Abstract
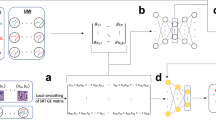
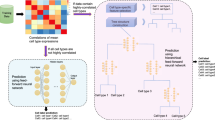
Single-cell multiomics data continues to grow at an unprecedented pace. Although several methods have demonstrated promising results in integrating several data modalities from the same tissue, the complexity and scale of data compositions present in cell atlases still pose a challenge. Here, we present scJoint, a transfer learning method to integrate atlas-scale, heterogeneous collections of scRNA-seq and scATAC-seq data. scJoint leverages information from annotated scRNA-seq data in a semisupervised framework and uses a neural network to simultaneously train labeled and unlabeled data, allowing label transfer and joint visualization in an integrative framework. Using atlas data as well as multimodal datasets generated with ASAP-seq and CITE-seq, we demonstrate that scJoint is computationally efficient and consistently achieves substantially higher cell-type label accuracy than existing methods while providing meaningful joint visualizations. Thus, scJoint overcomes the heterogeneity of different data modalities to enable a more comprehensive understanding of cellular phenotypes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All single-cell datasets used in this paper are publicly available. • Mouse atlas data. The scRNA-seq dataset was downloaded from Tabula Muris5 (https://tabula-muris.ds.czbiohub.org/). The sci-ATAC-seq dataset of Cusanovich et al.26 was downloaded from https://atlas.gs.washington.edu/mouse-atac/. • Human fetal atlas data. The scRNA-seq dataset from Cao et al.27 was downloaded from GSE156793. The scATAC-seq dataset from Domcke et al.28 was downloaded from GSE149683. • SNARE-seq data. The SNARE-seq dataset of adult mouse cerebral cortex14 was downloaded from GSE126074. • Multimodal PBMC data. The ASAP-seq and CITE-seq datasets from Mimitou et al.34 were obtained from GSE156478. • Human hematopoiesis data. The scRNA-seq and scATAC-seq datasets from Granja et al.40 were downloaded from https://github.com/GreenleafLab/MPAL-Single-Cell-2019.
Code availability
scJoint was implemented using PyTorch (v.1.0.0) with code available at https://github.com/SydneyBioX/scJoint.
References
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. 20, 257–272 (2019).
Berger, S. L. The complex language of chromatin regulation during transcription. Nature 447, 407–412 (2007).
Klemm, S. L., Shipony, Z. & Greenleaf, W. J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 20, 207–220 (2019).
Pott, S. & Lieb, J. D. Single-cell atac-seq: strength in numbers. Genome Biol. 16, 172 (2015).
Schaum, N. et al. Single-cell transcriptomics of 20 mouse organs creates a tabula muris: the tabula muris consortium. Nature 562, 367 (2018).
Regev, A. et al. Science forum: the human cell atlas. eLife 6, e27041 (2017).
Lopez, R., Regier, J., Cole, M. B., Jordan, M. I. & Yosef, N. Deep generative modeling for single-cell transcriptomics. Nat. Methods 15, 1053–1058 (2018).
Wang, J. et al. Data denoising with transfer learning in single-cell transcriptomics. Nat. Methods 16, 875–878 (2019).
Lin, Y. et al. scMerge leverages factor analysis, stable expression, and pseudoreplication to merge multiple single-cell RNA-seq datasets. Proc. Natl Acad. Sci. USA 116, 9775–9784 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with harmony. Nat. Methods 16, 1289–1296 (2019).
Wang, T. et al. Bermuda: a novel deep transfer learning method for single-cell RNA sequencing batch correction reveals hidden high-resolution cellular subtypes. Genome Biol. 20, 165 (2019).
Amodio, M. et al. Exploring single-cell data with deep multitasking neural networks. Nat. Methods 16, 1139–1145 (2019).
Xiong, L. et al. Scale method for single-cell atac-seq analysis via latent feature extraction. Nat. Commun. 10, 4576 (2019).
Chen, S., Lake, B. B. & Zhang, K. High-throughput sequencing of the transcriptome and chromatin accessibility in the same cell. Nat. Biotechnol. 37, 1452–1457 (2019).
Cao, J. et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 361, 1380–1385 (2018).
Jin, S., Zhang, L. & Nie, Q. scAI: an unsupervised approach for the integrative analysis of parallel single-cell transcriptomic and epigenomic profiles. Genome Biol. 21, 25 (2020).
Argelaguet, R. et al. MOFA+: a statistical framework for comprehensive integration of multi-modal single-cell data. Genome Biol. 21, 111 (2020).
Welch, J. D., Hartemink, A. J. & Prins, J. F. MATCHER: manifold alignment reveals correspondence between single cell transcriptome and epigenome dynamics. Genome Biol. 18, 138 (2017).
Amodio, M. & and Krishnaswamy, S. MAGAN: aligning biological manifolds. In Proc. 35th International Conference on Machine Learning (eds. Dy, J. & Krause, A.) 215–223 (PMLR, 2018).
Liu, J., Huang, Y., Vert, J.-P. & Noble, W. S. Jointly embedding multiple single-cell omics measurements. Algorithms Bioinform. 143, 10 (2019).
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887 (2019).
Duren, Z. et al. Integrative analysis of single-cell genomics data by coupled nonnegative matrix factorizations. Proc. Natl Acad. Sci. USA 115, 7723–7728 (2018).
Barkas, N. et al. Joint analysis of heterogeneous single-cell RNA-seq dataset collections. Nat. Methods 16, 695–698 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
DaiYang, K. et al. Multi-domain translation between single-cell imaging and sequencing data using autoencoders. Nat. Commun. 12, 31 (2021).
Cusanovich, D. A. et al. A single-cell atlas of in vivo mammalian chromatin accessibility. Cell 174, 1309–1324 (2018).
Cao, J. A human cell atlas of fetal gene expression. Science 370, eaba7721 (2020).
Domcke, S. A human cell atlas of fetal chromatin accessibility. Science 370, eaba7612 (2020).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Machine Learning Res. 9, 2579–2605 (2008).
McInnes, L., Healy, J. & Melville, J. UMAP: Uniform manifold approximation and projection for dimension reduction. Preprint at arXiv https://arxiv.org/abs/1802.03426 (2018).
Pliner, H. A. et al. Cicero predicts cis-regulatory DNA interactions from single-cell chromatin accessibility data. Mol. Cell 71, 858–871 (2018).
Stuart, T., Srivastava, A., Madad, S., Lareau, C. A. & Satija, R. Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341 (2021).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865 (2017).
Mimitou, E. P. et al. Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells. Nat. Biotechnol. 39, 1246–1258 (2021).
Kim, H. J., Lin, Y., Geddes, T. A., Yang, J. Y. H. & Yang, P. CiteFuse enables multi-modal analysis of CITE-seq data. Bioinformatics 36, 4137–4143 (2020).
Godfrey, D. I., MacDonald, H. R., Kronenberg, M., Smyth, M. J. & Van Kaer, L. NKT cells: what’s in a name? Nat. Rev. Immunol. 4, 231–237 (2004).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Hao, Y. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Abdelaal, T. et al. A comparison of automatic cell identification methods for single-cell RNA sequencing data. Genome Bol. 20, 194 (2019).
Granja, J. M. et al. Single-cell multiomic analysis identifies regulatory programs in mixed-phenotype acute leukemia. Nat. Biotechnol. 37, 1458–1465 (2019).
Wu, K. E., Yost, K. E., Chang, H. Y. & Zou, J. Babel enables cross-modality translation between multiomic profiles at single-cell resolution. Proc. Natl Acad. Sci. USA 118, e2023070118 (2021).
Maecker, H. T., McCoy, J. P. & Nussenblatt, R. Standardizing immunophenotyping for the human immunology project. Nat. Rev. Immunol. 12, 191–200 (2012).
Qiu, P. Embracing the dropouts in single-cell RNA-seq analysis. Nat. Commun. 11, 1169 (2020).
Jiang, R., Sun, T., Song, D. & Li, J. J. Zeros in scRNA-seq data: good or bad? how to embrace or tackle zeros in scRNA-seq data analysis? Preprint at bioRxiv (2020).
Acknowledgements
We gratefully acknowledge the following funding sources: Research Training Program Tuition Fee Offset and Stipend Scholarship and Chen Family Research Scholarship to Y.L.; Australian Research Council Discovery Project grant (DP170100654) and AIR@innoHK program of the Innovation and Technology Commission of Hong Kong to J.Y.H.Y.; Australian Research Council DECRA Fellowship (DE180101252) to Y.X.R.W; NIH grants R01 HG010359 and P50 HG007735 to W.H.W.
Author information
Authors and Affiliations
Contributions
T.-Y.W., W.H.W. and Y.X.R.W. conceived and designed this project; Y.L., T.-Y.W. and S.W. performed data preprocessing, model development and evaluation of results; J.Y.H.Y., W.H.W. and Y.X.R.W. supervised the execution; Y.L., J.Y.H.Y., W.H.W. and Y.X.R.W. wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Jingshu Wang, Nancy Zhang and Qing Nie for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–25, Tables 1 and 2 and Note.
Rights and permissions
About this article
Cite this article
Lin, Y., Wu, TY., Wan, S. et al. scJoint integrates atlas-scale single-cell RNA-seq and ATAC-seq data with transfer learning. Nat Biotechnol 40, 703–710 (2022). https://doi.org/10.1038/s41587-021-01161-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-021-01161-6
This article is cited by
-
Semi-supervised integration of single-cell transcriptomics data
Nature Communications (2024)
-
Inferring gene regulatory networks from single-cell multiome data using atlas-scale external data
Nature Biotechnology (2024)
-
Progress in single-cell multimodal sequencing and multi-omics data integration
Biophysical Reviews (2024)
-
DeepGAMI: deep biologically guided auxiliary learning for multimodal integration and imputation to improve genotype–phenotype prediction
Genome Medicine (2023)
-
scNAT: a deep learning method for integrating paired single-cell RNA and T cell receptor sequencing profiles
Genome Biology (2023)