Abstract
Phylogenetic trees provide a framework for organizing evolutionary histories across the tree of life and aid downstream comparative analyses such as metagenomic identification. Methods that rely on single-marker genes such as 16S rRNA have produced trees of limited accuracy with hundreds of thousands of organisms, whereas methods that use genome-wide data are not scalable to large numbers of genomes. We introduce updating trees using divide-and-conquer (uDance), a method that enables updatable genome-wide inference using a divide-and-conquer strategy that refines different parts of the tree independently and can build off of existing trees, with high accuracy and scalability. With uDance, we infer a species tree of roughly 200,000 genomes using 387 marker genes, totaling 42.5 billion amino acid residues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Microbial genomes are publically available via RefSeq (https://ftp.ncbi.nlm.nih.gov/refseq/release/). Microbial and simulated gene sequences/alignments, intermediate and output files from the analysis of biological and simulated data are openly available at Harvard Dataverse64 (https://doi.org/10.7910/DVN/BCUM6P). Microbial tree output files and postprocessing data are available at Zenodo65 (https://doi.org/10.5281/zenodo.8057941).
Code availability
The code is publicly available at https://github.com/balabanmetin/uDance25 under BSD 3-Clause license.
Change history
18 October 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41587-023-02027-9
References
Gonzalez, A. et al. Qiita: rapid, web-enabled microbiome meta-analysis. Nat. Methods 15, 796–798 (2018).
Zhu, Q. et al. Phylogeny-aware analysis of metagenome community ecology based on matched reference genomes while bypassing taxonomy. mSystems 7, e00167-22 (2022).
Nayfach, S., Shi, Z. J., Seshadri, R., Pollard, K. S. & Kyrpides, N. C. New insights from uncultivated genomes of the global human gut microbiome. Nature 568, 505–510 (2019).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Zhu, Q. et al. Phylogenomics of 10,575 genomes reveals evolutionary proximity between domains Bacteria and Archaea. Nat. Commun. 10, 5477 (2019).
Parks, D. H. et al. A complete domain-to-species taxonomy for Bacteria and Archaea. Nat. Biotechnol. 38, 1079–1086 (2020).
Mirarab, S., Nakhleh, L. & Warnow, T. Multispecies coalescent: theory and applications in phylogenetics. Annu. Rev. Ecol. Evol. Syst. 52, 247–268 (2021).
Davidson, R., Vachaspati, P., Mirarab, S. & Warnow, T. Phylogenomic species tree estimation in the presence of incomplete lineage sorting and horizontal gene transfer. BMC Genomics 16, S1 (2015).
Maddison, W. P. Gene trees in species trees. Syst. Biol. 46, 523–536 (1997).
Degnan, J. H. & Rosenberg, N. A. Gene tree discordance, phylogenetic inference and the multispecies coalescent. Trends Ecol. Evol. 24, 332–340 (2009).
Gogarten, J. P., Doolittle, W. F. & Lawrence, J. G. Prokaryotic evolution in light of gene transfer. Mol. Biol. Evol. 19, 2226–2238 (2002).
Creevey, C. J., Doerks, T., Fitzpatrick, D. A., Raes, J. & Bork, P. Universally distributed single-copy genes indicate a constant rate of horizontal transfer. PLoS ONE 6, e22099 (2011).
Yan, Z., Smith, M. L., Du, P., Hahn, M. W. & Nakhleh, L. Species tree inference methods intended to deal with incomplete lineage sorting are robust to the presence of paralogs. Syst. Biol. 71, 367–381 (2022).
Asnicar, F. et al. Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0. Nat. Commun. 11, 2500 (2020).
Kozlov, A. M., Darriba, D., Flouri, T., Morel, B. & Stamatakis, A. RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35, 4453–4455 (2019).
Mirarab, S. et al. ASTRAL: genome-scale coalescent-based species tree estimation. Bioinformatics 30, i541–i548 (2014).
Matsen, F. A., Kodner, R. B. & Armbrust, E. V. pplacer: linear time maximum-likelihood and Bayesian phylogenetic placement of sequences onto a fixed reference tree. BMC Bioinformatics 11, 538 (2010).
Rabiee, M. & Mirarab, S. INSTRAL: discordance-aware phylogenetic placement using quartet scores. Syst. Biol. 69, 384–391 (2020).
Wedell, E., Cai, Y. & Warnow, T. SCAMPP: scaling alignment-based phylogenetic placement to large trees. IEEE/ACM Trans. Comput. Biol. Bioinform. 20, 1417–1430 (2023).
Barbera, P. et al. EPA-ng: massively parallel evolutionary placement of genetic sequences. Syst. Biol. 68, 365–369 (2019).
Warnow, T. (ed.) Bioinformatics and Phylogenetics 121–150 (Springer, 2019).
Nelesen, S. M., Liu, K., Wang, L.-S., Linder, C. R. & Warnow, T. DACTAL: divide-and-conquer trees (almost) without alignments. Bioinformatics 28, i274–i282 (2012).
Huson, D. H., Nettles, S. M. & Warnow, T. J. Disk-covering, a fast-converging method for phylogenetic tree reconstruction. J. Comput. Biol. 6, 369–386 (1999).
Balaban, M. et al. Generation of accurate, expandable phylogenomic trees with uDance. GitHub https://github.com/balabanmetin/uDance (2023).
Balaban, M., Jiang, Y., Roush, D., Zhu, Q. & Mirarab, S. Fast and accurate distance-based phylogenetic placement using divide and conquer. Mol. Ecol. Resour. 22, 1213–1227 (2022).
Rabiee, M. & Mirarab, S. Forcing external constraints on tree inference using ASTRAL. BMC Genomics 21, 218 (2020).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree-2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Yin, J., Zhang, C. & Mirarab, S. ASTRAL-MP: scaling ASTRAL to very large datasets using randomization and parallelization. Bioinformatics 35, 3961–3969 (2019).
Vachaspati, P. & Warnow, T. ASTRID: accurate species TRees from internode distances. BMC Genomics 16, S3 (2015).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 6, 610–618 (2012).
Parks, D. H. et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 36, 996–1004 (2018).
Coleman, G. A. et al. A rooted phylogeny resolves early bacterial evolution. Science 372, eabe0511 (2021).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Sayyari, E. & Mirarab, S. Fast coalescent-based computation of local branch support from quartet frequencies. Mol. Biol. Evol. 33, 1654–1668 (2016).
Leebens-Mack, J. H. et al. One thousand plant transcriptomes and the phylogenomics of green plants. Nature 574, 679–685 (2019).
Jiang, Y., Balaban, M., Zhu, Q. & Mirarab, S. DEPP: deep learning enables extending species trees using single genes. Syst. Biol. 72, 17–34 (2023).
Jiang, Y., Tabaghi, P. & Mirarab, S. Learning hyperbolic embedding for phylogenetic tree placement and updates. Biology 11, 1256 (2022).
Nasko, D. J., Koren, S., Phillippy, A. M. & Treangen, T. J. RefSeq database growth influences the accuracy of k-mer-based lowest common ancestor species identification. Genome Biol. 19, 165 (2018).
Locey, K. J. & Lennon, J. T. Scaling laws predict global microbial diversity. Proc. Natl Acad. Sci. USA 113, 5970–5975 (2016).
Fullam A. et al. proGenomes3: approaching one million accurately and consistently annotated high-quality prokaryotic genomes. Nucleic Acids Res. 51, D760–D766 (2023).
Jukes, T. H. & Cantor, C. R. in Mammalian Protein Metabolism Vol. 3 (ed. Munro, H. N.) 21–132 (Academic Press, 1969).
Sonnhammer, E. L. L. & Hollich, V. Scoredist: a simple and robust protein sequence distance estimator. BMC Bioinformatics 6, 108 (2005).
Darriba, D. et al. ModelTest-NG: a new and scalable tool for the selection of DNA and protein evolutionary models. Mol. Biol. Evol. 37, 291–294 (2020).
Anisimova, M., Gil, M., Dufayard, J.-F., Dessimoz, C. & Gascuel, O. Survey of branch support methods demonstrates accuracy, power, and robustness of fast likelihood-based approximation schemes. Syst. Biol. 60, 685–699 (2011).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Zhang, C., Zhao, Y., Braun, E. L. & Mirarab, S. TAPER: pinpointing errors in multiple sequence alignments despite varying rates of evolution. Methods Ecol. Evol. 12, 2145–2158 (2021).
Sayyari, E., Whitfield, J. B. & Mirarab, S. Fragmentary gene sequences negatively impact gene tree and species tree reconstruction. Mol. Biol. Evol. 34, 3279–3291 (2017).
Mai, U. & Mirarab, S. TreeShrink: fast and accurate detection of outlier long branches in collections of phylogenetic trees. BMC Genomics 19, 272 (2018).
Balaban, M., Moshiri, N., Mai, U., Jia, X. & Mirarab, S. TreeCluster: clustering biological sequences using phylogenetic trees. PLoS ONE 14, e0221068 (2019).
Mölder, F. et al. Sustainable data analysis with Snakemake. F1000Res. 10, 33 (2021).
Mallo, D., De Oliveira Martins, L. & Posada, D. SimPhy: phylogenomic simulation of gene, locus, and species trees. Syst. Biol. 65, 334–344 (2016).
Fletcher, W. & Yang, Z. INDELible: a flexible simulator of biological sequence evolution. Mol. Biol. Evol. 26, 1879–1888 (2009).
Nguyen, N. D., Mirarab, S., Kumar, K. & Warnow, T. Ultra-large alignments using phylogeny-aware profiles. Genome Biol. 16, 124 (2015).
Yang, Z., Nielsen, R., Goldman, N. & Pedersen, A.-M. K. Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics 155, 431–449 (2000).
Haft, D. H. et al. RefSeq: an update on prokaryotic genome annotation and curation. Nucleic Acids Res. 46, D851–D860 (2018).
Segata, N., Börnigen, D., Morgan, X. C. & Huttenhower, C. PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nat. Commun. 4, 2304 (2013).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Darling, A. E. et al. PhyloSift: phylogenetic analysis of genomes and metagenomes. PeerJ 2, e243 (2014).
Orakov, A. et al. GUNC: detection of chimerism and contamination in prokaryotic genomes. Genome Biol. 22, 178 (2021).
Le, S. Q. & Gascuel, O. An improved general amino acid replacement matrix. Mol. Biol. Evol. 25, 1307–1320 (2008).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Wickett, N. J. et al. Phylotranscriptomic analysis of the origin and early diversification of land plants. Proc. Natl Acad. Sci. USA 111, 4859–4868 (2014).
Balaban, M. et al. Data for article: generation of accurate, expandable phylogenomic trees with uDance. Harvard Dataverse https://doi.org/10.7910/DVN/BCUM6P (2023).
Balaban, M. et al. Postprocessing data for article: generation of accurate, expandable phylogenomic trees with uDance. Zenodo https://doi.org/10.5281/zenodo.8057941 (2023).
Acknowledgements
This work was supported by the National Science Foundation (NSF; grants IIS 1845967 to S.M. and RAPID 20385.09 to R.K.) and National Institutes of Health (NIH; grant 1R35GM142725 to S.M.; grants U19AG063744, U24DK131617 and DP1-AT010885 to R.K.). It was also supported by the 2020 UCSD Center for Microbiome Innovation Grand Challenge Award to M.B. Computations were partially performed using Expanse at San Diego Supercomputing Center through allocations ASC150046 and BIO210103 from the Advanced Cyberinfrastructure Coordination Ecosystem—Services & Support (ACCESS) program, which is supported by NSF under grants 2138259, 2138286, 2138307, 2137603 and 2138296.
Author information
Authors and Affiliations
Contributions
S.M. and M.B. conceived and designed the uDance method. M.B., Y.J. and S.M. performed simulation studies. All authors contributed to building and analyses of the real biological dataset. All authors contributed to the writing of the paper. All authors reviewed and edited the paper.
Corresponding author
Ethics declarations
Competing interests
D.M. is a consultant for BiomeSense, Inc., has equity and receives income. The terms of these arrangements have been reviewed and approved by the University of California, San Diego in accordance with its conflict of interest policies. R.K. is a scientific advisory board member, and consultant for BiomeSense, Inc., has equity and receives income. He is a scientific advisory board member and has equity in GenCirq. He is a consultant and scientific advisory board member for DayTwo, and receives income. He has equity in and acts as a consultant for Cybele. He is a co-founder of Biota, Inc., and has equity. He is a cofounder of Micronoma, and has equity and is a scientific advisory board member. The terms of this arrangement have been reviewed and approved by the University of California, San Diego in accordance with its conflict of interest policies. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Gene tree estimation error in simulated datasets.
(a) nRF and (b) QD distance between estimated and true gene trees for all partition-gene pairs in all model conditions in the simulated dataset. RAxML-ng is used inside uDance (u) and on subsets whereas FT2 (a) is used on the full dataset. The calculation of errors is always on the subsets to obtain fair comparisons.
Extended Data Fig. 2 Inserting a small number of queries on a large tree using uDance in auto and fast mode.
We selected the 16,000 genomes from the analysis used in Fig. 2g and updated it with \(\{5\times {2}^{i}| 1\le i\le 8,i\in {\mathbb{Z}}\}\) genomes using uDance in the auto (standard) and fast insertion mode (where the only difference is that partition sizes are set to 100). We measured the delta error for each query, which is defined as the change in RF distance between the true tree and inferred tree after placement of the query sequence. We show the mean delta error versus CPU time for various query set sizes. The running time grows slower with the -fast mode without a significant sacrifice in accuracy. Whether the fast or the default modes is used, the accuracy is substantially higher when we allow the backbone tree to change (maxqs) compared to fixed backbone (incremental). In fact, the accuracy improves after addition if the update mode (maxqs) is used whereas the accuracy stays the same or degrades with the incremental model or simple placement.
Extended Data Fig. 3 10K tree and comparisons to other trees.
(a) The 10K ASTRAL tree decorated with GTDB taxonomy. (b) Pairwise quartet distance between the 10k, 16k, 200k, and GTDB trees, restricted to NCBI phyla and super-phyla.
Extended Data Fig. 4 ECDF of depth and the branch length of agreeing and disagreeing branches with backbone.
ECDF of depth and the branch length of agreeing and disagreeing branches between the backbone and output phylogenies for both 16k and 200k trees.
Extended Data Fig. 5 The paraphyletic Myxococcota phylum on the GTDB phylogenetic tree.
Green and red sequences represent the members of the phylum that are proximal Desulfobacteria and Proteobacteria respectively in the 200k tree.
Extended Data Fig. 6 Branch support (local posterior probability) patterns.
(a) ECDF of branch support across partitions of the 16k tree. (b) Branch support versus the diversity (average branch length) of all 78 partitions in the 200k tree. The dot and the range indicate median and 0.25-0.75 quantiles. Three colors correspond to clusters that are unusually small, unusually large, or typical in size. Fourteen of the 15 partitions with the lowest diversity are of size between 2,500 and 6,000. The largest partitions in the 200k tree are over-represented parts of the tree of life in the reference genomic library that did not break into smaller partitions by uDance because of their lower diversity. (c) Number of uncollapsed branches vs branch support (localPP) collapsing threshold for 16k and 200k trees.
Extended Data Fig. 7 Model heterogeneity across 16k dataset.
Divide-and-conquer approach permits heterogeneity of model of evolution parameters across the tree. In discrete four-category LG+GAMMA model, the rate heterogeneity across sites is modeled by a discrete approximation of the Gamma distribution. The first and the fourth discrete rate of LG+G model for every partition and gene pair in the 16k tree are shown in (a) and (b), respectively. The partition hierarchy created by uDance is shown on the left. A blank cell indicates a missing gene in a partition.
Extended Data Fig. 8 Schematic of Outgroup taxa selection strategy and stitching.
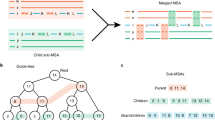
(a) Outgroup taxa selection strategy. Two to three taxa are chosen from the partition c2 (blue) to be added to the partition c1 (orange). (b) Finding junction node v in Constrained ASTRAL tree for color (partition) c. We illustrate the setup for Claims 2 and 3. (c) Stitching happens at junction nodes. After removing taxa placed on outgroup branches, other subtrees can be stitched to this subtree without any need for conceptual merge, but simply replacing the connecting nodes.
Extended Data Fig. 9 Data Selection and Quality Checks.
(a) Determining the backbone size in the simulated HD-100 dataset. ECDF of novelty of query sequences with respect to a backbone tree of N downsampled sequences induced from the full HGT dataset. With N = 1000 sequences selected using TreeCluster-max, for more than 95% of the query sequences, the novelty score is less than one. Novelty score is defined as two times the terminal branch length of the query when placed on the true location on the backbone tree. (b) The distribution of number of marker genes per sequence in WoL2 dataset. (c) Two dot plots comparing (1) contamination ratio-vs-CSS and (2) contamination ratio-vs-GUNC database identity for the species in the 16K tree that are ‘chimeric’ (CSS > 0.45). We colored each point based on whether the sequence passed QC in GTDB or not. Triangle points are the published WoL tree, and round points are the new 6K taxa we added in the 16K tree. In these figures, We annotated 17 taxa in the 16K tree that might be reducing the accuracy of uDance and APPLES-2 in large clusters (subtrees) that include some of the densely sampled species such as Salmonella, E. coli, TB, etc. The pattern is clear that these contaminated genomes can be characterized by a large contamination ratio, near 100% CSS, and high database identity. We do not remove high CSS taxa if their contamination percentage is low, since uDance performs whole-genome-based placement, and it’s tolerant to low levels of contamination. Removing taxa satisfying both CSS ≥ 0.5 and Contamination ratio ≥ 0.25 removes 195 taxa from the 16K tree. 171 of them (87%) fail QC in GTDB. Of 195, 37 taxa are also present in WoL tree. 29 of these 37 don’t pass GTDB QC. (d) Determining ‘best’ marker genes to be used with APPLES-2 in order to improve placement speed. We picked a local maxima of average Archaea occupancy at 68th marker gene, which also ensures that, on average, Archaea sequences have at least 20 marker genes. The set of Archaea used in computation of these two statistics is taken from WoL tree. The name of the genes (shown on x-axis) are not important and can be ignored.
Supplementary information
Supplementary Information
Supplementary Tables 1–4 and Supplementary Notes 1–4.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Balaban, M., Jiang, Y., Zhu, Q. et al. Generation of accurate, expandable phylogenomic trees with uDance. Nat Biotechnol (2023). https://doi.org/10.1038/s41587-023-01868-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41587-023-01868-8
This article is cited by
-
Generation of accurate, expandable phylogenomic trees with uDance
Nature Biotechnology (2023)
-
Greengenes2 unifies microbial data in a single reference tree
Nature Biotechnology (2023)