Abstract
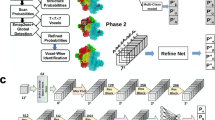
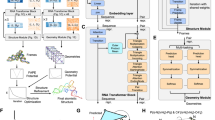
Many methods exist for determining protein structures from cryogenic electron microscopy maps, but this remains challenging for RNA structures. Here we developed EMRNA, a method for accurate, automated determination of full-length all-atom RNA structures from cryogenic electron microscopy maps. EMRNA integrates deep learning-based detection of nucleotides, three-dimensional backbone tracing and scoring with consideration of sequence and secondary structure information, and full-atom construction of the RNA structure. We validated EMRNA on 140 diverse RNA maps ranging from 37 to 423 nt at 2.0–6.0 Å resolutions, and compared EMRNA with auto-DRRAFTER, phenix.map_to_model and CryoREAD on a set of 71 cases. EMRNA achieves a median accuracy of 2.36 Å root mean square deviation and 0.86 TM-score for full-length RNA structures, compared with 6.66 Å and 0.58 for auto-DRRAFTER. EMRNA also obtains a high residue coverage and sequence match of 93.30% and 95.30% in the built models, compared with 58.20% and 42.20% for phenix.map_to_model and 56.45% and 52.3% for CryoREAD. EMRNA is fast and can build an RNA structure of 100 nt within 3 min.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw data of the evaluation results are provided in the article and supplementary tables. All published data sets used in this paper were taken from the EMDB and PDB (accession codes specified in the figure captions and in supplementary tables). The EMRNA input maps and output models used in the study are available at https://zenodo.org/records/10225107 ref. 67. Source data are provided with this paper.
Code availability
The EMRNA package is freely available for academic or noncommercial users at http://huanglab.phys.hust.edu.cn/EMRNA/ or https://zenodo.org/records/10540040 (ref. 68).
References
Nogales, E. The development of cryo-EM into a mainstream structural biology technique. Nat. Methods 13, 24–27 (2016).
Frank, J. Advances in the field of single-particle cryo-electron microscopy over the last decade. Nat. Protoc. 12, 209–212 (2017).
Cheng, Y. Single-particle cryo-EM—how did it get here and where will it go. Science 361, 876–880 (2018).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Terwilliger, T. C., Adams, P. D., Afonine, P. V. & Sobolev, O. V. A fully automatic method yielding initial models from high-resolution cryo-electron microscopy maps. Nat. Methods 15, 905–908 (2018).
Terwilliger, T. C., Adams, P. D., Afonine, P. V. & Sobolev, O. V. Cryo-EM map interpretation and protein model building using iterative map segmentation. Protein Sci. 29, 87–99 (2020).
de la Rosa-Trevín, J. M. et al. Scipion: a software framework toward integration, reproducibility and validation in 3D electron microscopy. J. Struct. Biol. 195, 93–99 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Lawson, C. L. et al. EMDataBank unified data resource for 3DEM. Nucleic Acids Res. 44, D396–D403v2016 (2016).
Ma, H., Jia, X., Zhang, K. & Su, Z. Cryo-EM advances in RNA structure determination. Signal Transduct. Target. Ther. 7, 58 (2022).
Su, Z. et al. Cryo-EM structures of full-length Tetrahymena ribozyme at 3.1 Å resolution. Nature 596, 603–607 (2021).
Bonilla, S. L., Vicens, Q. & Kieft, J. S. Cryo-EM reveals an entangled kinetic trap in the folding of a catalytic RNA. Sci. Adv. 8, eabq4144 (2022).
Luo, B. et al. Cryo-EM reveals dynamics of Tetrahymena group I intron self-splicing. Nat. Catal. 6, 298–309 (2023).
Li, S. et al. Topological crossing in the misfolded Tetrahymena ribozyme resolved by cryo-EM. Proc. Natl Acad. Sci. USA 119, e2209146119 (2022).
Zhang, X., Li, S., Pintilie, G., Palo, M. Z. & Zhang, K. Snapshots of the first-step self-splicing of Tetrahymena ribozyme revealed by cryo-EM. Nucleic Acids Res. 51, 1317–1325 (2023).
Li, S., Palo, M. Z., Zhang, X., Pintilie, G. & Zhang, K. Snapshots of the second-step self-splicing of Tetrahymena ribozyme revealed by cryo-EM. Nat. Commun. 14, 1294 (2023).
Liu, D., Thélot, F. A., Piccirilli, J. A., Liao, M. & Yin, P. Sub-3-Å cryo-EM structure of RNA enabled by engineered homomeric self-assembly. Nat. Methods 19, 576–585 (2022).
Baker, M. L. et al. Modeling protein structure at near atomic resolutions with Gorgon. J. Struct. Biol. 174, 360–373 (2011).
Lindert, S. et al. EM-fold: de novo folding of α-helical proteins guided by intermediate-resolution electron microscopy density maps. Structure 17, 990–1003 (2009).
Chen, M. & Baker, M. L. Automation and assessment of de novo modeling with Pathwalking in near atomic resolution cryoEM density maps. J. Struct. Biol. 204, 555–563 (2018).
Wang, R. Y. et al. De novo protein structure determination from near-atomic-resolution cryo-EM maps. Nat. Methods 12, 335–338 (2015).
Frenz, B., Walls, A. C., Egelman, E. H., Veesler, D. & DiMaio, F. RosettaES: a sampling strategy enabling automated interpretation of difficult cryo-EM maps. Nat. Methods 14, 797–800 (2017).
Terashi, G. & Kihara, D. De novo main-chain modeling for EM maps using MAINMAST. Nat. Commun. 9, 1618 (2018).
Si, D. et al. Deep learning to predict protein backbone structure from high-resolution cryo-EM density maps. Sci. Rep. 10, 4282 (2020).
He, J. & Huang, S. Y. Full-length de novo protein structure determination from cryo-EM maps using deep learning. Bioinformatics 37, 3480–3490 (2021).
Pfab, J., Phan, N. M. & Si, D. DeepTracer for fast de novo cryo-EM protein structure modeling and special studies on CoV-related complexes. Proc. Natl Acad. Sci. USA 118, e2017525118 (2021).
He, J. & Huang, S. Y. EMNUSS: a deep learning framework for secondary structure annotation in cryo-EM maps. Brief. Bioinform. 22, bbab156 (2021).
He, J., Lin, P., Chen, J., Cao, H. & Huang, S. Y. Model building of protein complexes from intermediate-resolution cryo-EM maps with deep learning-guided automatic assembly. Nat. Commun. 13, 4066 (2022).
Zhang, X., Zhang, B., Freddolino, P. L. & Zhang, Y. CR-I-TASSER: assemble protein structures from cryo-EM density maps using deep convolutional neural networks. Nat. Methods 19, 195–204 (2022).
Zhou, X. et al. Progressive assembly of multi-domain protein structures from cryo-EM density maps. Nat. Comput. Sci. 2, 265–275 (2022).
Jamali, K., Kimanius, D. & Scheres, S. H. A graph neural network approach to automated model building in cryo-EM maps. In The 11th International Conference on Learning Representations (ICLR, 2022).
Kappel, K. et al. De novo computational RNA modeling into cryo-EM maps of large ribonucleoprotein complexes. Nat. Methods 15, 947–954 (2018).
Kappel, K. et al. Accelerated cryo-EM-guided determination of three-dimensional RNA-only structures. Nat. Methods 17, 699–707 (2020).
Nguyen, T. H. D. et al. The architecture of the spliceosomal U4/U6. U5 tri-snRNP. Nature 523, 47–52 (2015).
Greber, B. J. et al. Architecture of the large subunit of the mammalian mitochondrial ribosome. Nature 505, 515–519 (2014).
Chaker-Margot, M., Barandun, J., Hunziker, M. & Klinge, S. Architecture of the yeast small subunit processome. Science 355, eaal1880 (2017).
Li, X. et al. Structure of ribosomal silencing factor bound to Mycobacterium tuberculosis ribosome. Structure 23, 1858–1865 (2015).
Chojnowski, G. et al. Brickworx builds recurrent RNA and DNA structural motifs into medium-and low-resolution electron-density maps. Acta Crystallogr. D 71, 697–705 (2015).
Nakamura, A. et al. Fast and automated protein–DNA/RNA macromolecular complex modeling from cryo-EM maps. Brief. Bioinform. 24, bbac632 (2023).
Wang, X., Terashi, G. & Kihara, D. CryoREAD: de novo structure modeling for nucleic acids in cryo-EM maps using deep learning. Nat. Methods 20, 1739–1747 (2023).
Das, R., Karanicolas, J. & Baker, D. Atomic accuracy in predicting and designing noncanonical RNA structure. Nat. Methods 7, 291–294 (2010).
Watkins, A. M., Rangan, R. & Das, R. FARFAR2: improved de novo Rosetta prediction of complex global RNA folds. Structure 28, 963–976 (2020).
He, J., Li, T. & Huang, S. Y. Improvement of cryo-EM maps by simultaneous local and non-local deep learning. Nat. Commun. 14, 3217 (2023).
Tian, C. et al. ff19SB: amino-acid-specific protein backbone parameters trained against quantum mechanics energy surfaces in solution. J. Chem. Theory Comput. 16, 528–552 (2019).
Zgarbová, M. et al. Refinement of the Cornell et al. nucleic acids force field based on reference quantum chemical calculations of glycosidic torsion profiles. J. Chem. Theory Comput. 7, 2886–2902 (2011).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Gong, S., Zhang, C. & Zhang, Y. RNA-align: quick and accurate alignment of RNA 3D structures based on size-independent TM-scoreRNA. Bioinformatics 35, 4459–4461 (2019).
Singh, J., Hanson, J., Paliwal, K. & Zhou, Y. RNA secondary structure prediction using an ensemble of two-dimensional deep neural networks and transfer learning. Nat. Commun. 10, 5407 (2019).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 1–14 (2011).
Zhang, K. et al. Practical blind image denoising via Swin-Conv-UNet and data synthesis. Mach. Intell. Res. 20, 822–836 (2023).
Liu, Z. et al. Swin transformer: hierarchical vision transformer using shifted windows. In Proc. IEEE/CVF International Conference on Computer Vision 9992–10002 (IEEE, 2021).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Capriotti, E. & Marti-Renom, M. A. Quantifying the relationship between sequence and three-dimensional structure conservation in RNA. BMC Bioinform. 11, 1–10 (2010).
Wang, Z., Bovik, A. C., Sheikh, H. R. & Simoncelli, E. P. Image quality assessment: from error visibility to structural similarity. IEEE Trans. Image Process. 13, 600–612 (2004).
Helsgaun K. An effective implementation of the Lin–Kernighan traveling salesman heuristic. Eur. J. Oper. Res. 126, 106–130 (2000).
Wayment-Steele, H. K. et al. RNA secondary structure packages evaluated and improved by high-throughput experiments. Nat. Methods 19, 1234–1242 (2022).
Kabsch, W. A solution for the best rotation to relate two sets of vectors. Acta Crystallogr. A 32, 922–923 (1976).
Lu, X. J., Bussemaker, H. J. & Olson, W. K. DSSR: an integrated software tool for dissecting the spatial structure of RNA. Nucleic Acids Res. 43, e142–e142 (2015).
Zhang, C. & Pyle, A. M. CSSR: assignment of secondary structure to coarse-grained RNA tertiary structures. Acta Crystallogr. D 78, 466–471 (2022).
Zhao, Y. et al. Automated and fast building of three-dimensional RNA structures. Sci. Rep. 2, 734 (2012).
Wang, J. & Xiao, Y. Using 3dRNA for RNA 3-D structure prediction and evaluation. Curr. Protoc. Bioinform. 57, 5–9 (2017).
Zhang, Y., Wang, J. & Xiao, Y. 3dRNA: 3D structure prediction from linear to circular RNAs. J. Mol. Biol. 434, 167452 (2022).
Ma, H. et al. Auto-DRRAFTER: automated RNA modeling based on cryo-EM density. Methods Mol. Biol. 2568, 193–211 (2023).
Zhang, K. et al. Cryo-EM structure of a 40 kDa SAM-IV riboswitch RNA at 3.7 Å resolution. Nat. Commun. 10, 5511 (2019).
Li, T. & Huang, S.-Y. EMRNA: Accurate RNA structure determination from cryo-EM maps by deep learning and integrated modeling. Zenodohttps://zenodo.org/records/10225107 (2023).
Li, T. & Huang, S.-Y. EMRNA program. Zenodo https://zenodo.org/records/10540040 (2024).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grant nos. 32161133002, 62072199 and 32071247) and the startup grant of Huazhong University of Science and Technology. The computation is completed in the HPC Platform of Huazhong University of Science and Technology.
Author information
Authors and Affiliations
Contributions
S.-Y.H. and Y.X. conceived and supervised the project. T.L., J.H., H.C., Y.Z. and S.-Y.H. designed and performed the experiments. S.-Y.H. and T.L. analyzed the data. H.C. and J.C. tested the program. T.L. and S.-Y.H. wrote the paper. All authors read and approved the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Texts 1–3, Figs. 1–15 and Note.
Supplementary Table
Supplementary Tables 1–10.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, T., He, J., Cao, H. et al. All-atom RNA structure determination from cryo-EM maps. Nat Biotechnol (2024). https://doi.org/10.1038/s41587-024-02149-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41587-024-02149-8