Abstract
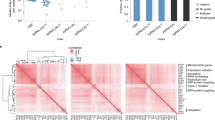
Type VI CRISPR enzymes are RNA-targeting proteins with nuclease activity that enable specific and robust target gene knockdown without altering the genome. To define rules for the design of Cas13d guide RNAs (gRNAs), we conducted massively parallel screens targeting messenger RNAs (mRNAs) of a green fluorescent protein transgene, and CD46, CD55 and CD71 cell-surface proteins in human cells. In total, we measured the activity of 24,460 gRNAs with and without mismatches relative to the target sequences. Knockdown efficacy is driven by gRNA-specific features and target site context. Single mismatches generally reduce knockdown to a modest degree, but spacer nucleotides 15–21 are largely intolerant of target site mismatches. We developed a computational model to identify optimal gRNAs and confirm their generalizability, testing 3,979 guides targeting mRNAs of 48 endogenous genes. We show that Cas13 can be used in forward transcriptomic pooled screens and, using our model, predict optimized Cas13 gRNAs for all protein-coding transcripts in the human genome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Screen data have been deposited at the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) with the accession no. GSE142675. All code and software to reproduce our entire analyses are available on our gitlab repository (https://gitlab.com/sanjanalab/cas13). Moreover, we provide precomputed gRNA predictions targeting all protein-coding transcripts in the human genome on our web-based repository (https://cas13design.nygenome.org). Other data and materials that support the findings of this research are available from the corresponding author upon reasonable request.
Code availability
The predictive on-target model as well as all code for the analyses presented in the letter is available on our gitlab repository (https://gitlab.com/sanjanalab/cas13).
References
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
Abudayyeh, O. O. et al. RNA targeting with CRISPR–Cas13. Nature 550, 280–284 (2017).
East-Seletsky, A. et al. Two distinct RNase activities of CRISPR-C2c2 enable guide-RNA processing and RNA detection. Nature 538, 270–273 (2016).
Smargon, A. A. et al. Cas13b Is a type VI-B CRISPR-associated RNA-guided RNase differentially regulated by accessory proteins Csx27 and Csx28. Mol. Cell 65, 618–630 (2017).
Konermann, S. et al. Transcriptome engineering with RNA-targeting type VI-D CRISPR effectors. Cell 173, 665–676 (2018).
Yan, W. X. et al. Cas13d Is a compact RNA-targeting type VI CRISPR effector positively modulated by a WYL-domain-containing accessory protein. Mol. Cell 70, 327–339 (2018).
Freije, C. A. et al. Programmable inhibition and detection of RNA viruses using Cas13. Mol. Cell 76, 1–12 (2019).
Poosala, P., Lindley, S. R., Anderson, K. M. & Anderson, D. M. Targeting toxic nuclear RNA foci with CRISPR-Cas13 to treat myotonic dystrophy. Preprint at bioRxiv https://doi.org/10.1101/716514 (2019).
Mahas, A., Aman, R. & Mahfouz, M. CRISPR-Cas13d mediates robust RNA virus interference in plants. Genome Biol. 20, 1–16 (2019).
Kushawah, G. et al. CRISPR-Cas13d induces efficient mRNA knock-down in animal embryos. Preprint at bioRxiv https://doi.org/10.1101/2020.01.13.904763 (2020).
Gootenberg, J. S. et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 356, 438–442 (2017).
Gootenberg, J. S. et al. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science 360, 439–444 (2018).
Cox, D. B. T. et al. RNA editing with CRISPR-Cas13. Science 358, 1019–1027 (2017).
Li, J.et al. Targeted mRNA demethylation using an engineered dCas13b-ALKBH5 fusion protein. Preprint at bioRxiv https://doi.org/10.1101/614859 (2019).
Wang, H. et al. CRISPR-mediated live imaging of genome editing and transcription. Science 365, 2–6 (2019).
Yang, L.-Z. et al. Dynamic imaging of RNA in living cells by CRISPR-technology. Mol. Cell 76, 1–17 (2019).
Jillette, N. & Cheng, A. W. CRISPR artificial splicing factors. Preprint at bioRxiv https://doi.org/10.1101/431064 (2018).
Anderson, K. M., Poosala, P., Lindley, S. R. & Anderson, D. M. Targeted cleavage and polyadenylation of RNA by CRISPR-Cas13. Preprint at bioRxiv https://doi.org/10.1101/531111 (2019).
Meeske, A. J., Nakandakari-Higa, S. & Marraffini, L. A. Cas13-induced cellular dormancy prevents the rise of CRISPR-resistant bacteriophage. Nature 570, 241–245 (2019).
Meeske, A. J. & Marraffini, L. A. RNA guide complementarity prevents self-targeting in type VI CRISPR systems. Mol. Cell 71, 791–801 (2018).
Liu, L. et al. The molecular architecture for RNA-guided RNA cleavage by Cas13a. Cell 170, 714–720 (2017).
Tambe, A., East-seletsky, A., Knott, G. J., Connell, M. R. O. & Doudna, J. A. RNA binding and HEPN-nuclease activation are decoupled in CRISPR-Cas13a. Cell Rep. 24, 1025–1036 (2018).
Zhang, C. et al. Structural basis for the RNA-guided ribonuclease activity of CRISPR-Cas13d. Cell 175, 212–223 (2018).
Zhang, B. et al. Two HEPN domains dictate CRISPR RNA maturation and target cleavage in Cas13d. Nat. Commun. 10, 2544 (2019).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR–Cas9 complex. Nature 517, 583–588 (2015).
Replogle, J. M.et al. Direct capture of CRISPR guides enables scalable, multiplexed, and multi-omic Perturb-seq. Preprint at bioRxiv https://doi.org/10.1101/503367 (2018).
Doench, J. G. et al. Rational design of highly active sgRNAs for CRISPR–Cas9-mediated gene inactivation. Nat. Biotechnol. 32, 1262–1267 (2014).
McFarland, J. M. et al. Improved estimation of cancer dependencies from large-scale RNAi screens using model-based normalization and data integration. Nat. Commun. 9, 1–13 (2018).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Briese, M. et al. A systems view of spliceosomal assembly and branchpoints with iCLIP. Nat. Struct. Mol. Biol. 26, 930–940 (2019).
Saulière, J. et al. CLIP-seq of eIF4AIII reveals transcriptome-wide mapping of the human exon junction complex. Nat. Struct. Mol. Biol. 19, 1124–1131 (2012).
Hauer, C. et al. Exon junction complexes show a distributional bias toward alternatively spliced mRNAs and against mRNAs coding for ribosomal proteins. Cell Rep. 16, 1588–1603 (2016).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Lorenz, R. et al. ViennaRNA package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–88 (2014).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, 212 (2009).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Kolde, R., Laur, S., Adler, P. & Vilo, J. Robust rank aggregation for gene list integration and meta-analysis. Bioinformatics 28, 573–580 (2012).
Agarwal, V., Subtelny, A. O., Thiru, P., Ulitsky, I. & Bartel, D. P. Predicting microRNA targeting efficacy in Drosophila. Genome Biol. 19, 1–23 (2018).
Krueger, J. & Rehmsmeier, M. RNAhybrid: microRNA target prediction easy, fast and flexible. Nucleic Acids Res. 34, 451–454 (2006).
Acknowledgements
We thank the entire Sanjana laboratory for support and advice. We thank R. Satija for critical feedback. We also thank D. Knowles for discussion about model building and M. Zaran for assistance with the web-tool server. N.E.S. is supported by New York University and New York Genome Center startup funds, National Institutes of Health (NIH)/National Human Genome Research Institute (grant nos. R00HG008171, DP2HG010099), NIH/National Cancer Institute (grant no. R01CA218668), Defense Advanced Research Projects Agency (grant no. D18AP00053), the Sidney Kimmel Foundation, the Melanoma Research Alliance, and the Brain and Behavior Foundation. A.M.-M. is supported by a CONACyT-Mexico Fellowship (no. 412653).
Author information
Authors and Affiliations
Contributions
H.H.W and N.E.S conceived the project. H.H.W., N.E.S and A.M.-M. designed the experiments. A.M.-M. and H.H.W. performed and analyzed the experiments. H.H.W. analyzed the screen data, and built the gRNA prediction software and online repository. X.G., M.L. and Z.D. helped with post-screen validation experiments. N.E.S. supervised the work. H.H.W. and N.E.S. wrote the manuscript with input from all the authors.
Corresponding author
Ethics declarations
Competing interests
The New York Genome Center and New York University have applied for patents relating to the work in this article. N.E.S. is an adviser to Vertex.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Materials
Supplementary Figs. 1–11, Tables 1–7 and Notes 1 and 2.
Supplementary Data 1
Guide RNA enrichments of sorted populations over input populations.
Supplementary Data 2
Combined on-target model input including all features.
Supplementary Data 3
Guide RNA predictions for protein-coding transcripts in GENCODE.
Supplementary Data 4
Oligonucleotide information.
Supplementary Data 5
Statistics of processed gRNA and cell numbers.
Supplementary Data 6
Sequencing read processing statistics.
Supplementary Data 7
Raw gRNA counts.
Supplementary Data 8
Final gRNA counts (after normalization; batch correction; outlier removal).
Rights and permissions
About this article
Cite this article
Wessels, HH., Méndez-Mancilla, A., Guo, X. et al. Massively parallel Cas13 screens reveal principles for guide RNA design. Nat Biotechnol 38, 722–727 (2020). https://doi.org/10.1038/s41587-020-0456-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0456-9
This article is cited by
-
Massively parallel profiling of RNA-targeting CRISPR-Cas13d
Nature Communications (2024)
-
Prediction of on-target and off-target activity of CRISPR–Cas13d guide RNAs using deep learning
Nature Biotechnology (2024)
-
Transcript-specific induction of stop codon readthrough using a CRISPR-dCas13 system
EMBO Reports (2024)
-
Engineered, nucleocytoplasmic shuttling Cas13d enables highly efficient cytosolic RNA targeting
Cell Discovery (2024)
-
Structures, mechanisms and applications of RNA-centric CRISPR–Cas13
Nature Chemical Biology (2024)