Abstract
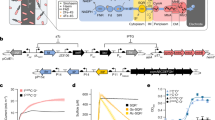
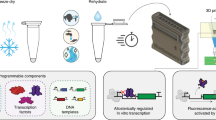
Cell-based biosensors have great potential to detect various toxic and pathogenic contaminants in aqueous environments. However, frequently they cannot meet practical requirements due to insufficient sensing performance. To address this issue, we investigated a modular, cascaded signal amplifying methodology. We first tuned intracellular sensory receptor densities to increase sensitivity, and then engineered multi-layered transcriptional amplifiers to sequentially boost output expression level. We demonstrated these strategies by engineering ultrasensitive bacterial sensors for arsenic and mercury, and improved detection limit and output up to 5,000-fold and 750-fold, respectively. Coupled by leakage regulation approaches, we developed an encapsulated microbial sensor cell array for low-cost, portable and precise field monitoring, where the analyte can be readily quantified via displaying an easy-to-interpret volume bar-like pattern. The ultrasensitive signal amplifying methodology along with the background regulation and the sensing platform will be widely applicable to many other cell-based sensors, paving the way for their real-world applications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data and plasmids supporting the findings are available from the corresponding author upon reasonable request.
References
van der Meer, J. R. & Belkin, S. Where microbiology meets microengineering: design and applications of reporter bacteria. Nat. Rev. Microbiol. 8, 511–522 (2010).
Kim, H. J., Jeong, H. & Lee, S. J. Synthetic biology for microbial heavy metal biosensors. Anal. Bioanal. Chem. 410, 1191–1203 (2018).
Wang, B., Barahona, M. & Buck, M. A modular cell-based biosensor using engineered genetic logic circuits to detect and integrate multiple environmental signals. Biosens. Bioelectron. 40, 368–376 (2013).
Stocker, J. et al. Development of a set of simple bacterial biosensors for quantitative and rapid measurements of arsenite and arsenate in potable water. Environ. Sci. Technol. 37, 4743–4750 (2003).
De Mora, K. et al. A pH-based biosensor for detection of arsenic in drinking water. Anal. Bioanal. Chem. 400, 1031–1039 (2011).
Cao, Y. et al. Programmable assembly of pressure sensors using pattern-forming bacteria. Nat. Biotechnol. 35, 1087–1093 (2017).
Riglar, D. T. et al. Engineered bacteria can function in the mammalian gut long-term as live diagnostics of inflammation. Nat. Biotechnol. 35, 653–658 (2017).
Mimee, M. et al. An ingestible bacterial-electronic system to monitor gastrointestinal health. Science 360, 915–918 (2018).
Courbet, A. et al. Detection of pathological biomarkers in human clinical samples via amplifying genetic switches and logic gates. Sci. Transl. Med. 7, 289ra83 (2015).
Watstein, D. M. & Styczynski, M. P. Development of a pigment-based whole-cell zinc biosensor for human serum. ACS Synth. Biol. 7, 267–275 (2018).
Duan, F. & March, J. C. Engineered bacterial communication prevents Vibrio cholerae virulence in an infant mouse model. Proc. Natl Acad. Sci. USA 107, 11260–11264 (2010).
Hwang, I. Y. et al. Reprogramming microbes to be pathogen-seeking killers. ACS Synth. Biol. 3, 228–237 (2014).
Ho, C. L. et al. Engineered commensal microbes for diet-mediated colorectal-cancer chemoprevention. Nat. Biomed. Eng. 2, 27–37 (2018).
Zhang, F., Carothers, J. M. & Keasling, J. D. Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat. Biotechnol. 30, 354–359 (2012).
Cerminati, S., Soncini, F. C. & Checa, S. K. Selective detection of gold using genetically engineered bacterial reporters. Biotechnol. Bioeng. 108, 2553–2560 (2011).
Belkin, S. et al. Remote detection of buried landmines using a bacterial sensor. Nat. Biotechnol. 35, 308–310 (2017).
Dana, G. V., Kuiken, T., Rejeski, D. & Snow, A. A. Synthetic biology: four steps to avoid a synthetic-biology disaster. Nature 483, 29–29 (2012).
Shemer, B., Koshet, O., Yagur-Kroll, S. & Belkin, S. Microbial bioreporters of trace explosives. Curr. Opin. Biotechnol. 45, 113–119 (2017).
Landry, B. P. et al. Phosphatase activity tunes two-component system sensor detection threshold. Nat. Commun. 9, 1433 (2018).
Kim, H. J. et al. Development of a highly specific and sensitive cadmium and lead microbial biosensor using synthetic CadC-T7 genetic circuitry. Biosens. Bioelectron. 79, 701–708 (2016).
Wang, B., Barahona, M. & Buck, M. Engineering modular and tunable genetic amplifiers for scaling transcriptional signals in cascaded gene networks. Nucleic Acids Res. 42, 9484–9492 (2014).
Wang, B., Barahona, M. & Buck, M. Amplification of small molecule-inducible gene expression via tuning of intracellular receptor densities. Nucleic Acids Res. 43, 1955–1964 (2015).
Volpetti, F., Petrova, E. & Maerkl, S. J. A microfluidic biodisplay. ACS Synth. Biol. 6, 1979–1987 (2017).
Prindle, A. et al. A sensing array of radically coupled genetic ‘biopixels’. Nature 481, 39–44 (2012).
Ang, J. et al. Tuning response curves for synthetic biology. ACS Synth. Biol. 2, 547–567 (2013).
Merulla, D. & van der Meer, J. R. Regulatable and modulable background expression control in prokaryotic synthetic circuits by auxiliary repressor binding sites. ACS Synth. Biol. 5, 36–45 (2016).
Chen, Y. et al. Tuning the dynamic range of bacterial promoters regulated by ligand-inducible transcription factors. Nat. Commun. 9, 64 (2018).
Salis, H. M., Mirsky, E. A. & Voigt, C. A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Fernandez-Rodriguez, J. & Voigt, C. A. Post-translational control of genetic circuits using Potyvirus proteases. Nucleic Acids Res. 44, 6493–6502 (2016).
Cameron, D. E. & Collins, J. J. Tunable protein degradation in bacteria. Nat. Biotechnol. 32, 1276–1281 (2014).
Ravenscroft, P. Predicting the Global Extent of Arsenic Pollution of Groundwater and its Potential Impact on Human Health (UNICEF, New York, 2007).
Wang, B. & Buck, M. Customizing cell signaling using engineered genetic logic circuits. Trends Microbiol. 20, 376–384 (2012).
Quiles-Puchalt, N. et al. A super-family of transcriptional activators regulates bacteriophage packaging and lysis in Gram-positive bacteria. Nucleic Acids Res. 41, 7260–7275 (2013).
Rhodius, V. A. et al. Design of orthogonal genetic switches based on a crosstalk map of σs, anti-σs, and promoters. Mol. Syst. Biol. 9, 702 (2013).
Liu, Q. et al. Orthogonality and burdens of heterologous and gate gene circuits in E. coli. ACS Synth. Biol. 7, 553–564 (2018).
Potvin-Trottier, L., Lord, N. D., Vinnicombe, G. & Paulsson, J. Synchronous long-term oscillations in a synthetic gene circuit. Nature 538, 514–517 (2016).
Chang, C. C. et al. Structural basis of the mercury(II)-mediated conformational switching of the dual-function transcriptional regulator MerR. Nucleic Acids Res. 43, 7612–7623 (2015).
Wackwitz, A. et al. Internal arsenite bioassay calibration using multiple bioreporter cell lines. Microb. Biotechnol. 1, 149–157 (2008).
Nielsen, A. A. K. et al. Genetic circuit design automation. Science 352, aac7341 (2016).
Buffi, N. et al. Development of a microfluidics biosensor for agarose-bead immobilized Escherichia coli bioreporter cells for arsenite detection in aqueous samples. Lab Chip 11, 2369–2377 (2011).
Willsky, G. R. & Malamy, M. H. Effect of arsenate on inorganic phosphate transport in Escherichia coli. J. Bacteriol. 144, 366–374 (1980).
Pardee, K. et al. Rapid, low-cost detection of Zika virus using programmable biomolecular components. Cell 165, 1255–1266 (2016).
Rampley, C. P. N. et al. Development of simcells as a novel chassis for functional biosensors. Sci. Rep. 7, 7261 (2017).
Cevenini, L. et al. A novel bioluminescent NanoLuc yeast-estrogen screen biosensor (nanoYES) with a compact wireless camera for effect-based detection of endocrine-disrupting chemicals. Anal. Bioanal. Chem. 410, 1237–1246 (2018).
Wen, K. Y. et al. A cell-free biosensor for detecting quorum sensing molecules in P. aeruginosa-infected respiratory samples. ACS Synth. Biol. 6, 2293–2301 (2017).
Nikel, P. I., Martínez-García, E. & De Lorenzo, V. Biotechnological domestication of pseudomonads using synthetic biology. Nat. Rev. Microbiol. 12, 368–379 (2014).
Choi, P. J., Cai, L., Frieda, K. & Xie, X. S. A stochastic single-molecule event triggers phenotype switching of a bacterial cell. Science 322, 442–446 (2008).
Ma, D. et al. Low-cost detection of norovirus using paper-based cell-free systems and synbody-based viral enrichment. Synth. Biol. 3, ysy018 (2018).
Liu, X. et al. 3D printing of living responsive materials and devices. Adv. Mater. 30, 107821 (2018).
Shetty, R. P., Endy, D. & Knight, T. F. Engineering BioBrick vectors from BioBrick parts. J. Biol. Eng. 2, 5 (2008).
Ochman, H., Gerber, A. S. & Hartl, D. L. Genetic applications of an inverse polymerase chain reaction. Genetics 120, 621–623 (1988).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Armbruster, D. A. & Pry, T. Limit of blank, limit of detection and limit of quantitation. Clin. Biochem. Rev. 29, S49–S52 (2008).
Wang, B., Kitney, R. I., Joly, N. & Buck, M. Engineering modular and orthogonal genetic logic gates for robust digital-like synthetic biology. Nat. Commun. 2, 508 (2011).
Acknowledgements
We thank M. Billah (Khulna University) and his colleagues for facilitating our collections of groundwater samples in Bangladesh. This work was supported by UK BBSRC project grant (no. BB/N007212/1), Leverhulme Trust research grant (no. RPG-2015-445), Wellcome Trust Seed Award in Science (no. 202078/Z/16/Z) and EPSRC/BBSRC Global Challenges Research Fund Awards. X.W. was supported by scholarships from the China Scholarship Council and Scottish Universities Life Sciences Alliance. F.V., E.P. and S.J.M. were supported by the Ecole Polytechnique Federale de Lausanne, a Swiss National Science Foundation Grant (no. CR23I2 140697) and a SystemsX.ch Special Opportunity Grant (no. 2015/325).
Author information
Authors and Affiliations
Contributions
B.W. conceived and led the study. X.W. designed the experiments with inputs and supervision from B.W. and C.F. X.W. performed all the experiments and data analysis excluding the microfluidics-based experiments. F.V., E.P. and S.J.M. designed and performed the microfluidics-based experiments. All authors took part in the interpretation of results and preparation of materials for the manuscript. B.W. and X.W. wrote the manuscript with comments from all authors.
Corresponding author
Ethics declarations
Competing interests
B.W. and X.W. filed a patent application based on the technology invention in this work.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–5, Supplementary Figures 1–11
Rights and permissions
About this article
Cite this article
Wan, X., Volpetti, F., Petrova, E. et al. Cascaded amplifying circuits enable ultrasensitive cellular sensors for toxic metals. Nat Chem Biol 15, 540–548 (2019). https://doi.org/10.1038/s41589-019-0244-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0244-3
This article is cited by
-
Customizing cellular signal processing by synthetic multi-level regulatory circuits
Nature Communications (2023)
-
Biosensor-based high-throughput screening enabled efficient adipic acid production
Applied Microbiology and Biotechnology (2023)
-
Using fungible biosensors to evolve improved alkaloid biosyntheses
Nature Chemical Biology (2022)
-
Programming cell-free biosensors with DNA strand displacement circuits
Nature Chemical Biology (2022)
-
Reprogrammed tracrRNAs enable repurposing of RNAs as crRNAs and sequence-specific RNA biosensors
Nature Communications (2022)