5 Analysis of high-speed timelapse SEPT6-GFP data in MV3.
(A) Autocorrelation curves for the local mean curvature and SEPT6-GFP data presented in Fig. 2l; the fluorescence signal was normalized to account for photobleaching as described in Methods. (B) Temporal cross-correlation functions as shown in Fig. 2l, here with extended range of time lags. (C) Mean rates of absolute change in SEPT6-GFP intensity for positive and negative intracellular mean curvature contigs, expressed as a function of the dynamicity of the contigs they occur within. Dynamicity is defined as the total number of timepoints showing intensity increases and decreases that occur within a contig, divided by the total number of timepoints within that contig. Derived from the same dataset used in Fig. 2l. Vertical dashed line represents the threshold for low and high dynamic groups as shown in Figs. 2l and S7B.
Extended Data Fig. 6 A mutation that disrupts inter-oligomeric polymerization blocks septin localization to the cell surface.
Top, a cartoon illustrating the mechanism underlying the altered function of the SEPT2(33–306) mutant. Bottom left above, mean local SEPT6-GFP intensity and intracellular mean curvature as a function of distance from bleb edges in cells expressing SEPT2(33–306), calculated from 1293 blebs across 8 MV3 cells. Error bands indicate 95% confidence intervals. Bottom left below, difference in mean local SEPT6-GFP intensity compared to local maxima as a function of distance from bleb edge for cells expressing SEPT2(33–306) and control cells (as seen in Fig. 2e). Bottom right, Western blot of SEPT2 in MV3 cells with and without SEPT2(33–306) expression. Calculated mass of SEPT2 is 41.5kD and SEPT2(33–306) is 31.7 kD. Annotations show positions of the indicated ladder bands.
Extended Data Fig. 7 Changes in septin expression upon MAPKi treatment of A375 cells.
Top, Western blots of SEPT2, SEPT6, SEPT7, and SEPT9 abundance (the 4 septins with highest expression in A375 cells, as measured by mass spectrometry, see Extended Data Table 2) upon combined treatment with Dabrafenib and Trametinib at doses indicated. Three independent experiments were performed, with data from all replicates shown alongside vinculin loading controls. Bottom, densitometry analysis of Western blots showing fold change in septin expression as dose response curves. Dashed lines indicate individual experiments (color-coded according to the color bars indicating Western blot replicates); solid red lines indicate means of all replicates.
Extended Data Fig. 8 Viability effects of SEPT2(33–306) expression.
Viability in adhered and detached melanoma cells expressing SEPT2(33–306). Cells grown for 24 h and assayed for cell death using ethidium homodimer staining. All treatment groups grown and assayed in simultaneous paired experiments. Dots represent individual experiments. Cell counts for all replicates (see Extended Data Table 1 for individual counts): MV3 (951, 872), A375 (653, 1302). MV3 and A375 datasets were tested using one-sided Paired T Tests, p = 1.02x10−3 and p = 0.388.
Extended Data Fig. 9 Effect of SEPT6 expression on NRAS Localization.
(A) Observed NRAS-GFP % enrichment at the cortex (voxels within 0.96 µm of surface) of individual unperturbed and septin-inhibited MV3 cells. Basal/FCF tested with one-sided Welch’s T-test (p = 4.27x10−5), normality tested with Shapiro-Wilk (p = 0.219 & 0.219), variance tested with two-tailed F test (p = 0.0054). Basal/SEPT2(33–306) tested with two sample one-sided t-test using pooled variance (p = 0.0101), normality tested with Shapiro-Wilk (p = 0.219 & 0.724), variance tested with two-tailed F test (p = 0.768). Dashed lines separate quartiles. (B) Observed NRAS-GFP percent enrichment at the cortex (voxels within 0.96 µm of surface) of individual MV3 cells with and without expression of SEPT6-HALO using the same pLVX-IRES-Hyg vector as the SEPT2(33–306) construct. Included as a negative control demonstrating that the NRAS mislocalization in SEPT2(33–306)-expressing cells is not due to effects arising from the protein expression construct. Significance tested with two sample one-sided T-test with pooled variance (p = 0.738), normality tested with Shapiro-Wilk (p = 0.315 & 0.426), variance tested with two-tailed F test (p = 0.181). Dashed lines separate quartiles. (C) Cell death upon expression of dominant negative NRAS(S17N) for adhered and detached MV3 cells. Cells grown as in Fig. 1a for 24 h. Data were normalized by subtracting paired negative control values from each treatment group. Dots represent individual experiments. Cell counts for all replicates, with control counts in parentheses (see Extended Data Table 1 for individual counts): Adh. 1105(1039), Det. 799(734).
Extended Data Fig. 10 Anoikis resistance in non-malignant cells.
(A) SEPT6-GFP localization in a recently detached HEK cell, embedded in soft bovine collagen. Top, maximum intensity projection (MIP); bottom, individual 0.16 µm z-slice from the same cell. (B) Additional fraction of MV3 cells showing caspase activation after 4 h of treatment with 10 µg/mL WGA (compared to matched, same-day negative controls) for adhered and detached cells. Total cell counts with control in parenthesis: Replicate 1: Adh 143(220), Det 310(544); Replicate 2: Adh 182(152), Det 256(215); Replicate 3: Adh 134(157), Det 89(267). (C) The same data shown in Fig. 5c, without normalization. Solid, dashed, and dotted lines represent paired, same-day experiments. Total cell counts for each condition, with adhered group in parenthesis: Replicate 1: Con 49(89), DYN2(K44A) 38(55), DYN2(K44A)+WGA 32(25), DYN2(K44A)+FCF 27(33); Replicate 2: Con 23(25), DYN2(K44A) 28(39), DYN2(K44A)+WGA 27(43), DYN2(K44A)+FCF 28(34); Replicate 3: Con 92(54), DYN2(K44A) 121(84), DYN2(K44A)+WGA 101(115), DYN2(K44A)+FCF 141(70).
Extended Data Fig. 11 Graphical Abstract.
Summary illustration of the information flow via bleb signaling as well as the mechanisms through which pro-survival bleb signaling hubs are constructed.
Supplementary information
Supplementary Fig. 1
Uncropped images of western blots shown in this study.
Supplementary Table 1
Summary of mass spectrometry results from MV3 lysate and SEPT6 pulldown experiments. Proteins that were identified with only 1 PSM are highlighted in orange, proteins that were identified but not quantified are highlighted in blue, and potential contaminants from experimenter are highlighted in yellow. Details of column titles are as follows—Protein FDR confidence: High (1% false discovery rate), Medium (5% false discovery rate) or Low (>5% false discovery rate); Master: if more than one protein in a group has the same score, and an equal number of PSMs, and an equal number of peptides, the protein with the longest sequence is designated as the master protein; Accession: protein accession number (from UniProtKB); Description: description taken from UniProt; Coverage (%): percentage of the protein sequence that was covered by the peptides identified for the protein; #PSMs: number of peptide spectrum matches, or the number of spectra assigned to peptides that contributed to the inference of the protein; MW [kDa]: molecular weight of the protein based on the sequence from Uniprot; Abundance: the sum of the peak intensities for each peptide identified for that protein (these values can be used to compare relative abundance of a protein between samples).
Supplementary Table 2
Summary of mass spectrometry results from SEPT6 BioID proximity labelling experiment. Proteins that were identified with only 1 PSM are highlighted in orange, proteins that were identified but not quantified are highlighted in blue, and potential contaminants from experimenter are highlighted in yellow. Details of column titles are as follows—Protein FDR confidence: High (1% false discovery rate), Medium (5% false discovery rate) or low (>5% false discovery rate); Master: if more than one protein in a group has the same score, and an equal number of PSMs, and an equal number of peptides, the protein with the longest sequence is designated as the master protein; Accession: protein accession number (from UniProtKB); Description: description taken from UniProt; Coverage (%): percentage of the protein sequence that was covered by the peptides identified for the protein; #PSMs: number of peptide spectrum matches, or the number of spectra assigned to peptides that contributed to the inference of the protein; MW [kDa]: molecular weight of the protein based on the sequence from Uniprot; Abundance: the sum of the peak intensities for each peptide identified for that protein (these values can be used to compare relative abundance of a protein between samples).
Supplementary Video 1
Mammalian cells become rounded and blebby after detachment. Top: three common cell lines (MEF cells, human epithelial kidney 293 cells and human bronchial epithelial cells) grown under standard adherent conditions in plastic Petri dishes. Bottom: Cells from the same cultures 3 min after treatment with trypsin and gentle pipetting to cause detachment. All time lapses played at ≈15× speed.
Supplementary Video 2
Conversion of raw light-sheet microscope data to 3D surfaces using u-Shape3D. Poorly adhered MV3 cell embedded in soft bovine collagen. Septins visualized with mouse SEPT6–GFP probe.
Supplementary Video 3
Septins ‘pulse’ at the curvy bases of blebs. Poorly adhered MV3 cell embedded in soft bovine collagen. Septins visualized with mouse SEPT6–GFP probe. Five-min time lapse acquired at ≈0.83 Hz played twice, the second time with arrowheads and insets highlighting septin pulses.
Supplementary Video 4
Iterative blebbing produces stable septin structures. Poorly adhered MV3 cell embedded in soft bovine collagen. Septins visualized with mouse SEPT6–GFP probe. Five-min time lapse acquired at ≈0.83 Hz. Three orthogonal views show tightly spaced iterative blebbing producing septin pulses that combine to form a bright stable structure that maintains signal intensity independent of local blebbing.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Weems, A.D., Welf, E.S., Driscoll, M.K. et al. Blebs promote cell survival by assembling oncogenic signalling hubs. Nature 615, 517–525 (2023). https://doi.org/10.1038/s41586-023-05758-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-05758-6
This article is cited by
-
Pan-cancer evaluation of regulated cell death to predict overall survival and immune checkpoint inhibitor response
npj Precision Oncology (2024)
-
Blebs signal survival
Nature Reviews Cancer (2023)
-
Anoikis resistance through blebs
Nature Reviews Molecular Cell Biology (2023)
-
Bleb protrusions help cancer cells to cheat death
Nature (2023)
-
Cellular harmonics for the morphology-invariant analysis of molecular organization at the cell surface
Nature Computational Science (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.