Abstract
DNA methylation and Polycomb are key factors in the establishment of vertebrate cellular identity and fate. Here we report de novo missense mutations in DNMT3A, which encodes the DNA methyltransferase DNMT3A. These mutations cause microcephalic dwarfism, a hypocellular disorder of extreme global growth failure. Substitutions in the PWWP domain abrogate binding to the histone modifications H3K36me2 and H3K36me3, and alter DNA methylation in patient cells. Polycomb-associated DNA methylation valleys, hypomethylated domains encompassing developmental genes, become methylated with concomitant depletion of H3K27me3 and H3K4me3 bivalent marks. Such de novo DNA methylation occurs during differentiation of Dnmt3aW326R pluripotent cells in vitro, and is also evident in Dnmt3aW326R/+ dwarf mice. We therefore propose that the interaction of the DNMT3A PWWP domain with H3K36me2 and H3K36me3 normally limits DNA methylation of Polycomb-marked regions. Our findings implicate the interplay between DNA methylation and Polycomb at key developmental regulators as a determinant of organism size in mammals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The human next-generation sequencing data used in the manuscript are available on request from the relevant Data Access Committee from the European Genome-phenome Archive (EGA). Exome data are available under accession EGAS00001003231. Human RNA-seq, RRBS and ChIP-seq data are available under accession EGAS00001003232. The data are not publicly available to ensure protection of subject sequence data confidentiality through controlled access. Processed data files and mouse RNA-seq and RRBS data are available in GEO under accession GSE120558.
References
Klingseisen, A. & Jackson, A. P. Mechanisms and pathways of growth failure in primordial dwarfism. Genes Dev. 25, 2011–2024 (2011).
Bicknell, L. S. et al. Mutations in the pre-replication complex cause Meier-Gorlin syndrome. Nat. Genet. 43, 356–359 (2011).
Bicknell, L. S. et al. Mutations in ORC1, encoding the largest subunit of the origin recognition complex, cause microcephalic primordial dwarfism resembling Meier-Gorlin syndrome. Nat. Genet. 43, 350–355 (2011).
Guernsey, D. L. et al. Mutations in origin recognition complex gene ORC4 cause Meier-Gorlin syndrome. Nat. Genet. 43, 360–364 (2011).
Burrage, L. C. et al. De novo GMNN mutations cause autosomal-dominant primordial dwarfism associated with Meier-Gorlin syndrome. Am. J. Hum. Genet. 97, 904–913 (2015).
Rauch, A. et al. Mutations in the pericentrin (PCNT) gene cause primordial dwarfism. Science 319, 816–819 (2008).
Griffith, E. et al. Mutations in pericentrin cause Seckel syndrome with defective ATR-dependent DNA damage signaling. Nat. Genet. 40, 232–236 (2008).
Martin, C. A. et al. Mutations in PLK4, encoding a master regulator of centriole biogenesis, cause microcephaly, growth failure and retinopathy. Nat. Genet. 46, 1283–1292 (2014).
Conlon, I. & Raff, M. Size control in animal development. Cell 96, 235–244 (1999).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Tatton-Brown, K. et al. Mutations in the DNA methyltransferase gene DNMT3A cause an overgrowth syndrome with intellectual disability. Nat. Genet. 46, 385–388 (2014).
Tlemsani, C. et al. SETD2 and DNMT3A screen in the Sotos-like syndrome French cohort. J. Med. Genet. 53, 743–751 (2016).
Okamoto, N., Toribe, Y., Shimojima, K. & Yamamoto, T. Tatton-Brown-Rahman syndrome due to 2p23 microdeletion. Am. J. Med. Genet. A. 170A, 1339–1342 (2016).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Dhayalan, A. et al. The Dnmt3a PWWP domain reads histone 3 lysine 36 trimethylation and guides DNA methylation. J. Biol. Chem. 285, 26114–26120 (2010).
Sankaran, S. M., Wilkinson, A. W., Elias, J. E. & Gozani, O. A PWWP domain of histone-lysine N-methyltransferase NSD2 binds to dimethylated Lys-36 of histone H3 and regulates NSD2 function at chromatin. J. Biol. Chem. 291, 8465–8474 (2016).
Qin, S. & Min, J. Structure and function of the nucleosome-binding PWWP domain. Trends. Biochem. Sci. 39, 536–547 (2014).
Rondelet, G., Dal Maso, T., Willems, L. & Wouters, J. Structural basis for recognition of histone H3K36me3 nucleosome by human de novo DNA methyltransferases 3A and 3B. J. Struct. Biol. 194, 357–367 (2016).
Kungulovski, G. et al. Application of histone modification-specific interaction domains as an alternative to antibodies. Genome Res. 24, 1842–1853 (2014).
Du, J., Johnson, L. M., Jacobsen, S. E. & Patel, D. J. DNA methylation pathways and their crosstalk with histone methylation. Nat. Rev. Mol. Cell Biol. 16, 519–532 (2015).
Manzo, M. et al. Isoform-specific localization of DNMT3A regulates DNA methylation fidelity at bivalent CpG islands. EMBO J. 36, 3421–3434 (2017).
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Cao, R. et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 298, 1039–1043 (2002).
Kuzmichev, A., Jenuwein, T., Tempst, P. & Reinberg, D. Different EZH2-containing complexes target methylation of histone H1 or nucleosomal histone H3. Mol. Cell 14, 183–193 (2004).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Li, Y. et al. Genome-wide analyses reveal a role of Polycomb in promoting hypomethylation of DNA methylation valleys. Genome. Biol. 19, 18 (2018).
Jeong, M. et al. Large conserved domains of low DNA methylation maintained by Dnmt3a. Nat. Genet. 46, 17–23 (2014).
Long, H. K. et al. Epigenetic conservation at gene regulatory elements revealed by non-methylated DNA profiling in seven vertebrates. eLife 2, e00348 (2013).
Bartke, T. et al. Nucleosome-interacting proteins regulated by DNA and histone methylation. Cell 143, 470–484 (2010).
Wu, H. et al. Dnmt3a-dependent nonpromoter DNA methylation facilitates transcription of neurogenic genes. Science 329, 444–448 (2010).
Sidoli, S. et al. Middle-down hybrid chromatography/tandem mass spectrometry workflow for characterization of combinatorial post-translational modifications in histones. Proteomics 14, 2200–2211 (2014).
Yuan, W. et al. H3K36 methylation antagonizes PRC2-mediated H3K27 methylation. J. Biol. Chem. 286, 7983–7989 (2011).
Streubel, G. et al. The H3K36me2 methyltransferase Nsd1 demarcates PRC2-mediated H3K27me2 and H3K27me3 domains in embryonic stem cells. Mol. Cell 70, 371–379 (2018).
Smallwood, S. A. & Kelsey, G. De novo DNA methylation: a germ cell perspective. Trends Genet. 28, 33–42 (2012).
Pollard, S. M., Benchoua, A. & Lowell, S. Neural stem cells, neurons, and glia. Methods Enzymol. 418, 151–169 (2006).
Meissner, A. et al. Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis. Nucleic Acids Res. 33, 5868–5877 (2005).
Tippmann, S. C. et al. Chromatin measurements reveal contributions of synthesis and decay to steady-state mRNA levels. Mol. Syst. Biol. 8, 593 (2012).
Challen, G. A. et al. Dnmt3a is essential for hematopoietic stem cell differentiation. Nat. Genet. 44, 23–31 (2011).
Jeong, M. et al. Loss of Dnmt3a immortalizes hematopoietic stem cells in vivo. Cell Rep. 23, 1–10 (2018).
Blackledge, N. P. et al. CpG islands recruit a histone H3 lysine 36 demethylase. Mol. Cell 38, 179–190 (2010).
Wiehle, L. et al. Tet1 and Tet2 protect DNA methylation canyons against hypermethylation. Mol. Cell. Biol. 36, 452–461 (2015).
Gu, T. et al. DNMT3A and TET1 cooperate to regulate promoter epigenetic landscapes in mouse embryonic stem cells. Genome. Biol. 19, 88 (2018).
Boulard, M., Edwards, J. R. & Bestor, T. H. FBXL10 protects Polycomb-bound genes from hypermethylation. Nat. Genet. 47, 479–485 (2015).
Goll, M. G. & Bestor, T. H. Eukaryotic cytosine methyltransferases. Annu. Rev. Biochem. 74, 481–514 (2005).
Voigt, P., Tee, W. W. & Reinberg, D. A double take on bivalent promoters. Genes Dev. 27, 1318–1338 (2013).
Klose, R. J., Cooper, S., Farcas, A. M., Blackledge, N. P. & Brockdorff, N. Chromatin sampling—an emerging perspective on targeting polycomb repressor proteins. PLoS Genet. 9, e1003717 (2013).
Pereira, J. D. et al. Ezh2, the histone methyltransferase of PRC2, regulates the balance between self-renewal and differentiation in the cerebral cortex. Proc. Natl Acad. Sci. USA 107, 15957–15962 (2010).
Kurotaki, N. et al. Haploinsufficiency of NSD1 causes Sotos syndrome. Nat. Genet. 30, 365–366 (2002).
Luscan, A. et al. Mutations in SETD2 cause a novel overgrowth condition. J. Med. Genet. 51, 512–517 (2014).
Tatton-Brown, K. et al. Germline mutations in the oncogene EZH2 cause Weaver syndrome and increased human height. Oncotarget 2, 1127–1133 (2011).
Gibson, W. T. et al. Mutations in EZH2 cause weaver syndrome. Am. J. Hum. Genet. 90, 110–118 (2012).
Cohen, A. S. et al. A novel mutation in EED associated with overgrowth. J. Hum. Genet. 60, 339–342 (2015).
Awad, S. et al. Mutation in PHC1 implicates chromatin remodeling in primary microcephaly pathogenesis. Hum. Mol. Genet. 22, 2200–2213 (2013).
Tatton-Brown, K. et al. Mutations in epigenetic regulation genes are a major cause of overgrowth with intellectual disability. Am. J. Hum. Genet. 100, 725–736 (2017).
Wood, A. R. et al. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat. Genet. 46, 1173–1186 (2014).
Ernst, J. & Kellis, M. Chromatin-state discovery and genome annotation with ChromHMM. Nat. Protoc. 12, 2478–2492 (2017).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Murray, J. E. et al. Extreme growth failure is a common presentation of ligase IV deficiency. Hum. Mutat. 35, 76–85 (2014).
de Bruin, C. et al. An XRCC4 splice mutation associated with severe short stature, gonadal failure, and early-onset metabolic syndrome. J. Clin. Endocrinol. Metab. 100, E789–E798 (2015).
Guerois, R., Nielsen, J. E. & Serrano, L. Predicting changes in the stability of proteins and protein complexes: a study of more than 1000 mutations. J. Mol. Biol. 320, 369–387 (2002).
Triche, T. J. Jr., Weisenberger, D. J., Van Den Berg, D., Laird, P. W. & Siegmund, K. D. Low-level processing of Illumina Infinium DNA Methylation BeadArrays. Nucleic Acids Res. 41, e90 (2013).
Fortin, J. P., Triche, T. J. Jr & Hansen, K. D. Preprocessing, normalization and integration of the Illumina HumanMethylationEPIC array with minfi. Bioinformatics 33, 558–560 (2017).
Illingworth, R. S., Holzenspies, J. J., Roske, F. V., Bickmore, W. A. & Brickman, J. M. Polycomb enables primitive endoderm lineage priming in embryonic stem cells. eLife 5, e14926 (2016).
Orlando, D. A. et al. Quantitative ChIP-Seq normalization reveals global modulation of the epigenome. Cell Rep. 9, 1163–1170 (2014).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Rohde, C., Zhang, Y., Reinhardt, R. & Jeltsch, A. BISMA—fast and accurate bisulfite sequencing data analysis of individual clones from unique and repetitive sequences. BMC Bioinformatics 11, 230 (2010).
Bock, C. et al. BiQ Analyzer: visualization and quality control for DNA methylation data from bisulfite sequencing. Bioinformatics 21, 4067–4068 (2005).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome. Biol. 11, R25 (2010).
Acknowledgements
We are grateful to families and clinicians for their involvement and participation. We thank W. Bickmore, R. Meehan, N. Hastie, T. Baubec and I. Adams for helpful discussions, G. Kelsey for discussion of unpublished data, P. Madapura, G. Taylor, L. Duthie and R. Illingworth for technical advice, and E. Freyer, A. Meynert, Institute of Genetics & Molecular Medicine FACS and Sequencing Cores, Bioresearch & Veterinary Services at the University of Edinburgh, Wellcome Centre for Cell Biology mass-spectroscopy facility and the WTCRF for technical support. A.P.J. is supported by the Medical Research Council UK (MRC, U127580972) and the European Research Council (ERC), through ERC Starter Grant 281847; and now by the European Union’s Horizon 2020 research and innovation program ERC Advanced Grant (grant agreement 788093). D.S. is a Cancer Research UK Career Development fellow (reference C47648/A20837), and work in his laboratory is also supported by a MRC university grant to the MRC Human Genetics Unit. J.A.M. is supported by a MRC Career Development Award (MR/M02122X/1). P.H. was supported by a fellowship within the Postdoc Program of the German Academic Exchange Service (DAAD). V.H. is supported by funding from the National Institutes of Health and Eunice Kennedy Shriver National Institute of Child Health and Human Development (R01HD078592). T.A. is supported by Wellcome Trust funding to R.C.A. (200885). J.R. is supported by the Wellcome Trust through a Senior Research Fellowship (103139) and a multiuser equipment grant (108504). The Wellcome Centre for Cell Biology is supported by core funding from the Wellcome Trust (203149).
Author information
Authors and Affiliations
Contributions
C.V.L., R.C.C., D.A.P. and A.D. performed molecular genetic studies to identify patient mutations. P.H. designed the cell biology experiments, and performed these with C.V.L., A.F., F.T., F.K. and M.A.M.R. P.H., A.F. and C.-A.M. characterized the mouse model. J.A.M. performed structural modeling. T.A. performed mass spectrometry with support from R.C.A. and J.R. V.C.-D., C.-T.F., K.G., V.H., L.I., S.P.R., G.S. and A.D. ascertained subjects, obtained samples and/or assisted with clinical studies. D.S. performed computational analyses of DNA methylation and ChIP-seq and RNA-seq data sets. P.H., D.S. and A.P.J. wrote the manuscript with input from M.A.M.R., and all authors reviewed it. D.S. and A.P.J. planned and supervised the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Recombinant DNMT3A-PWWPW330R is stably expressed but has impaired H3K36me2 and H3K36me3 binding.
a, Left, Coomassie blue–stained SDS–PAGE gels with eluates from GST-PWWP protein purifications show that PWWPWT and PWWPW330R can be purified from bacterial lysates, in contrast to PWWP domains containing the TBRS overgrowth mutations W297del and I310N. Representative image of n = 2 independent experiments. Right, immunoblot probed with anti-GST antibody on total bacterial protein lysates (T) and purified GST-PWWP protein after elution (E) from glutathione beads, demonstrating that PWWPW297del and PWWPI310N are expressed but present at greatly reduced levels. b, MODified Histone Peptide Array representing 384 distinct or combinatorial histone modifications probed with recombinant PWWP proteins as indicated. Shown is an independent experimental replicate (from Fig. 2e; n = 2 independent experiments) for 10 nM PWWP protein and also after probing the array with a higher concentration (100 nM; n = 1 experiment) of PWWPW330R protein. A Myc-tag-positive control was detected with a Myc tag antibody on the latter. Each peptide is spotted in duplicate (left and right halves of the array). Circles on the left half indicate the position of the indicated histone modification (H3K36me2 and H3K36me3) and Myc-positive control peptide on the array.
Supplementary Figure 2 Hypomethylated DMRs in DNMT3AW330R/+ patient fibroblasts are not reproducible across tissues.
a, Heatmap of differentially methylated regions (DMRs; n = 738) with DNA methylation loss in patient fibroblasts (P1, P2) compared to control fibroblasts (C1, C2). Loss is not evident in control (C3, C4), overgrowth (O1, O2) or patient peripheral blood leukocytes (blood samples). Right, DNA methylation loss is also not evident in fibroblasts from microcephalic dwarfism (MD) patients with PCNT mutations (PCNT1, PCNT2). b, Quantification of DNA methylation loss for the DMRs depicted in a. Box, 25th–75th percentile; whiskers, full data range; center line, median; ∆%mCpG, percent change in methylation relative to mean of controls. c, Hypermethylated DMRs are not evident in fibroblasts from MD patients with PCNT mutations. Heatmaps reproduced from Fig. 3b (left) compared with those of DNA methylation levels in control and PCNT patient fibroblasts (PCNT1, PCNT2) (right) over DMRs hypermethylated in DNMT3AW330R/+ patients. d, Quantification of the heatmap data from c for control and PCNT patients. Box, 25th–75th percentile; whiskers, full data range; center line, median. C1–C5: healthy controls; P1, P2: DNMT3AW330R/+ MD patients; O1, O2: TBRS overgrowth patients; PCNT1, PCNT2: MD patients with PCNT mutation.
Supplementary Figure 3 P3 exhibits the same DNA hypermethylation pattern.
a, Regions with methylation gain identified in patient 1 (P1) and patient 2 (P2) fibroblasts (DNMT3AW330R/+) also gain DNA methylation in patient 3 (P3; DNMT3AD333N/+) peripheral blood leukocytes (PBLs; ‘blood samples’). Heatmaps for DMRs with methylation gain in P1 and P2 fibroblasts, and P2 and P3 PBLs, excluding DMRs absent from P3 RRBS data. C1–C4, healthy controls. b, Regions that lose DNA methylation in fibroblasts identified in P1 and P2 do not lose DNA methylation in P3 PBLs. Heatmaps for DMRs with methylation loss in P1 and P2 fibroblasts, and P2 and P3 PBLs, excluding DMRs absent from P3 RRBS data. c, Quantification of DNA methylation for DMRs depicted for P3 in a and b. Box, 25th–75th percentile; whiskers, full data range; center line, median; ∆%mCpG P3-C3, percent change of methylation in patient 3 relative to control 3. n = 282 hypo-DMRs and n = 845 hyper-DMRs. d, Representative genome browser views of hypermethylated DMRs (n = 845) demonstrating increased DNA methylation at the same developmental genes in patient P3 (compare Fig. 3f). CpGs with coverage depth of ≥5 reads in both samples are plotted. All tracks are scaled 0–100%. mCpG, DNA methylation; CGI, CpG island.
Supplementary Figure 4 Hypermethylated DMRs lose H3K27me3 in patient cells.
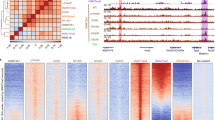
a,b, H3K27me3 is decreased at hypermethylated DMRs in patient cells. a, Heatmaps of normalized H3K27me3 ChIP-seq reads for each control and patient fibroblast cell line centered on DMRs, ranked for mean H3K27me3 levels in control fibroblasts. Scale indicates normalized read counts. Window size, 250 bp. b, Quantification of the data shown in a. Median H3K27me3 profile at hypermethylated DMRs (n = 1,131 DMRs). Median (dark) and 95% confidence intervals (light) for control (blue) and patient (red) dermal fibroblasts plotted. Window size, 250 bp. H3K27me3 is significantly decreased at DMRs in patient samples (two-sided Wilcoxon rank-sum test, patients versus controls P = 5.2 × 10–70, tested across the length of each DMR). c, H3K27me3 is reduced in patients at Polycomb-marked DMVs. P value, two-sided Wilcoxon rank-sum test, Polycomb positive (+) (n = 524) versus negative (–) DMVs (n = 628). d–f, H3K37me3 is most reduced at DMRs in patient cells. d, Heatmaps for mean normalized H3K27me3 ChIP-seq reads in control (C1, C2 mean) and patient (P1, P2 mean) fibroblasts. Maps are centered on reproducible H3K27me3 peaks that overlap Infinium DNA methylation probe sites. e, Quantification of the heatmap in d demonstrates a significantly greater decrease in H3K27me3 levels at hypermethylated DMRs compared to H3K27me3 peaks at other genomic locations. P value, two-sided Wilcoxon rank-sum test, hyper-DMRs (n = 498 H3K27me3 peaks) versus other sites (n = 3,317 H3K27me3 peaks) for log2 patients/controls. c,e, The y axes indicate the difference between patients and controls: 0, no change; >0, increased in patients; <0, decreased in patients. Box, 25th–75th percentile; whiskers, 1.5× interquartile range. f, ChIP-Rx confirms decreased H3K27me3 levels at hypermethylated DMRs in patient 1. Median spike-in normalized H3K27me3 profile at hypermethylated DMRs. Median (dark) and 95% confidence intervals (light) for control 1 (C1, blue) and patient 1 (P1, red) dermal fibroblasts are plotted. Window size, 250 bp. H3K27me3 is significantly decreased at DMRs (n = 1,128) in the patient sample (two-sided Wilcoxon rank-sum test, patient versus control P = 2.35 × 10–42, tested across the length of each DMR for normalized read counts). g, Expression levels of core PRC2 components are not changed in patient fibroblasts. Plotted are RPKM values for control (C1, C2) and patient (P1, P2) dermal fibroblasts derived from RNA-seq. h, Protein levels of the H3K27me2/3 histone methyltransferase EZH2 are not reduced in patient fibroblasts. Immunoblotting for EZH2 of control and patient fibroblast cell lysates and an EZH2 siRNA knockdown control experiment in HeLa cells confirming antibody specificity is shown. Representative of n = 2 independent experiments. Immunoblots are cropped. i, Histone post-translational modification quantification by mass spectrometry demonstrates that global levels of H3K27me3 are unchanged between patient and control dermal fibroblasts. Plotted is the relative peptide abundance for control (control 1 and 2) and patient (patient 1 and 2) dermal fibroblasts. Each dot represents an independently extracted sample from n = 2 independent cell cultures of each control and patient line (total n = 4). Bar height indicates the mean.
Supplementary Figure 5 H3K4me3 is reduced at regions that gain DNA methylation in patient fibroblasts.
a, Heatmaps of normalized H3K4me3 ChIP-seq reads in control (C1, C2) and patient (P1, P2) fibroblasts centered on DMRs with DNA methylation gain, ranked for DMR H3K4me3 levels in controls (C1, C2 mean). Scale indicates normalized read counts. Window size, 250 bp. b, Quantification of the data shown in a. Median H3K4me3 levels at hypermethylated DMRs (n = 1,129 DMRs). Median (dark) and 95% confidence intervals (light) for control (blue) and patient (red) dermal fibroblasts are plotted. Window size, 250 bp. H3K4me3 is significantly decreased at DMRs in patient samples (two-sided Wilcoxon rank-sum test, patients versus controls P=4.33 × 10–130, tested across the length of the DMR for normalized read counts). c, H3K4me3 is reduced in patients at Polycomb-marked DMVs. P value, two-sided Wilcoxon rank-sum test, Polycomb positive (+) (n = 524 DMVs) versus negative (–) DMVs (n = 628). d,e, Comparison of H3K4me3 at DMRs with other sites. d, Heatmaps for mean normalized H3K4me3 ChIP-seq reads in control (C1, C2 mean) and patient (P1, P2 mean) fibroblasts. Maps centered on reproducible H3K4me3 peaks that overlap Infinium DNA methylation probe sites. e, Quantification of the heatmap in d shows a significant decrease in H3K4me3 levels at hypermethylated DMRs compared to H3K4me3 peaks at other genomic locations. P value, two-sided Wilcoxon rank-sum test, hyper-DMRs (n = 720 H3K4me3 peaks) versus other sites (n = 14,850 H3K4me3 peaks) for log2 patients/controls). c,e, The y axes indicate the difference between patients and controls: 0, no change; >0, increased in patients; <0, decreased in patients. For the Polycomb-marked DMV definition, see Methods. Box, 25th–75th percentile; whiskers, 1.5× interquartile range.
Supplementary Figure 6 H3K36me3 is depleted at hyper-DMRs and DMVs.
a, Representative genome browser views of H3K36me3 ChIP-Rx-seq experiment. Few H3K36me3 sequencing reads are detected at FOXA1 and SOX1 DMVs. The region surrounding the housekeeping gene GAPDH is plotted as a positive control for comparison, demonstrating high levels of H3K36me3 over gene bodies. DNA methylation (magenta, scale 0–100%, all tracks), H3K27me3 (green, scale 0–4 scaled read counts per 107 reads, all tracks), H3K4me3 (yellow, scale 0–8 scaled read counts per 107 reads, all tracks) in control (C1, C2) and patient (P1, P2) dermal fibroblasts. DNA methylation, H3K27me3 and H3K4me3 data for SOX1 and FOXA1 (Fig. 3f) are reproduced again here for comparison with H3K36me3 coverage. b, Mean H3K36me3 ChIP-Rx levels at DMVs in control fibroblasts. Normalized read counts are plotted, with all DMVs scaled to the same length. The start of the DMV (0%) to the end of the DMV (100%) is highlighted in gray. The horizontal line indicates no enrichment compared to ChIP input control. c, H3K36me3 peaks are significantly depleted at hypermethylated DMRs. Bar graph, mean percentage of Infinium Methylation probes overlapping H3K36me3 peaks in control (C1, C2) and patient (P1, P2) fibroblasts. Data points, percentage of probes overlapping H3K36me3 peaks, for each respective cell line. All, all probes on the array (n = 403,064 probes). Hyper-DMRs, probes within hypermethylated DMRs (n = 9,085). Error bars, s.d. P value, two-sided Fisher’s exact tests all versus hyper-DMR probes.
Supplementary Figure 7 Bisulfite sequencing of NPC differentiation.
a,b, Locus-specific bisulfite sequencing analysis of Foxa1 (a) and Sox1 (b) of LIF/serum-maintained mESCs. Blocks, independent cell lines; blue and red squares, unmethylated and methylated CpGs, respectively; white squares, undetermined methylation status; columns, CpG sites; rows, individual sequences. Data shown for Foxa1 are representative of n = 2 independent experiments, het cell line n = 1 experiment. Sox1, n = 1 experiment. c, RRBS sequencing confirms DNA methylation gain at DMRs genome-wide upon differentiation. Heatmap of differentially methylated regions (DMRs) after 9-day NPC differentiation. d, Quantification of DNA methylation levels for the hyper-DMRs depicted in c. Datapoints, median percent change in methylation. ∆%mCpG, percent change in methylation relative to the mean for control. P value, two-tailed t test. Horizontal bar, mean per group.
Supplementary Figure 8 Reduced gene expression in hypermethylated DMRs.
a,b, Reproducibility of RNA-seq between samples. a, Scatterplots with pairwise comparisons for control (C1, C2) and patient (P1, P2) fibroblast RNA-seq data sets. Each dot represents the expression level for a gene (n = 11,963 genes) for both samples, measured in log2 CPM. b, Scatterplots with pairwise comparisons for wild-type (WT1, WT2, WT3) and Dnmt3aW326R/W326R (hom1, hom2, hom3) 9-d NPC differentiation RNA-seq data sets. Each dot represents the expression level for a gene (n = 13,022 genes) for both samples, measured in log2 CPM. CPM, counts per million; R, Pearson correlation coefficient. c, Reduced expression levels for genes associated with hyper-DMRs are evident in fibroblasts. Plotted, log2 CPM ratios, box, 25th–75th percentile; whiskers, 1.5× interquartile range from box; center line, median. P value, two-sided Wilcoxon rank-sum test, All genes (n = 11,495) with coverage in beadarrays versus genes associated with hypo-DMRs (n = 286) or hyper-DMRs (n = 435). d, Model linking transcriptional changes in stem/progenitor cells due to DNMT3A mutations with organ/organism size. Gain- and loss-of-function mutations in DNMT3A are proposed to impact on stem cell fate dynamics. Owing to increased expression of multipotency genes and a decrease in differentiation/neurogenic gene expression, loss of DNMT3A promotes self-renewal of stem/progenitor cells at the expense of differentiation31,39. This increase in stem cell/progenitor pool size would be expected to result in increased final cell number generated during development, that is, overgrowth. Gain-of-function DNMT3A mutations in the PWWP domain cause an opposite transcriptional bias and therefore might be expected to accelerate differentiation at the expense of self-renewal, depleting progenitor pools and resulting in reduced final cell number and brain/organism size. ~46 rounds of cell division are required to generate the ~1014 cells that make up the human body; therefore, small effects over an extended timeframe in cell fate choice (progenitor versus differentiation) could account for substantial differences in final size.
Supplementary Figure 9 Dnmt3aW326R/+ mice are small with reduced brain weight, cortical surface area, and exhibit hypermethylation at DMRs/DMVs.
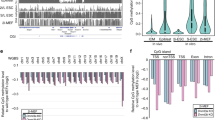
a, Schematic of the CRISPR–Cas9-mediated homology-directed repair strategy for generation of Dnmt3aW326R mice. Alignment of antisense repair template (ODN) to the genomic DNA (gDNA) sequence of Dnmt3a. The targeting sequence of gRNA is shaded in yellow, the PAM site in blue. The triplet coding for tryptophan 326 (TGG) is mutated to arginine (AGA), simultaneously destroying the PAM site. b, cDNA sequencing of Dnmt3aW326R/+ and wild-type littermate (8 week old, female) confirms that the Trp326Arg mutant allele is present and expressed in mice. Sequence electropherograms of cDNA samples with the site of mutation indicated (c.TGG>AGA, p.Trp326Arg). c, Brain weight of female Dnmt3aW326R/+ mice compared to wild-type littermates at 8 weeks of age. n = 3, each group. d, Body weight and cortical area for 5-month-old Dnmt3aW326R/+ mice compared to wild-type littermates. n = 7 wild-type and n = 9 Dnmt3aW326R/+ animals. (The body weight of these animals is also plotted in Fig. 6b at a time point corresponding to 6 weeks of age.) c,d, P value, two-tailed t test. Horizontal bar, mean per group. e, Bisulfite sequencing of the Sox1 locus for cortex samples from Dnmt3aW326R/+ and wild-type littermates (n = 3/group; female, age 8 weeks). Blocks, individual mice; blue and red squares, unmethylated and methylated CpGs, respectively; white squares, undetermined methylation status; columns, CpG sites; rows individual sequences. f–h, RRBS confirms gain of DNA methylation at NPC DMRs in adult mouse cortexes. f, Heatmap of hypermethylated DMRs identified in in vitro NPC differentiation showing gains in Dnmt3aW326R/+ cerebral cortex. Each column in the heatmap represents an individual mouse. g, Quantification of the data shown in f. Plotted is the median percent change in methylation (∆%mCpG) relative to the mean for controls for all hypermethylated regions per sample. P value, two-tailed t test. Horizontal red bar, mean per group. n = 3, each group. h, Genome browser view of a region that gains DNA methylation in Dnmt3aW326R/+ mouse cortex. Scale 0–100%. M1–M6, individual mice.
Supplementary Figure 10 Global description of DNA methylation data sets.
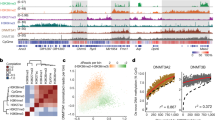
a, Density histograms of methylation levels (beta values) for Infinium beadarray data sets. Plotted are the density distributions of methylation levels for all examined probes. Each line represents an independent sample. C1–C5: healthy controls; P1, P2: DNMT3AW330R/+ microcephalic dwarfism patients; O1, O2: TBRS overgrowth patients; PCNT1, PCNT2: microcephalic dwarfism patients with PCNT mutation. Bandwidth is defined as the s.d. of the smoothing kernel. b, Median methylation levels (beta values) do not significantly differ between patient and control groups. Each dot represents a sample. Blood samples, n = 2 controls, n = 1 microcephalic dwarfism patient (P2), n = 2 TBRS patients. Fibroblasts, n = 3 controls, n = 2 DNMT3A microcephalic dwarfism patients (P1, P2), n = 2 PCNT microcephalic dwarfism patients. Horizontal bar, mean per group. c, Density histogram of methylation levels for patient 3 and control 3 reduced-representation bisulfite sequencing data sets. Plotted are the density distributions of methylation levels for all CpGs with coverage ≥10. d,e, Genic DNA methylation profiles are similar between patients and controls. Scaled metagene profiles for all autosomal genes plotted (n = 18,754). Each line represents the mean of an individual sample. TSS, transcription start site; TES, transcription end site. f,g, Density histograms of methylation levels for NPC differentiation (f) and mouse cortex (g) reduced-representation bisulfite sequencing data sets. Plotted are the density distributions of methylation levels for all CpGs with coverage ≥10. Each line represents an independent sample. h, Overall methylation level for NPC differentiation and cortex samples from 8-week-old mice in reduced-representation bisulfite sequencing data sets. Plotted is the weighted mean percentage methylation. P value, two-tailed t test. Each dot represents an individual sample. NPCs, n = 3 independent wild-type samples and n = 4 independent W326R samples. Cortexes, n = 3 per group. Horizontal bar, mean per group.
Supplementary Figure 11 Uncropped immunoblot and gel images.
Uncropped immunoblot and Coomassie gel images from main and supplementary figures. Red frames indicate the cropped area.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11, Supplementary Tables 1, 4 and 6–9, and Supplementary Note
Supplementary Table 2
Hypermethylated DMRs from human patient fibroblasts
Supplementary Table 3
Hypomethylated DMRs from human patient fibroblasts
Supplementary Table 5
Hypermethylated DMRs from mouse neural differentiation
Supplementary Table 10
Hypomethylated DMRs from mouse neural differentiation
Rights and permissions
About this article
Cite this article
Heyn, P., Logan, C.V., Fluteau, A. et al. Gain-of-function DNMT3A mutations cause microcephalic dwarfism and hypermethylation of Polycomb-regulated regions. Nat Genet 51, 96–105 (2019). https://doi.org/10.1038/s41588-018-0274-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0274-x
This article is cited by
-
Interrogating epigenetic mechanisms with chemically customized chromatin
Nature Reviews Genetics (2024)
-
Combined and differential roles of ADD domains of DNMT3A and DNMT3L on DNA methylation landscapes in mouse germ cells
Nature Communications (2024)
-
DNMT3B PWWP mutations cause hypermethylation of heterochromatin
EMBO Reports (2024)
-
Base editor scanning charts the DNMT3A activity landscape
Nature Chemical Biology (2023)
-
Regulation, functions and transmission of bivalent chromatin during mammalian development
Nature Reviews Molecular Cell Biology (2023)