Abstract
Structural variants (SVs) can contribute to oncogenesis through a variety of mechanisms. Despite their importance, the identification of SVs in cancer genomes remains challenging. Here, we present a framework that integrates optical mapping, high-throughput chromosome conformation capture (Hi-C), and whole-genome sequencing to systematically detect SVs in a variety of normal or cancer samples and cell lines. We identify the unique strengths of each method and demonstrate that only integrative approaches can comprehensively identify SVs in the genome. By combining Hi-C and optical mapping, we resolve complex SVs and phase multiple SV events to a single haplotype. Furthermore, we observe widespread structural variation events affecting the functions of noncoding sequences, including the deletion of distal regulatory sequences, alteration of DNA replication timing, and the creation of novel three-dimensional chromatin structural domains. Our results indicate that noncoding SVs may be underappreciated mutational drivers in cancer genomes.
Similar content being viewed by others
Main
Structural variants (SVs), including inversions, deletions, duplications, and translocations, are a hallmark of most cancer genomes1. The discovery of recurrent SVs and their molecular consequences for gene organization and expression has greatly advanced our knowledge of oncogenesis. Numerous oncogenes have been identified as the products of recurrent translocations and have provided successful targets for drug therapies2,3,4,5,6, particularly for hematopoietic malignancies.
Despite their importance, identifying SVs in cancer genomes remains challenging. Historically, G-band karyotyping has been the major method and is routinely performed in the clinic7. However, it is an inherently low-resolution and low-throughput method that cannot characterize extensively rearranged genomes. Microarrays are another commonly used method for detecting gains and losses of genetic material8, but they do not provide precise location of rearrangements and cannot detect balanced rearrangements. Targeted approaches such as fluorescence in situ hybridization (FISH) and PCR are also used extensively in the clinic. However, such methods require a priori knowledge of the rearrangements and hence are not suitable for de novo SV detection. Recently, high-throughput sequencing-based methods, such as RNA sequencing (RNA-seq) and whole-genome sequencing (WGS), have emerged as attractive methods for SV identification; they can identify gene fusions and genomic rearrangements with high resolution9,10,11,12,13,14,15,16. Despite their success, these short-read-based approaches cannot effectively detect SVs in the repetitive regions in the genome and have limited power to determine complex SVs in a haplotype-resolved manner.
Here we propose an integrative framework to comprehensively detect SVs by using a combination of technologies, including WGS, next-generation optical mapping (Irys system; Bionano Genomics), and high-throughput chromosome conformation capture (Hi-C). In addition, we developed a novel algorithm that uses Hi-C data to detect SVs genome-wide. By integrating the results from different platforms, we compiled a list of high-confidence SVs in eight human cancer cell lines (Table 1). We observed that optical mapping and Hi-C excelled at detecting large and complex structural alterations, whereas high-coverage WGS was adept at identifying SVs with high resolution. We identified numerous instances of three-dimensional (3D) genome organization alterations as a result of structural genome variation, such as the formation or dissolution of topologically associating domains (TADs), suggesting a critical role for structural variation in gene misregulation in oncogenesis.
Results
An integrated approach for SV detection
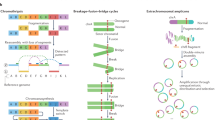
To evaluate the ability of different platforms in detecting SVs, we compared WGS, optical mapping, and Hi-C data in eight cancer cell lines and one karyotypically normal control (GM12878) (Fig. 1a and Supplementary Table 1). We generated WGS data in seven cancer cell lines with an average coverage of >30×, and downloaded the data for LNCaP and GM12878 cells from a previous study17 and the Platinum Genome Dataset (Illumina), respectively. We built an in-house pipeline that integrates the results from the LUMPY, Delly, and Control-FREEC software (see URLs)18,19,20 for initial SV detection, and then performed extensive data filtering (Supplementary Fig. 1 and Supplementary Tables 2,3). Next, we performed optical mapping in the same nine cell lines with an average coverage of approximately 100×. We used RefAligner 6119 (Bionano Genomics) and pipeline 6498 to conduct de novo assembly and SV detection, and we designed an in-house pipeline to perform further data filtering (Supplementary Fig. 2 and Supplementary Table 4). Lastly, we performed Hi-C experiments in 14 cancer cell lines and analyzed an additional 21 previously published data sets21,22,23,24,25,26,27. We developed a novel algorithm to use Hi-C data to identify rearrangement events, including translocations, inversions, deletions, and tandem duplications (Supplementary Table 5 and Supplementary Figs. 3 and 4). After comparing and merging the results from each platform, we identified thousands of insertions and deletions (>50 bp), hundreds of tandem duplications and interchromosomal translocations, and tens of inversions (Supplementary Table 2). We compiled a list of high-confidence SVs that were predicted by at least two methods (Supplementary Table 6). An example is shown in Fig. 1b, where a translocation between chromosomes 2 and 3 in Caki2 cells was detected by all three methods. This translocation was also validated by observation of dramatic shifts in DNA replication timing profiles in the same region. Finally, we observed that the cancer genomes displayed many more rearrangement events compared with normal cells, as illustrated by circular genome structural profiles28 (Fig. 1c, Supplementary Fig. 5).
a, The pipeline of SV detection, validation, and functional analysis. b, An example of the same translocation (TL) detected by different technologies in Caki2 cells (hg38 coordinates: chr2:204,260,308 and chr3:179,694,900). c, Cancer genomes possess more CNVs and translocations in comparison with karyotypically normal GM12878 cells. The tracks from outer to inner circles are chromosome coordinates, copy number, duplications (red), and deletions (blue), and rearrangements including inversions, interchromosomal TLs and unclassified rearrangements. The outward red bars in the CNV track indicate gain of copies (>2, 2–8 copies); the inward blue bars indicate loss of copies (<2, 0–2 copies). CNVs are profiled by WGS with a 50-kb bin size. Duplications, deletions, and translocations are detected by at least two methods from WGS, Irys, and Hi-C.
Detection of large-scale rearrangements using Hi-C data
Several groups have reported unusual interchromosomal interactions in Hi-C data and suggested these signals are the results of SVs21,29,30,31; however, to identify the breakpoints, they mainly relied on visual inspection32,33. Software tools have recently been developed to identify copy number alterations (CNAs) or interchromosomal translocations in Hi-C data sets32,33,34,35. However, to our knowledge, no algorithm has been developed that can use Hi-C for genome-wide detection of a full range of SVs, including deletions, inversions, tandem duplications, and interchromosomal translocations.
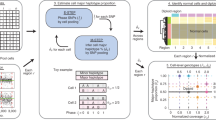
In a Hi-C experiment in karyotypically normal cells, interchromosomal interactions are rare (left panel in Fig. 2a). However, this pattern does not hold in cancer cells. For example, in Caki2 cancer cells, we observed strong “interchromosomal” interactions (right panel in Fig. 2a), which might be due to the fusion of chromosomes 6 and 8. The challenge is to determine whether the increased signals are due to rearrangement or normal variation in 3D genome organization. We first developed probabilistic models for “normal” 3D genome organization features, including genomic distance between loci, TADs, A/B compartments, and the increased interactions between small chromosomes and between subtelomeric regions (Supplementary Fig. 3 and Supplementary Methods (see Supplementary Note)). In the event of a rearrangement, the two rearranged regions are genetically fused, altering the linear distance between loci. This leads to local clusters of deviations from the expected interaction frequencies, and such patterns can be used to detect SVs (Fig. 2a,b). To systematically identify this signature, we developed an iterative approach to pinpoint local clusters of interaction frequencies suggestive of the presence of rearrangements. The method progressively reduces the bin size to refine the resolution of breakpoints to as high as 1 kb (Supplementary Fig. 6).
a,b, Interchromosomal (a) and intrachromosomal (b) rearrangements detected using Hi-C data (marked by the arrow sign). In panel (a), GM12878 heat maps are shown at 100-kb resolution; Caki2 are shown at 1 Mb resolution. c, A complex translocation (chr6-chr16-chr6) in K562 cells validated by FISH. Similar results for FISH validation experiments were performed using 20 independent metaphase nuclei. Scale bars (white) represent 5 μM. d, Number of interchromosomal and intrachromosomal rearrangements detected by Hi-C in 29 cancer genomes and 9 normal genomes. e, An example of the impact of translocations (TLs) on replication timing (RT). When plotted to the reference genome, the RT profiles of chr5 and chr10 of SK-N-MC show abrupt shifts at the TL breakpoints (←, left panels), and they are smoothly connected due to their juxtaposition in the cancer genome (right panel, normal chr10 is absent in SK-N-MC). The solid black (chr10) and red (chr5) lines indicate loess-smoothened RT data. Since the RT experiments were designed for validation purposes, one replicate was performed.
We first evaluated our algorithm with a well-characterized chronic myelogenous leukemia cell line (K562) and compared the results with the published karyotype. Of the 19 Hi-C predicted rearrangements, 11 can be confirmed and the remaining 8 are novel36. Since these eight events were found in both of the two replicate experiments that were performed in two independent laboratories, they are not likely a product of clonal evolution. Several of the events are complex rearrangements. One event is between chromosome 16 and two different regions of chromosome 6 (Fig. 2c). Another is a rearrangement involving chromosomes 1, 6, 18, and 20 (Supplementary Fig. 7). We performed FISH experiments to validate the novel predicted translocations; 18 of the 19 predicted translocations using Hi-C data were validated by either FISH or previous karyotyping (Supplementary Table 7), suggesting that our algorithm can identify large-scale structural variation with high specificity.
To further evaluate the algorithm, we performed Hi-C in ES cells derived from the Tc1 mouse (Supplementary Fig. 8a), which are engineered to carry a copy of human chromosome 2137. In the process of establishing this cell line, human chromosome 21 was subject to gamma irradiation37, leading to massive genomic rearrangements, a subset of which have been previously identified using PCR and Sanger sequencing38. We evaluated the sensitivity of our algorithm at various sequencing depths by subsampling, and found that the algorithm can achieve decent sensitivity with as few as 5–10 million reads. The performance reaches a plateau at approximately 100 million sequencing read pairs with a sensitivity of 90% (Supplementary Fig. 8b). The predicted breakpoints are internally consistent when at least 50 million reads are available (Supplementary Fig. 8c,d). We noticed that sometimes Hi-C and WGS call breakpoints in the same regions but report different strandedness (Supplementary Fig. 8e–h). The discrepancies usually involve complex events, where Hi-C reports the larger-scale SVs and WGS reports the smaller SVs for the sample complex event. To evaluate the effect of sample heterogeneity, we simulated mixed tumor/normal samples by combining Hi-C reads from K562 and GM12878 cells at different fractions while keeping the total sequencing depth at 100 million reads. We observed a limited loss of sensitivity even with tumor fractions as low as 30%, indicating that the performance of Hi-C-based SV finding is robust under conditions of moderate sample heterogeneity (Supplementary Fig. 8i).
Finally, we expanded our Hi-C analysis to 27 cancer cell lines and 9 karyotypically normal cell lines (Fig. 2d). On average, we reported 25 rearrangements in cancer cells and virtually no such events in normal cells, with an interchromosomal to intrachromosomal rearrangement ratio of roughly 2:1 (424 versus 274 in all cell lines). Our algorithm appears to identify mostly large SVs, with only 4.3% of intrachromosomal SVs being <2 Mb in size (Supplementary Fig. 8j). This is likely because it is challenging to distinguish the strong Hi-C signals resulting from structural variation from those strong local interaction signals within the same TAD.
Validation of Hi-C breakpoints by replication timing
Next, we compared our Hi-C-defined breakpoints with altered DNA replication timing as an independent functional test. Eukaryotic genomes replicate via the synchronous firing of clusters of origins, which together produce multi-replicon domains, each of which completes replication in a short (45–60 min) burst during the S phase of the cell cycle39,40. Genome-wide profiling of replication timing shows that these domains can be replicated at different times during S phase, with adjacent earlier and later replicating domains punctuated by regions of replication timing transition39,40. Consequently, translocations that fuse the domains of early and late replication can result in earlier replication of the late replicating domain and/or delayed replication of the early replicating domain41,42. When mapped to the reference genome, these changes appear as abrupt shifts in replication timing profiles that have the potential to validate breakpoints (Fig. 2e and Supplementary Fig. 9a). Our Hi-C pipeline identified 249 translocations (at the 10-kb or 100-kb resolution) in 10 cell lines with available replication timing. Among them, 75 translocations were associated with an abrupt shift in replication timing. Since an abrupt shift is only expected for translocations between domains that replicate at different times, we classified the genome into regions that are constitutively early (CE) replicating, constitutively late (CL) replicating, and regions that switch replication timing during development (S), using 48 replication timing profiles of noncancerous cell lines and differentiation intermediates43,44 (Supplementary Methods and Supplementary Fig. 9b). Among the 249 translocations detected by Hi-C, 9 were CE to CL fusions and 32 were CE to CE or CL to CL fusions. As expected, an abrupt shift in timing was identified in CE to CL with a much higher frequency (~67%) than in CE to CE or CL to CL fusions (~13%) (Supplementary Fig. 9c). Translocations between CE and CE were observed with a frequency three times higher than expected by chance (Supplementary Fig. 9c), which is consistent with previous reports linking chromosomal breakpoints to early replication and higher transcriptional activity45,46. Overall, replication timing can provide functional validation of a specific class of translocation events that fuse regions that are replicated at different times in S phase.
Cross-platform comparison and integration of SV detection
To systematically evaluate the performance of different platforms, we compared the SVs predicted by Hi-C, optical mapping, WGS, fusion transcripts, karyotyping36,45,48,49,50,51,52,53,54, and paired-end tag sequencing (PET-seq)55,56 (Supplementary Fig. 10 and Supplementary Table 8). We defined rearrangements detected by at least two different methods as high-confidence SVs. To approximate sensitivity and specificity, we defined the contribution of a method as the fraction of high-confidence SVs that are detected by this method; the overlap rate refers to the proportion of SVs from one method that overlap with high-confidence SVs.
Overall, we observed that 20% of all interchromosomal translocations were identified by at least two platforms (Supplementary Fig. 11a–b). Compared with previously known karyotypes in each lineage, many of the observed translocations are novel. For example, to our knowledge, 14 out of 26 translocations in T47D cells found in this study have not been reported before (Supplementary Fig. 11c). We selected eight of them for further validation, and all of them were confirmed by PCR (Supplementary Table 7). Hi-C is a method with significant contribution and a high overlap rate (48 and 66%), and with better performance for interchromosomal translocations (53 and 66%) than intrachromosomal SVs (43 and 71%) (Supplementary Table 9 and Supplementary Fig. 12). Integration of Hi-C, optical mapping, and WGS increases the overall contribution to 90% (their individual contributions are 48, 40, and 64%, respectively). Karyotyping has a high overlap rate with the high-confidence calls for all kinds of large SVs (88%) and relatively good contribution for interchromosomal translocations (56%).
Next, we merged the results across different platforms in the same cell line into a final high-confidence SV list and refined the breakpoints using the highest resolution available (Supplementary Table 8 and Supplementary Fig. 12c). More importantly, we resolved the SV type for a subset of unclassified large intrachromosomal rearrangements detected by WGS and optical mapping. For example, Irys reported 24 unclassified intrachromosomal rearrangements (≥5 Mb) in T47D cells. By comparing this with Hi-C or WGS data, we were able to identify the SV types for 9 of them (37.5%).
We also identified thousands of gains or losses of genetic material by optical mapping and WGS in each cancer cell line. Optical mapping detects fewer but larger deletions than WGS (Supplementary Table 10). In T47D cells, WGS detected 2,943 deletions with a median size of 552 bp, while Irys reported 1,128 deletions with a median size of 1,335 bp (Fig. 3a,b). Eighty-five percent (2,495 of 2,945) of WGS-detected deletions are missed by Irys. Among them, 78% are <1 kb. These are likely to be missed by optical mapping because its resolution is limited by the minimum distance between two nicking sites; 3% of the deletions predicted by Irys overlap with multiple smaller WGS deletions, and in those cases, the summed size of these WGS deletions is close to the Irys-detected deletion (Supplementary Fig. 13a–c and Supplementary Table 11). Fifty-eight percent of the Irys-detected deletions are not captured by WGS. We tested a subset of deletions detected by Irys, and 87.5% (14 of 16) were validated by PCR (Supplementary Table 7). Further, optical mapping can identify deletions within repetitive regions where WGS reads are not mapped (Fig. 3c) and in regions with lower mappability around the breakpoints (Supplementary Fig. 13d). We detected many megabase-scale deletions in the cancer cell lines. In contrast, the largest deletion we found in the GM12878 cells was a 700-kb event associated with potential V(D)J recombination (Supplementary Fig. 14). We found that WGS, Irys, and Hi-C can detect different sets of interchromosomal and large-scale rearrangements (Supplementary Fig. 15). Besides mappability, we observed that both Hi-C and Irys are particularly powerful at detecting rearrangements involved with unalignable junctions (Supplementary Fig. 16a,b), which could come from a third chromosome that is too short to be recognized, the non-templated addition of bases to the genome, or exogenous DNA sequences, such as those from viruses.
a, Overlap of deletions in T47D cells detected by optical mapping and WGS. b, Size distribution of deletions detected by optical mapping (n = 1,108) and WGS (n = 2964, P = 1.33 × 10−36, two-sided Wilcoxon rank-sum test). For the boxplots, the box represents the interquartile range (IQR), and the whiskers extend to 1.5 times the IQR or to the maximum/minimum if <1.5 × IQR. c, Optical mapping detects a 6-kb deletion within chrX:96,041,289-96,072,340 that is missed by WGS. d, Reconstruction of the complex local structure of a derivative chromosome in K562 cells through the integration of optical mapping, Hi-C, and WGS. The rearranged allele consists of 5 regions: A (chr13:80.5–80.8 Mb); B (chr13:89.7–93.3 Mb); C (chr13:107.8–108 Mb); D (chr9:130.7–131.3 Mb), and an unalignable region. Further, segment B consists of three smaller regions (B1, B2, and B3 in the figure). We reconstructed a global view of the genome structures in this region by stitching several optical mapping contigs together (middle panel). Each junction of the optical mapping genome map can be validated with Hi-C data. WGS data can provide base-pair resolution breakpoints for specific breakpoint junctions. Each line in the WGS panel represents a read pair. WGS reads that support the breakpoint site are marked as purple (forward strand) and red (reverse strand). e, Strategy of using Hi-C to reconstruct SVs. Hi-C shows increased interaction frequency if two translocated regions are directly joined (→) or if they are not immediately linked adjacent (*), but are linked to the same rearranged allele.
In summary, we found that an integrative approach combining complementary methods is essential to gain a more comprehensive understanding of structural variation in cancer genomes (Table 2). An example is in shown in Fig. 3d, where we used optical mapping to thread the putative local structure, the WGS calls to pinpoint breakpoints, and the Hi-C data to validate the linkage of several adjacent rearrangements on the same allele (Fig. 3e, Supplementary Fig. 17).
Better estimation of gaps in the human genome
We noticed that optical mapping can be used to better estimate the size of gap regions. We detected a number of deletions in multiple samples, including GM12878, when we used the hg19 reference genome, but these deletions disappeared when we processed data with a more recent version of the reference genome (GRCh38). Further investigation shows that many such “deletions” identified in the hg19 consist of gaps in the reference genome and that the size of these gaps has been corrected in the GRCh38 build. The corrected size in GRCh38 is very similar to our predictions (Supplementary Table 12). However, we noticed that there remain several such “deletions” over gap regions even in the GRCh38 build, indicating that either these gap sizes can be further refined or they represent polymorphisms in the population. We compared our results with two recent studies that also re-estimated the genomic gaps in the GRCh38 reference genome57,58. While overall our data show consistency with previous results (Supplementary Table 13), we observed differences due to possible population polymorphisms, including a gap region where we reported a range of sizes from 889 bp to 1,535 bp across 9 different individual cell lines. (The estimation is 1,299 bp by Pendleton et al.57 and 705 bp from Seo et al.58, respectively.)
Functional consequences of SVs in cancer genomes
To investigate the functional consequences of SVs, we first analyzed the RNA-seq data of 11 cancer cell lines to identify fused gene transcripts. We detected many RNA-seq read pairs whose two ends are mapped to different chromosomes, crossing the translocation breakpoints identified in this study (Supplementary Table 14). We also discovered many novel fusion transcripts involving bona fide oncogenes, such as EVI1-CFAP70 in the T47D cells. Whether these gene fusion events contribute to oncogenic potential remains to be further investigated.
CNAs represent another class of genetic variation in cancer. We profiled the CNAs in the T47D breast cancer cell line and compared them with the WGS data of 560 breast cancer patients16. Eight out of the top ten frequently mutated oncogenes in patients were also amplified in the T47D cancer cells; tumor suppressor genes such as ATRX and CDKN1B displayed loss of copies (Fig. 4a), suggesting that the T47D cells reflect the CNA landscape in breast cancer. We further compared the RNA-seq data in T47D and human mammary epithelial cells (HMECs) and found that loss of heterozygosity (LOH) and homozygous deletions lead to significantly reduced gene expression, which was also observed in other cancer cell lines (Supplementary Fig. 18a–d). We found exon deletions in 25 COSMIC (Catalogue of Somatic Mutations in Cancer; see URLs) tumor-related genes, and the majority (76%) showed decreased transcription (Supplementary Fig. 18e). We noticed widespread amplification of known oncogenes (such as MYC) and loss of cell cycle checkpoint genes (such as CDKN2A and CDKN2B, Supplementary Fig. 19). We found over 100 highly amplified (≥5 copies) or deleted genes in cancer cells that were not reported in COSMIC, suggesting potential roles in cancer (Supplementary Fig. 20).
a, Copy number changes in T47D cells of RefSeq genes, sorted by copy number. Genes that are frequently mutated in breast cancer are labeled if they show amplification (red dots) or deletion (yellow dots). The right panel of this figure displays the density plot of gene copy numbers. b, An approximately 3.4-kb deletion (chr3:179,546,826–179,550,207) in T47D overlaps an HMEC-specific enhancer. Hi-C data from HMEC indicates that there is an interaction between the deleted enhancer and the promoter of gene GNB4. This enhancer–promoter linkage is also reported in GM12878 cells by the Capture Hi-C data. According to the WGS data, the local region is amplified and has six copies in the T47D cells, but the enhancer is deleted in five of the six copies. c, Compared with HMEC, all the genes in this region in T47D are upregulated potentially due to the local amplification, except for GNB4, whose expression is reduced by ~50%. d, Functional pathway analysis of deleted enhancers (n = 1,859) by the Genome Regulatory Architecture tools (P value from two-sided binomial test). FDR, false discovery rate. ESR1, estrogen receptor 1. e, Genes with deleted enhancers show reduced expression levels (two-sided Wilcoxon rank-sum test). Genes with exon deletions or copy number loss are excluded; 534 genes are linked by Capture Hi-C data to at least one deleted enhancer (green) and 10,677 genes are linked to enhancers that show no deletions (gray). For the boxplots, the box represents the interquartile range (IQR), and the whiskers extend to 1.5 × IQR or to the maximum/minimum if <1.5 × IQR.
Deletions in cancer and normal cell lines differed in their likelihood of disrupting repetitive or functional elements. GM12878 cells are more enriched for deletions in repetitive elements when compared with cancer cell lines (70 versus 50%; the expected value based on genomic background is 50%) (Supplementary Table 15). Interestingly, deletions of genes and enhancers are depleted in GM12878 cells relative to the genomic background (12 versus 60, empirical P value <0.001, Supplementary Table 16), while the cancer cell lines do not show such depletion of enhancer deletions (Supplementary Fig. 21a).
To identify deletions specific to cancer genomes, we compared the observed deletions with the Database of Genomic Variants (see URLs), which compiles known polymorphic SVs identified by previous studies. Ninety-five percent of the deletions identified in GM12878 cells have been previously reported, suggesting they are polymorphisms in the population. The fraction of polymorphic deletions in cancer cells is lower at 90% (Supplementary Fig. 18f and Supplementary Fig. 22a), likely due to the presence of somatic mutations. In total, cancer cells suffer a greater loss of genetic material compared with normal cells (Supplementary Fig. 22b). Further analysis shows that polymorphic deletions are enriched for repetitive elements (70 versus 50% genomic background) and depleted of exons (1.5 versus 4% genomic background) (Supplementary Fig. 22c–d). In the six cancer cell lines where we can find control cells with enhancer annotations, we found that the polymorphic deletions are resistant to enhancer loss (empirical P < 0.005 in all cell lines; Supplementary Fig. 21b). In contrast, the novel deletions are not enriched in repeats or depleted of enhancers or exons (Supplementary Figs. 21c and 22). Instead, they are enriched in COSMIC tumor-related genes (Supplementary Fig. 18f)59, suggesting that a subset of the deletions are potentially pathogenic. We confirmed that copy number changes detected by optical mapping and WGS are highly consistent (Supplementary Fig. 23).
Next, we investigated whether SVs can influence the expression of cancer-related genes by disrupting distal regulatory elements. For this analysis, we focused on the comparison between T47D breast cancer cells and HMECs. We predicted enhancers in HMECs using H3K27ac chromatin immunoprecipitation sequencing data from the ENCODE consortium (see URLs) and compared the enhancers with deleted regions in T47D to identify potential deleted enhancers in cancer cells (Supplementary Table 17). We show an example in Fig. 4b, where a 3.4-kb deletion downstream of the GNB4 gene overlaps with a breast tissue-specific enhancer. This region has six copies due to genomic amplification, five of which carry this deletion; only one copy of the enhancer remains undisrupted. Evidence by Hi-C in HMECs and Capture Hi-C data60 suggests that GNB4 is potentially regulated by this enhancer. More importantly, it is the only gene in this region with decreased expression; the expression of the rest of the genes in this region are highly upregulated, possibly due to the increased copy number (Fig. 4c). Further, we found that globally, deleted enhancers are located near genes involved in breast cancer-relevant pathways (Fig. 4d) and genes linked to these deleted enhancers show a reduced level of expression (Fig. 4e). Overall, these results suggest that deletions in cancer genomes may frequently affect enhancers and potentially contribute to oncogenesis.
The impact of structural variations on 3D genome organization
Genetic mutations can disrupt TADs and create “neo-TADs” that lead to misregulated gene expression in developmental disorders61,62. Several groups have also shown that alterations that affect TAD boundaries or transcriptional repressor CTCF binding sites at specific loci can create new chromatin structural domains leading to misregulation of nearby oncogenes through “enhancer hijacking”61,64,65. However, the extent to which SVs alter 3D genome structures genome-wide in cancer cells remains unclear.
Having identified SVs in 20 cancer cell lines with Hi-C data, we systematically investigated the consequences of structural variation on TAD structure. We observed that neo-TADs are frequently formed as the result of large-scale genomic rearrangements in cancer cells. An example is shown in Fig. 5a, where the fusion between chromosomes 9 and 18 forms a neo-TAD in PANC-1 cells. Further, we found that many neo-TADs induced by SVs in cancer cells contain known cancer driver genes, such as ERBB2, ETV1, ETV4, MYC and TERT (Supplementary Fig. 24). To address whether neo-TAD formation is a general consequence of SV rearrangements in cancer genomes, we performed an aggregate analysis of all breakpoint-crossing Hi-C signals in each cell line. As shown in Fig. 5b, we observed that interchromosomal Hi-C signals form a sharp triangular shape (dashed line), suggesting the formation of a fusion-TAD as a result of the rearrangement (details in Supplementary Methods). This pattern was not observed when we performed the same analysis using shuffled TADs with randomized boundary positions (right panel in Fig. 5b). These results indicate that structural variations in cancers can rewire TAD structure and lead to TAD fusion and altered regulatory environments (Fig. 5c).
a, Fusion-TAD formation as a result of a translocation in PANC-1 cells. The left box shows the rearranged region on chromosome 9, while the right box shows the rearranged region on chromosome 18. The breakpoint fusion lies in the middle. The triangular Hi-C heat maps show intrachromosomal interactions. The diamond heat map shows the breakpoint-crossing Hi-C signal, indicating the presence of a TAD fusion. b, Aggregate analysis of TAD fusions. Breakpoint-crossing Hi-C signals were averaged and centered on bins between the nearest TAD boundaries (left) or shuffled TAD boundaries (right, randomization performed 1,000 times). The dashed lines show expected neo-TAD borders based on the intersections of the nearest breakpoint-proximal TAD boundaries. c, Model for neo-TAD formation. TADs are rearranged due to breaks and fusions, juxtaposing regulatory sequences with nontarget genes. d, Violin plots showing the distribution of allelic expression bias for genes within rearranged (n = 1,004) or non-rearranged (n = 74,184) TADs. The vertical bars represent the median (the P value is from a two-sided Wilcoxon rank-sum test). e, RNA-seq for MYCN/N-Myc (green) and MYC/c-Myc (red) in neuroblastoma cell lines. The cell lines with TAD fusions at the MYC locus show high levels of MYC expression (marked in red); the cell line that lacks a TAD fusion at the MYC locus lacks MYC expression (yellow). RPKM, reads per kilobase of exon per million reads mapped. f, Hi-C data from SK-N-SH cells showing a TAD fusion at the MYC locus. g, Hi-C data in SK-N-AS cells showing a TAD fusion at the MYC locus.
Next, we investigated the impact of neo-TADs on gene expression. Across eight cancer cell lines, we observed that genes within TADs containing a rearrangement show greater allelic bias than genes within non-rearranged TADs, suggesting that at least a subset of these events likely lead to altered gene expression in cis (Fig. 5d). We examined the Hi-C data in three neuroblastoma cell lines and compared MYC expression. Among them, SK-N-DZ has high MYCN/N-myc expression, and the other two lines (SK-N-SH and SK-N-AS) have high MYC/c-Myc expression (Fig. 5e). Remarkably, in the two neuroblastoma cell lines that had high MYC expression (SK-N-AS and SK-N-SH), we identified the presence of translocations in the vicinity of the MYC gene. Copy number segmentation from the Cancer Cell Line Encyclopedia (see URLs) indicates that there is no MYC amplification in these two cell lines. Instead, we observed the formation of neo-TADs that encompass the MYC gene in both cases (Fig. 5f,g), suggesting that the formation of neo-TADs may be involved in MYC activation. Determining whether any individual neo-TAD represents a recurrent alteration in a given cancer cell type, or how neo-TADs may ultimately contribute to oncogenesis, remains to be elucidated. However, our analysis suggests that creation of neo-TADs is a common consequence of rearrangements in cancer genomes.
Discussion
Detecting SVs in cancer genomes remains a challenge for geneticists and cancer biologists. Here we developed an algorithm that, for the first time, can use Hi-C data to identify a full range of SVs in cancer cells genome-wide. Our algorithm shows high accuracy for detecting interchromosomal translocations and large intrachromosomal rearrangements, even with as little as approximately 1 × genome coverage. Currently, our approach has limited power in detecting alterations <1 Mb in size. On the other hand, we have demonstrated that optical mapping excels at detecting complex SVs and resolving local genome structure, although it cannot detect small deletions and insertions (<1 kb). WGS has the highest resolution in detecting structural variation but is less successful in detecting SVs in poorly mappable regions of the genome or in resolving complex SVs. Ultimately, only an integrative approach that employs complementary technologies can give the most comprehensive view of the cancer genome.
In examining regions affected by SVs, we identified extensive deletions of distal enhancers, which are located in proximity to genes known to be mutated in cancer and important for pathways in cancer biology. To what extent such distal noncoding mutations are recurrent in cancer genomes remains unclear, but this represents an important, less explored aspect of cancer genomics. By analyzing the 3D genome structure surrounding the SVs, we observed frequent creation of neo-TADs as a result of genomic rearrangements in cancer genomes. We have developed a Web-based tool for users to visualize and examine such neo-TADs (available at the 3D Genome Browser; see URLs). There has been ample evidence that the juxtaposition of active regulatory sequences to known oncogenes can contribute to tumorigenesis. Our results indicate that at least part of this effect may result from the creation of novel structural domains in cancer genomes. Whether all SVs generate fusion-TADs, and the extent to which TAD fusion events are recurrent and act as driver mutations in cancer genomes will be an important question for future studies to address.
URLs
LUMPY, https://github.com/arq5x/lumpy-sv. Delly, https://github.com/dellytools/delly. Control-FREEC, http://boevalab.com/FREEC/. COSMIC, https://cancer.sanger.ac.uk/cosmic. Database of Genomic Variants, http://dgv.tcag.ca/dgv/app/home. ENCODE, http://encodeproject.org/. 3D Genome Browser, http://3dgenome.org/. Cancer Cell Line Encyclopedia, https://portals.broadinstitute.org/ccle. 1000 Genomes Consortium GRCh38, http://www.internationalgenome.org/data-portal/search?q=GRCh38. SpeedSeq framework, https://github.com/hall-lab/speedseq. Samblaster, https://anaconda.org/bioconda/samblaster. Replication Domain genome browser and analysis tool, www.replicationdomain.com. European Nucleotide Archive, https://www.ebi.ac.uk/ena. Sequence Read Archive, https://www.ncbi.nlm.nih.gov/sra. Basic Local Alignment Search Tool (BLAST), https://blast.ncbi.nlm.nih.gov/Blast.cgi. Genomic Regions Enrichment of Annotations Tool, http://great.stanford.edu/public/html/splash.php. Github code for SV identification, https://github.com/dixonlab/hic_breakfinder.
Methods
Cell culture
K562 cells (ATCC) were cultured in IMDM supplemented with 10% fetal bovine serum (FBS) and antibiotics. T47D cells (ATCC), NCI-H460 cells (ATCC), A549 cells (ATCC), LNCaP (ATCC), and GM12878 cells (Coriell Institute) were cultured in RPMI-1640 medium (Corning, 10-043-CV) supplemented with 10% FBS and antibiotics, or 15% FBS and antibiotics (GM12878). Caki2 cells (ATCC), G-401 cells (ATCC) were cultured in McCoy’s 5A (Modified) Medium supplemented with 10% FBS and antibiotics. PANC-1 cells (ATCC) were cultured in DMEM supplemented with 10% FBS and antibiotics. SK-N-MC (ATCC), RPMI-7951 (ATCC) cells were cultured in Eagle’s Minimum Essential Medium supplemented with 10% FBS and antibiotics. SK-N-AS cells (ATCC) were cultured in DMEM supplemented with 10% FBS, 0.1 mM MEM Non-Essential Amino Acids solution (Gibco) and antibiotics. All cell lines cultured as part of the ENCODE data generation (A549, Caki2, G-401, LNCaP, NCI-H460, PANC-1, RPMI-7951, SJCRH30 (ATCC), SK-MEL-5 (ATCC), SK-N-DZ (ATCC), SK-N-MC, and T47D) were cultured using standardized protocols, the details of which can be found on the ENCODE consortium website (see URLs).
Optical mapping experiments
Ten million cells of T47D, Caki2, K562, SK-N-MC, A549, NCI-H460, PANC-1, and LNCaP were pelleted and then washed three times with PBS buffer solution. Cells equivalent to 600 ng of DNA were embedded in 2% agarose (Bio-Rad Laboratories) and solidified at 4 °C for 45 min. Cells within plugs were lysed in 2 ml cell lysis buffer (Bionano Genomics) containing 167 µl proteinase K (Qiagen) for 48 h, and washed twice with Tris-EDTA buffer solution, pH 8 (Tris-EDTA) for 15 min per wash. DNA plugs were purified with 2 ml 5% RNAase (Qiagen) for 2 h, washed in Tris-EDTA for 15 min × 6 times, melted and equilibrated on 43 °C for 45 min with 2 µl of GELase (Epicentre). DNA was transferred onto a membrane floating in Tris-EDTA and concentrated by dialysis for 135 min. DNA was then equilibrated at room temperature overnight; 900 ng DNA was digested by 30 U nicking enzyme BspQI (New England Biolabs) in 1 × buffer III (Bionano Genomics), 37 °C for 4 h, and labeled with 1 × labeling mix (Bionano Genomics) and 15 U Taq DNA polymerase (New England Biolabs) in 1 × labeling buffer (Bionano Genomics) at 72 °C for 60 min. Nick-labeled DNA was repaired in 1 × repair mix (Bionano Genomics), 1 × Thermo polymerase buffer (New England Biolabs), 50 µM β-nicotinamide adenine dinucleotide (New England Biolabs), and 3 µl 120 U Taq DNA ligase (New England Biolabs) at 37 °C for 30 min. DNA staining was finally performed with the final solution containing 1 × flow buffer, 1 × dithiothreitol (Bionano Genomics), and 3 µl DNA stain (Bionano Genomics), at room temperature overnight. On average, each sample underwent 7 rounds of data collection on the Irys platform (Bionano Genomics) to reach 100 × reference coverage. For each round, 160 ng prepared DNA was loaded onto an Irys chip that contained two flow cells, and each round contained 30 cycles of data collection.
Hi-C experiments and sequence read alignment
Hi-C in K562 and SK-N-AS cells was performed using the in situ Hi-C protocol21 from five million cells using the MboI enzyme. Hi-C experiments in all ENCODE cells lines were performed by following the original Hi-C protocol using the type-2 restriction enzyme HindIII66. Hi-C experiments were performed as biological replicates to ensure experimental reproducibility. Hi-C libraries were sequenced using the HiSeq 2000 and HiSeq 2500 sequencing systems (Illumina) and processed to FASTQ format files using standard processing pipelines. Read pairs were aligned independently using the BWA-MEM algorithm to a custom GRCh38 genome assembly. The base for this assembly is available through the 1000 genomes consortium (see URLs) and contains “decoy” sequences representing common viral sequences and contigs assembled in certain individual genome assemblies but not found in the current reference. For our purposes, we also removed any alternate haplotype sequences from the reference. After initial alignment, individual reads were paired using a custom in-house pipeline, and PCR duplicate reads were removed using Picard Tools (https://broadinstitute.github.io/picard/). The aligned sequences were then processed into raw Hi-C matrices using multiple bin sizes (1 Mb, 100 kb, 40 kb, 10 kb). For the K562 and SK-N-AS libraries, these were also processed into 1-kb matrices. This was not done for the ENCODE cell line data because HindIII is expected to cut approximately every 4 kb in the genome, such that analyzing data with bin sizes below 4 kb yields highly variable interaction matrices. We then computed the raw one-dimensional coverage of bins and removed any bins in the bottom 2.5%. Hi-C matrices were then normalized using iterative correction; we specifically retained the vector of intrinsic biases, q, for use in downstream breakpoint calling. Thus, the normalized interaction frequency ni,j between any two bins i and j are given by the following equation:
\(n_{i,j} = \frac{{o_{i,j}}}{{q_i \times q_j}}\)where oi,j is the observed number of reads between bins i and j, and qi and qj are the biases of bins i and j, respectively.
For Hi-C data analyzed using 1-Mb matrices, the A/B compartment patterns are a prominent feature of the data that complicate downstream breakpoint finding. To account for these features, we performed further processing of the 1-Mb matrices. Specifically, we renormalized the raw 1-Mb matrices using iterative correction but with all intrachromosomal interactions set to zero. We then performed principal component analysis on this balanced matrix. Specifically, using R, we computed the covariance matrix of our initial normalized matrix and then extracted the first eigenvector. We then computed a new matrix, D, representing the fraction of the original matrix derived from the first principal component by the following equation: \(B = Xvv^T\), where X is the original normalized matrix, v is the first eigenvector, and vT is the transposition of the first eigenvector. Each element bi,j of B represents the additive increase or decrease in interaction frequency due to A/B compartment patterns between bins i and j, and is used in modeling the interaction frequencies for breakpoint identification. For further details regarding the algorithms for SV detection using Hi-C data, please see the Supplementary Methods.
SV detection and filtration from WGS
SV detection
SVs were detected by three independent pipelines. In the first pipeline, paired-end sequencing reads were first aligned using the BWA-MEM algorithm (v0.7.15-r1140) to a GRCh38 human reference genome (version GCA000001405.015) with alternate haplotypes removed. Duplicate reads were removed using Picard Tools. Reads with a mapping quality of at least 20 were retained for SV detection. SV calls were generated from this mapped data using Delly (v0.7.7) with default parameters (-q 20). Delly detects deletions, inversions, tandem duplications, insertions, and interchromosomal translocations.
In the second pipeline, paired-end reads were processed using the SpeedSeq framework (see URLs). Paired-end reads were aligned to the GRCh38 reference genome using the BWA-MEM algorithm in the same manner as the first pipeline. Duplicated reads were removed by “samblaster” (v0.1.24; see URLs). Discordant and split reads were extracted by samblaster for SV detection. SV calls were generated using LUMPY (v0.2.13) with default parameters (speedseq sv -g -t 64 -x). LUMPY reports SVs as deletions, inversions, duplications, interchromosomal translocations, and unresolved break ends. In both pipelines, telomeric, centromeric, and 12 heterochromatic regions were masked for SV detection using the blacklisted regions provided by the Delly software.
Copy number profiles were generated using Control-FREEC (v11.0; see URLs). For all cell lines, we used a set of common parameters (ploidy = 2 for normal cells NA12878, pseudodiploid cells SK-N-MC, and hypotriploid cells T47D, A549, LNCaP, NCI-H460 and Caki2; ploidy = 3 for triploid cells K562 and hypertriploid cells PANC-1; breakPointThreshold = 0.8, coefficientOfVariation = 0.062, mateOrientation = FR). For A549, Caki2, LNCaP, NCI-H460, and PANC-1, sex was set to “XY”; for K562, NA12878, SK-N-MC, and T47D, sex was set to “XX”. The predicted copy number for each 50-kb bin was used for making genome-profile plots generated with the Circos software available from http://circos.ca/. Regions with copy loss (copy number = 0 or 1) that were not captured by SV detection using Delly or LUMPY (by excluding those that reciprocally overlapped by at least 50% with the deletions called by Delly and LUMPY) were included in the set of detected deletions.
SV filtration
To reduce false positive calls, the following filtration steps were applied for the Delly and LUMPY SV calls. First, we required all SV calls to be supported by at least three split reads or three spanning paired-end reads. Insertions or deletions <50 bp were removed, as were SV calls that mapped to chromosome Y or to the mitochondrial genome. SV calls from Delly and LUMPY were then merged, and only SVs that were identified by both methods were retained. We used separate criteria to call SVs overlapping between the two methods depending on the type of SV. For deletions, calls were merged between the two pipelines if they had a reciprocal overlap (RO) ≥50%. We used the coordinates provided by LUMPY for this merged deletion set. For inversions, calls identified by both LUMPY and Delly were merged if they had an RO ≥0.9. The final merged coordinates were based on the coordinates from the LUMPY calls. Translocations were merged between the two pipelines if the paired break ends mapped within ±50 bp of each other and if the strand of the break ends matched. The final coordinates were based on the calls from LUMPY. Regions annotated as insertions were identified by Delly alone, since LUMPY does not annotate SVs as insertions. No specific filtration for insertion was applied.
Additional filtration was applied to specific types of SVs. For deletions, we removed deletions that had at least 50% reciprocal overlap (RO ≥50%) with known gap regions (±50 bp), or at least a 1 bp overlap with centromere regions (±1 kb). Recurrent deletions that were larger than 1 Mb and presented in more than one cell line with an RO ≥99.9% were removed. Large deletions (≥100 kb) that did not show consistent decrease of read depth compared with adjacent regions were also removed (less than one difference of read depth between deletions and flanking 10-kb regions). For inversions, recurrent inversions that were longer than 100 kb and were present in more than one cell line (defined by RO ≥99.9%) were removed. For translocations, recurrent translocations that were present in more than one cell line (defined by both break ends being within ± 50 bp) were filtered out.
We required a minimal number of supporting split reads and paired-end reads (SR + PE) for translocation calls that we varied according to the sequencing depth and the ploidy of the WGS sample. (Cells with polyploidy can harbor an SV in only one copy of the DNA; thus, the SV is only present in a small fraction WGS reads.) Due to high sequencing coverage (~80×) in the LNCaP sample, we only kept translocations with at least 15 SRs (PE + SR). For the GM12878 cells (coverage of 50×), since they are diploid, we used a more stringent filter of 20 SRs, with at least two being split reads. For all other cell lines, which had similar read depth and ploidy, we required at least five SRs to call a translocation. We further compiled a list of high-coverage regions (coverage >500×) in NA12878 cells, which are largely characterized by repetitive genomic elements. In our initial analysis, we observed that such regions have high rates of translocation calls. However, given their extreme outlier coverage and association with repetitive elements, these are most likely simply anomalous alignments. We filtered out translocations whose breakpoint ends were located in those regions. In addition, for unclassified intrachromosomal rearrangements called by LUMPY, we removed calls with a quality score <100. Finally, for tandem duplications, we required ten SRs for LNCaP, five for GM12878, and three for all other samples.
Detection of SVs based on optical mapping
Cell line- or sample-specific genomic maps were generated through de novo assembly of DNA optical reads using RefAligner 6119 and pipeline 6498. We required that DNA reads be no shorter than 150 kb with at least 9 labels per molecule, and the signal-to-noise ratio no less than 2.75, while the maximum backbone intensity should be 0.6. The assembly pipeline was applied with the following parameters: iterations = 5; initial assembly P value threshold = 1 × 10−11; extension and refinement P value threshold = 1 × 10−11. De novo assembly noise were specifically false positive density/100 kb = 1.0; false negative rate = 0.1; siteSD = 0.15; scalingSD = 0; relativeSD = 0.03; resolutionSD = 0.25.
SV detection was performed after the completion of de novo assembly by comparing the assembled contigs to the GRCh38 reference genome GRCh38 using the built-in module “runSV”. All centromere regions were skipped during SV identification. Deletions, insertions, and inversions were detected with the default settings using a P value threshold of 1 × 10−12. In the default output, any intrachromosomal SVs larger than 5 Mb were defined as “unclassified” intrachromosomal rearrangements. Unclassified intrachromosomal rearrangements and interchromosomal translocations were detected using a less stringent P value threshold of 1 × 10−8.
For further details on the filtration and classification of SVs detected by optical mapping, please see the Supplementary Information File.
Profiling of gene copies using optical mapping
To identify genes that had undergone CNA, we compared copy number profiles from optical mapping in the four primary normal tissues and eight cancer cell lines with the RefSeq gene list from the NCBI Reference Sequence Database (https://www.ncbi.nlm.nih.gov/refseq/). The longest isoform was used to characterize copy number changes. For each gene, the average copy number profiles of each 50-kb bin spanned by the gene was considered as the copy number of that gene. The copy number variations (CNVs) of genes were also profiled using WGS normalized coverage (Control-FREEC) in T47D and Caki2 for differential gene expression analysis.
Re-prediction of gap sizes
To gain a list of candidate unresolved gap regions, recurrent deletions detected by optical mapping at least twice in cancer cells lines and at least once in normal cells were collected from 12 samples, including 8 cancer cell lines (T47D, Caki2, K562, A549, NCI-H460, PANC-1, LNCaP, SK-N-MC) and 4 normal cells (GM12878, 3078entB, 3045entB, and 3391entB). Recurrent deletions were then intersected with hg19 gaps using “bedtools”. Only gaps where at least 80% of the gap overlapped with a deletion and the gap accounted for at least 30% of the deletion were retained for gap size re-estimation. When using hg19 as the reference genome, the gap size was predicted by subtracting the deletion size from the gap size in hg19. To evaluate the predictions, the gap regions were lifted over to GRCh38; the sizes of the same regions in GRCh38 were compared with our prediction and the size in hg19. Some gaps ultimately have a negative value, meaning that the size of the deletion is shorter than the annotated gap in the reference genome, potentially because of the variation across populations.
To predict the size of unresolved gaps in GRCh38, we repeated our analysis of deletions overlapping gap regions using GRCh38 as the reference genome as described previously. In some cases, the re-estimated size of the same gap could vary among different cell lines, and the degree of variation was relatively small with respect to the overall change of the perceived scale of gap size. Therefore, we report the median, maximum, and minimal gap size of each gap from our estimation, since this variation can represent polymorphisms of gap sizes in the population. We then annotated which genes were spanned by those adjusted gaps and could be affected by intersecting re-estimated gaps with the gene list in GRCh38. We further compared our gap size predictions in GRCh38 with the results from previous publications57,58.
Genome-wide DNA replication timing
Genome-wide replication timing was measured in A549, Caki2, G401, LNCaP, NCI-H460, SK-N-MC and T47D using the repli-seq method68. Briefly, asynchronously cycling cells were pulse-labeled with the nucleotide analog 5-bromo-2′-deoxyuridine (BrdU; Sigma-Aldrich, B5002). The cells were then sorted into early and late S phase fractions on the basis of DNA content using flow cytometry. BrdU-labeled DNA from each fraction was immunoprecipitated, amplified, and sequenced using the HiSeq 2500 Sequencing System (Illumina). The replication timing was then measured as the log2 ratio of early over late reads in 5-kb bins. For the K562, MCF7, and SK-N-SH cell lines, raw data for six-fraction repli-seq were downloaded from the ENCODE portal (see URLs). The data were transformed to match the early/late repli-seq by combining G1, S1, and S2 fractions to represent the early S phase, and S3, S4, and G2 fractions to represent the late S phase. Smoothed replication timing profiles around the breakpoints were produced by loess smoothing replication timing data separately for the upstream and the downstream segments from the breakpoints predicted by Hi-C (Figs. 1b, 2e).
Classification of the human genome into constitutive and switching regions
Forty-eight human replication timing data sets (ENCODE, Replication Domain; see URLs) were used for the annotation of the human genome into constitutive and switching regions. The data sets were windowed into 50-kb bins. Then, the following criteria were used for the annotation. A threshold above 0.15 was used to identify early replicating bins; below −0.15 was used to identify a late replicating bin for each data set. If a bin was early in two or more cell types and late in two or more cell types, those bins were classified as “switching” (S). The remaining bins were then evaluated as being either CE, CL, or left unclassified (N/A). If a bin was early in at least 46 out of 48 cell types, it was classified as CE. If a bin was late in at least 46 out of 48 cell types, it was classified as CL.
Quantifying abrupt shifts in replication timing
Genome-wide replication timing profiles in cancer genomes show several abrupt shifts in replication timing associated with translocations. We sought to quantify the frequency of these abrupt shifts. To this end, we made a pipeline to detect abrupt shifts next to translocations identified by Hi-C. For each predicted translocation, unsmoothed replication timing data in 5-kb bins from ±200 kb of the breakpoint were used to scan for abrupt shifts. A span of ±200 kb was chosen because the resolution of Hi-C translocation calls started at 100 kb. Then, for every 5-kb bin, the difference between the median of the preceding 20 bins and succeeding 20 bins were calculated. Outliers were removed from this metric by a median filter (span = 5). Then, a threshold of 0.6 was used to determine the presence or absence of an abrupt shift. While the threshold was chosen empirically, the results showed the same trend across a wide range of thresholds.
Fusion transcripts
We downloaded paired-end RNA-seq data for 14 cell lines from the ENCODE, European Nucleotide Archive, or Sequence Read Archive databases (Supplementary Table 1). We used three different pipelines (Tophat-Fusion (v2.1.0)69, STAR-Fusion (v1.1.0)70, and EricScript (v0.5.5))71 to identify fusion transcripts. For Tophat-Fusion, paired-end reads were aligned to a GRCh38 reference genome (version GCA000001405.015) to identify fusion events. Tophat-Fusion was run on the following parameters: --no-coverage-search -r 50 --mate-std-dev 80 --max-intron-length 100000 --fusion-min-dist 1000 --fusion-anchor-length 13. Tophat-Fusion outputs a list of potential fusion events, which were then processed by “tophat-fusion-post” to filter out false positives by aligning the sequences flanking fusion junctions against BLAST (Basic Local Alignment Search Tool) databases (see URLs). Fusion events were further filtered by requiring at least three split reads or three spanning read pairs. In STAR-Fusion, a built-in GRCh38 reference genome with GENCODE v26 annotation was used. Fusion transcripts were detected by STAR-Fusion with default parameters. To reduce false positives, fusion events with fusion fragments per million total reads <0.1 were removed. EricScript detects fusion transcripts by aligning the reads to a pre-built reference transcriptome (Ensembl v.84) provided by the authors. Further, candidate fusions are required to be supported by at least three spanning read pairs and three split reads. We also included a fourth set of fusion transcripts from Kljin et al72. The final set of fusion transcripts was obtained by considering the union of fusion calls from the three pipelines and the fourth set of fusion events identified by Kljin et al.
Identification of allelic imbalance in expression
To evaluate the effects of TAD fusion events on altered gene expression in cis, we tested whether TADs containing rearrangements showed different patterns of allele-specific gene expression compared to TADs that lack rearrangements. For each cell line where we had WGS (A549, Caki2, K562, NCI-H460, LNCaP, PANC-1, SK-N-MC, T47D), we aligned RNA-seq data to the genome using STAR (https://github.com/alexdobin/STAR). We then implemented the WASP allele-specific pipeline73 for filtering and realigning reads to identify reads that showed inherent allelic mapping biases. We then computed the number of reads that aligned to each allele at each single-nucleotide variant within an exon of any GENCODE gene using “samtools mpileup”. The number of reads aligning to each allele was normalized by the total number of reads (reads per million (RPM) mapped reads), to account for sequencing depth differences between cell lines. To compute the degree of bias in expression between alleles, we used a simple chi-squared statistic. To account for potential differences in copy number between alleles, the expected value of the chi-squared statistic for each single-nucleotide variant was derived from the observed ratio of coverage between alleles from WGS. Specifically, the expected value for each allele was calculated as the fraction of reads from WGS aligning to that allele multiplied by the sum of the RNA-seq RPM values across both alleles.
Gene ontology analysis of deleted enhancers
To perform ontology analysis of enhancer deletions, the locations of high-confidence deletions in T47D cells was intersected with H3K27ac-defined enhancers in HMEC cells. After removal of duplicates, the loci of deleted enhancers were lifted over from hg38 to hg19 and gene ontology analysis was performed by GREAT v.3.0.0 (Genomic Regions Enrichment of Annotations Tool, http://great.stanford.edu/public/html/) using the hg19 reference as background [76] (GREAT requires the use of the hg19 reference). The association rule was set as “basal plus extension”, with “proximal 5.0 kb upstream”, “1.0 kb downstream”, and “plus distal up to 1000.0 kb”.
Differential gene expression from gene dosage or enhancer deletion
To evaluate the effects of gene dosage and enhancer deletions on gene expression, we evaluated the expression of genes in T47D or Caki2 cell lines where we detected CNA of the gene itself or of linked enhancers. For T47D, we used RNA-seq data from HMEC cells as a normal control, and for Caki2, we used RNA-seq data from primary kidney tissue as a normal control. We downloaded the FASTQ format files of paired-end RNA-seq data from T47D, HMEC, Caki2, and primary kidney from the Sequence Read Archive database or ENCODE. Each sample contained two replicates. The raw reads were aligned, and differential expression analysis was performed using “Tophat” and “cufflinks74. To analyze the impact of gene dosage on expression, we grouped genes into four classes: homozygous deletions (zero copies); genes with LOH (one copy); normal genes (two copies); and amplified genes (≥three copies) according to the CNV profiles from WGS. We calculated the expression (fragments per kilobase of transcript per million (FPKM) mapped reads) fold change of all genes in each category relative to the control sample.
To analyze the impact of enhancer deletions on gene expression, we first filtered genes and removed those with deletions of exons or entire genes to control for the impact of gene dosage on expression. We further filtered the genes and focused only on the 9,672 genes with evidence of expression in HMEC cells (FPKM ≥1). Enhancers were annotated as homozygous deletion or LOH based on WGS coverage, and were examined for linkage to filtered genes from any significant interactions identified by Capture Hi-C in GM12878 cells. The expression fold change between T47D and HMEC cells was then computed for the 530 genes with a copy number loss of linked enhancers and was compared with 9,142 unaffected genes using the Wilcoxon signed-rank test.
TAD fusions
To evaluate the effects of SVs on TAD structure, we analyzed breakpoint-crossing Hi-C signals. Our initial observations identified cases where the nearest TAD boundaries to the breakpoint were being “fused” together to create a new TAD. To evaluate whether such TAD fusion events were generally the case, we analyzed whether the breakpoint-crossing Hi-C signal between the nearest TAD boundaries showed local enrichment, which is characteristic of “normal” TADs.
We began this analysis with a list of breakpoints within each cell type. For each breakpoint, we identified the nearest breakpoint-proximal TAD boundary based on TAD calls from H1 human embryonic stem cells (hESCs). We chose TAD calls from H1 hESCs because we wanted to use TAD calls from an independent, non-rearranged cell type, in case the rearrangement was altering TAD calls within the rearranged cell line. We should note that TAD calls are highly stable between cell types, such that these results are similar regardless of the source of the TAD calls. We then identified the predicted “peak” of the TAD “triangle” by identifying the bin representing the interaction between each of the nearest breakpoint-proximal TAD boundaries. The bin representing the interaction between each of the breakpoint-proximal TAD boundaries was then considered as the center of a sub-matrix. We calculated the average interaction frequency of all bins within the 41 × 41 bin sub-matrix centered on the TAD boundary-interacting bin. Each bin was then normalized to this average interaction frequency, such that the new sub-matrix would represent a fold change above the average value in the sub-matrix. This was then log-transformed (with a pseudocount of 1 added to avoid taking the log0 and to minimize the effects of noisy low-frequency interactions). The reason for normalizing to the mean of the sub-matrix is to account for the differences in interaction frequencies that would be expected due to genomic distance alone. In other words, without normalizing to the central bin, the aggregated Hi-C data would be dominated by short-distance interactions. The log fold change sub-matrix was then averaged for all breakpoints in all cell types, yielding a single aggregate log fold change sub-matrix. For display purposes, this was then exponentiated to represent these values again as a fold change. This process was also applied to a random set of TAD boundaries. Random TAD boundaries analysis was performed by first randomly permuting the TAD boundaries from H1 hESCs, using the following approach: for TADs on chromosomes affected by SVs, we generated a random number between 1 and the size of the chromosome where it was located. This number was then added to the start and end coordinates of every TAD on the chromosome. If the randomly generated TAD was larger than the size of the chromosome, the size of the chromosome in bp was then subtracted. This is done to preserve the observed size and spacing of TADs in the random data set to limit any artifacts or bias of randomization. This set of permuted TADs was then used for the input into the same process already described to evaluate the chromatin interactions across the breakpoints. The only data to be randomized were the positions of TADs; both the SVs and chromatin interaction maps used for the plots were from the true cancer cell lines in this study. This randomization was repeated 1,000 times.
Statistics
We used the Wilcoxon rank-sum test to compare distributions between two groups because this is a nonparametric distribution that does not make underlying assumptions of normality. We also used permutation to calculate empirical P values, which does not make any assumptions on the underlying distribution of the data.
Reporting Summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Code availability
Code for Hi-C-based SV identification can be accessed through GitHub (see URLs). We used publicly available software for WGS SV detection (LUMPY, DELLY, control-FREEC). We used RefAligner 6119 and pipeline 6498 for SV detection from optical mapping experiments. Custom data processing scripts can be made available on request.
Data availability
Hi-C and replication timing data generated in this study have been deposited on the ENCODE portal and can be accessed without restrictions (see URLs). Details of specific accession numbers for each data set can be found in the Supplementary Methods section.
References
Futreal, P. A. et al. A census of human cancer genes. Nat. Rev. Cancer 4, 177–183 (2004).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Soda, M. et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature 448, 561–566 (2007).
Kwak, E. L. et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N. Engl. J. Med. 363, 1693–1703 (2010).
Rowley, J. D. Letter: a new consistent chromosomal abnormality in chronic myelogenous leukaemia identified by quinacrine fluorescence and Giemsa staining. Nature 243, 290–293 (1973).
Kantarjian, H. et al. Hematologic and cytogenetic responses to imatinib mesylate in chronic myelogenous leukemia. N. Engl. J. Med. 346, 645–652 (2002).
Wan, T. S. Cancer cytogenetics: methodology revisited. Ann. Lab. Med. 34, 413–425 (2014).
Zack, T. I. et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, 1134–1140 (2013).
Mardis, E. R. & Wilson, R. K. Cancer genome sequencing: a review. Hum. Mol. Genet. 18, R163–168 (2009).
Inaki, K. et al. Transcriptional consequences of genomic structural aberrations in breast cancer. Genome Res. 21, 676–687 (2011).
Maher, C. A. et al. Transcriptome sequencing to detect gene fusions in cancer. Nature 458, 97–101 (2009).
Zhang, J. et al. INTEGRATE: gene fusion discovery using whole genome and transcriptome data. Genome Res. 26, 108–118 (2016).
Campbell, P. J. et al. Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nat. Genet. 40, 722–729 (2008).
Alkan, C., Coe, B. P. & Eichler, E. E. Genome structural variation discovery and genotyping. Nat. Rev. Genet. 12, 363–376 (2011).
Peifer, M. et al. Telomerase activation by genomic rearrangements in high-risk neuroblastoma. Nature 526, 700–704 (2015).
Nik-Zainal, S. et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534, 47–54 (2016).
Xu, H. et al. Integrative analysis reveals the transcriptional collaboration between EZH2 and E2F1 in the regulation of cancer-related gene expression. Mol. Cancer Res. 14, 163–172 (2016).
Layer, R. M., Chiang, C., Quinlan, A. R. & Hall, I. M. LUMPY: a probabilistic framework for structural variant discovery. Genome Biol. 15, R84 (2014).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, i333–i339 (2012).
Boeva, V. et al. Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics 28, 423–425 (2012).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Wang, Z. et al. The properties of genome conformation and spatial gene interaction and regulation networks of normal and malignant human cell types. PLoS One 8, e58793 (2013).
Barutcu, A. R. et al. Chromatin interaction analysis reveals changes in small chromosome and telomere clustering between epithelial and breast cancer cells. Genome Biol. 16, 214 (2015).
Barutcu, A. R. et al. RUNX1 contributes to higher-order chromatin organization and gene regulation in breast cancer cells. Biochim. Biophys. Acta 1859, 1389–1397 (2016).
Taberlay, P. C. et al. Three-dimensional disorganization of the cancer genome occurs coincident with long-range genetic and epigenetic alterations. Genome Res. 26, 719–731 (2016).
Guo, Y. et al. CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell 162, 900–910 (2015).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechno.l 31, 1119–1125 (2013).
Engreitz, J. M., Agarwala, V. & Mirny, L. A. Three-dimensional genome architecture influences partner selection for chromosomal translocations in human disease. PLoS One 7, e44196 (2012).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Seaman, L. et al. Nucleome analysis reveals structure–function relationships for colon cancer. Mol. Cancer Res. 15, 821–830 (2017).
Harewood, L. et al. Hi-C as a tool for precise detection and characterisation of chromosomal rearrangements and copy number variation in human tumours. Genome Biol. 18, 125 (2017).
Wu, H. J. & Michor, F. A computational strategy to adjust for copy number in tumor Hi-C data. Bioinformatics 32, 3695–3701 (2016).
Chakraborty, A. & Ay, F. Identification of copy number variations and translocations in cancer cells from Hi-C data. Bioinformatics 34, 338–345 (2017).
Naumann, S., Reutzel, D., Speicher, M. & Decker, H. J. Complete karyotype characterization of the K562 cell line by combined application of G-banding, multiplex-fluorescence in situ hybridization, fluorescence in situ hybridization, and comparative genomic hybridization. Leuk. Res. 25, 313–322 (2001).
O’Doherty, A. et al. An aneuploid mouse strain carrying human chromosome 21 with Down syndrome phenotypes. Science 309, 2033–2037 (2005).
Gribble, S. M. et al. Massively parallel sequencing reveals the complex structure of an irradiated human chromosome on a mouse background in the Tc1 model of Down syndrome. PLoS One 8, e60482 (2013).
Rhind, N. & Gilbert, D. M. DNA replication timing. Cold Spring Harb. Perspect. Biol. 5, a010132 (2013).
Dileep, V., Rivera-Mulia, J. C., Sima, J. & Gilbert, D. M. Large-scale chromatin structure-function relationships during the cell cycle and development: insights from replication timing. Cold Spring Harb. Symp. Quant. Biol. 80, 53–63 (2015).
Pope, B. D. et al. Replication-timing boundaries facilitate cell-type and species-specific regulation of a rearranged human chromosome in mouse. Hum. Mol. Genet. 21, 4162–4170 (2012).
Ryba, T. et al. Abnormal developmental control of replication-timing domains in pediatric acute lymphoblastic leukemia. Genome Res. 22, 1833–1844 (2012).
Dileep, V. et al. Topologically associating domains and their long-range contacts are established during early G1 coincident with the establishment of the replication-timing program. Genome Res. 25, 1104–1113 (2015).
Rivera-Mulia, J. C. et al. Dynamic changes in replication timing and gene expression during lineage specification of human pluripotent stem cells. Genome Res. 25, 1091–1103 (2015).
Sima, J. & Gilbert, D. M. Complex correlations: replication timing and mutational landscapes during cancer and genome evolution. Curr. Opin. Genet. Dev. 25, 93–100 (2014).
Chiarle, R. et al. Genome-wide translocation sequencing reveals mechanisms of chromosome breaks and rearrangements in B cells. Cell 147, 107–119 (2011).
Struski, S. et al. Identification of chromosomal loci associated with non-P-glycoprotein-mediated multidrug resistance to topoisomerase II inhibitor in lung adenocarcinoma cell line by comparative genomic hybridization. Genes Chromosomes Cancer 30, 136–142 (2001).
Strefford, J. C. et al. A combination of molecular cytogenetic analyses reveals complex genetic alterations in conventional renal cell carcinoma. Cancer Genet. Cytogenet. 159, 1–9 (2005).
Peng, K. J. et al. Characterization of two human lung adenocarcinoma cell lines by reciprocal chromosome painting. Dongwuxue Yanjiu 31, 113–121 (2010).
Beheshti, B., Karaskova, J., Park, P. C., Squire, J. A. & Beatty, B. G. Identification of a high frequency of chromosomal rearrangements in the centromeric regions of prostate cancer cell lines by sequential giemsa banding and spectral karyotyping. Mol. Diagn. 5, 23–32 (2000).
Liu, J. et al. Modeling of lung cancer by an orthotopically growing H460SM variant cell line reveals novel candidate genes for systemic metastasis. Oncogene 23, 6316–6324 (2004).
Espino, P. S., Pritchard, S., Heng, H. H. & Davie, J. R. Genomic instability and histone H3 phosphorylation induction by the Ras-mitogen activated protein kinase pathway in pancreatic cancer cells. Int. J. Cancer 124, 562–567 (2009).
Sirivatanauksorn, V. et al. Non-random chromosomal rearrangements in pancreatic cancer cell lines identified by spectral karyotyping. Int. J. Cancer 91, 350–358 (2001).
Rondón-Lagos, M. et al. Differences and homologies of chromosomal alterations within and between breast cancer cell lines: a clustering analysis. Mol. Cytogenet. 7, 8 (2014).
Hillmer, A. M. et al. Comprehensive long-span paired-end-tag mapping reveals characteristic patterns of structural variations in epithelial cancer genomes. Genome Res. 21, 665–675 (2011).
Hampton, O. A. et al. Long-range massively parallel mate pair sequencing detects distinct mutations and similar patterns of structural mutability in two breast cancer cell lines. Cancer Genet. 204, 447–457 (2011).
Pendleton, M. et al. Assembly and diploid architecture of an individual human genome via single-molecule technologies. Nat. Methods 12, 780–786 (2015).
Seo, J. S. et al. De novo assembly and phasing of a Korean human genome. Nature 538, 243–247 (2016).
Forbes, S. A. et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 43, D805–811 (2015).
Mifsud, B. et al. Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C. Nat. Genet. 47, 598–606 (2015).
Franke, M. et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 538, 265–269 (2016).
Lupiáñez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Hnisz, D. et al. Activation of proto-oncogenes by disruption of chromosome neighborhoods. Science 351, 1454–1458 (2016).
Northcott, P. A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014).
Weischenfeldt, J. et al. Pan-cancer analysis of somatic copy-number alterations implicates IRS4 and IGF2 in enhancer hijacking. Nat. Genet. 49, 65–74 (2017).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Marchal, C. et al. Genome-wide analysis of replication timing by next-generation sequencing with E/L Repli-seq. Nat. Protoc. 13, 819–839 (2018).
Kim, D. & Salzberg, S. L. TopHat-Fusion: an algorithm for discovery of novel fusion transcripts. Genome Biol. 12, R72 (2011).
Haas, B. et al. STAR-Fusion: fast and accurate fusion transcript detection from RNA-Seq. Preprint at https://www.biorxiv.org/content/early/2017/03/24/120295 (2017).
Benelli, M. et al. Discovering chimeric transcripts in paired-end RNA-seq data by using EricScript. Bioinformatics 28, 3232–3239 (2012).
Klijn, C. et al. A comprehensive transcriptional portrait of human cancer cell lines. Nat. Biotechnol. 33, 306–312 (2015).
van de Geijn, B., McVicker, G., Gilad, Y. & Pritchard, J. K. WASP: allele-specific software for robust molecular quantitative trait locus discovery. Nat. Methods 12, 1061–1063 (2015).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Acknowledgements
This work was supported by NIH grants R35GM124820, R01HG009906, and U01CA200060 (F.Y.), R24DK106766 (R.C.H. and F.Y.), GM083337 (D.M.G.), GM085354 (D.M.G.), DK107965 (D.M.G.), U54HG004592 (J.D. and J.A.S.), HG003143 and DK107980 (J.D.), U41HG007000 (W.S.N.), and DP5OD023071 (J. D.). This work was also supported by European Research Council (No. 615584 to D.T.O.and C.E.), Cancer Research UK (Nos. 20412 and 22398 to D.T.O. and C.E.), Wellcome Trust (No. 84459 to D.T.O. and C.E.), and Wellcome Trust (No. 106985/Z/15/Z to S.H.). J.D. is an investigator of the Howard Hughes Medical Institute. J.R.D. is also supported by the Leona M. and Harry B. Helmsley Charitable Trust grant No. 2017-PG-MED001. F.A. was supported by Institute Leadership Funds from La Jolla Institute for Allergy and Immunology. F.Y. is also supported by the Leukemia Research Foundation and Penn State Clinical and Translational Science Institute. We thank the ENCODE Data Coordination Center for helping with Hi-C and replication time data deposition. We would also like to thank Jan Karlseder and Nausica Arnault for help with the FISH experiments.
Author information
Authors and Affiliations
Contributions
J.X., J.R.D., F.S., and F.Y. led the overall integrative analysis. J.X. and S.F. performed the WGS data analysis. J.R.D. led the overall Hi-C analysis. ENCODE Hi-C data were generated by Y. Z. and analyzed by B.R.L., H.O., and J.D. J.R.D., V.T.L., J.X., and F.Y. performed the additional Hi-C and FISH experiments. J.X., F.Y., A.C. and F.A. contributed to Hi-C analysis. J.X., D.V.B., R.C., J.B., L.Z., C.P., J.R.B., and F.Y. performed the optical mapping and data analysis. V.D., T.S., J.C., and D.G. led the replication timing analysis. C.E. and D.O. prepared the Tc1 material. D.P. and S.H. prepared the Hi-C experiments on Tc1 cells and the preliminary analysis. G.Y., L.Z., H.Y., T.L., S.I., L.A., C.P., R.K., M.B., K.L., M.D., J.S., and D.G. analyzed the data. J.R.D., J.X., V.D., F.S., F.A., R.C.H., W.S.N., J.D., D.G., and F.Y. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–24
Supplementary Table 1
List of cell/tissue types with performed experiments and analysis
Supplementary Table 2
Number of SVs detected by WGS, Hi-C and optical mapping in eight cancer cell lines and NA12878
Supplementary Table 3
SVs detected by WGS in eight cancer cell lines and NA12878
Supplementary Table 4
SVs detected by optical mapping in eight cancer cell lines and NA12878
Supplementary Table 5
SVs detected by Hi-C in 36 cell lines
Supplementary Table 6
High-confidence SV calls from integration
Supplementary Table 7
Validated translocations and deletions in K562, Caki and T47D cells
Supplementary Table 8
Cross comparison of large intrachromosomal rearrangements (≥1 Mb) and interchromosomal translocations
Supplementary Table 9
Contribution by each method and their overlapping percentage with high-confidence SVs
Supplementary Table 10
Integration of intrachromosomal rearrangements (<1 Mb)
Supplementary Table 11
Irys-detected deletions encompass multiple smaller WGS-detected deletions with the same total deletion sizes
Supplementary Table 12
Optical mapping predicts the size of unresolved genome gap in hg19
Supplementary Table 13
Optical mapping provides estimation of gap size in hg38 and comparison to previous gap assessment of hg38
Supplementary Table 14
SV-induced fused genes detected by RNA-seq
Supplementary Table 15
Summary of genes, repetitive elements and insulators overlapping with high-confidence deletions
Supplementary Table 16
Frequency of enhancer deletions versus simulated expectation in cancer cells and normal cells
Supplementary Table 17
Deleted potential enhancers and insulators in T47D, Caki2, K562 and NCIH460
Rights and permissions
About this article
Cite this article
Dixon, J.R., Xu, J., Dileep, V. et al. Integrative detection and analysis of structural variation in cancer genomes. Nat Genet 50, 1388–1398 (2018). https://doi.org/10.1038/s41588-018-0195-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0195-8
This article is cited by
-
A benchmarking framework for the accurate and cost-effective detection of clinically-relevant structural variants for cancer target identification and diagnosis
Journal of Translational Medicine (2024)
-
Alteration of chromosome structure impacts gene expressions implicated in pancreatic ductal adenocarcinoma cells
BMC Genomics (2024)
-
Large-scale genomic rearrangements boost SCRaMbLE in Saccharomyces cerevisiae
Nature Communications (2024)
-
A graph neural network-based interpretable framework reveals a novel DNA fragility–associated chromatin structural unit
Genome Biology (2023)
-
Hi-C analysis of genomic contacts revealed karyotype abnormalities in chicken HD3 cell line
BMC Genomics (2023)