Abstract
Redox cycles have been reported in ultradian, circadian and cell cycle-synchronized systems. Redox cycles persist in the absence of transcription and cyclin-CDK activity, indicating that cells harbor multiple coupled oscillators. Nonetheless, the causal relationships and molecular mechanisms by which redox cycles are embedded within ultradian, circadian or cell division cycles remain largely elusive. Yeast harbor an ultradian oscillator, the yeast metabolic cycle (YMC), which comprises metabolic/redox cycles, transcriptional cycles and synchronized cell division. Here, we reveal the existence of robust cycling of H2O2 and peroxiredoxin oxidation during the YMC and show that peroxiredoxin inactivation disrupts metabolic cycling and abolishes coupling with cell division. We find that thiol-disulfide oxidants and reductants predictably modulate the switching between different YMC metabolic states, which in turn predictably perturbs cell cycle entry and exit. We propose that oscillatory H2O2-dependent protein thiol oxidation is a key regulator of metabolic cycling and its coordination with cell division.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All datasets generated or analyzed during this study are included in this Article and its Supplementary Information. Source data are provided with this paper.
References
Takahashi, J. S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 18, 164–179 (2017).
Dunlap, J. C. Molecular bases for circadian clocks. Cell 96, 271–290 (1999).
Gaucher, J., Montellier, E. & Sassone-Corsi, P. Molecular cogs: interplay between circadian clock and cell cycle. Trends Cell Biol. 28, 368–379 (2018).
Nakajima, M. et al. Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science 308, 414–415 (2005).
Tomita, J., Nakajima, M., Kondo, T. & Iwasaki, H. No transcription-translation feedback in circadian rhythm of KaiC phosphorylation. Science 307, 251–254 (2005).
Sweeney, B. M., Tuffli, C. F. Jr & Rubin, R. H. The circadian rhythm in photosynthesis in Acetabularia in the presence of actinomycin D, puromycin and chloramphenicol. J. Gen. Physiol. 50, 647–659 (1967).
Woolum, J. C. A re-examination of the role of the nucleus in generating the circadian rhythm in Acetabularia. J. Biol. Rhythms 6, 129–136 (1991).
Causton, H. C., Feeney, K. A., Ziegler, C. A. & O’Neill, J. S. Metabolic cycles in yeast share features conserved among circadian rhythms. Curr. Biol. 25, 1056–1062 (2015).
Edgar, R. S. et al. Peroxiredoxins are conserved markers of circadian rhythms. Nature 485, 459–464 (2012).
O’Neill, J. S. & Reddy, A. B. Circadian clocks in human red blood cells. Nature 469, 498–503 (2011).
O’Neill, J. S. et al. Circadian rhythms persist without transcription in a eukaryote. Nature 469, 554–558 (2011).
Cho, C. S., Yoon, H. J., Kim, J. Y., Woo, H. A. & Rhee, S. G. Circadian rhythm of hyperoxidized peroxiredoxin II is determined by hemoglobin autoxidation and the 20S proteasome in red blood cells. Proc. Natl Acad. Sci. USA 111, 12043–12048 (2014).
Henslee, E. A. et al. Rhythmic potassium transport regulates the circadian clock in human red blood cells. Nat. Commun. 8, 1978 (2017).
Tu, B. P., Kudlicki, A., Rowicka, M. & McKnight, S. L. Logic of the yeast metabolic cycle: temporal compartmentalization of cellular processes. Science 310, 1152–1158 (2005).
Tu, B. P. et al. Cyclic changes in metabolic state during the life of a yeast cell. Proc. Natl Acad. Sci. USA 104, 16886–16891 (2007).
Burnetti, A. J., Aydin, M. & Buchler, N. E. Cell cycle start is coupled to entry into the yeast metabolic cycle across diverse strains and growth rates. Mol. Biol. Cell 27, 64–74 (2016).
Papagiannakis, A., Niebel, B., Wit, E. C. & Heinemann, M. Autonomous metabolic oscillations robustly gate the early and late cell cycle. Mol. Cell 65, 285–295 (2017).
Chen, Z., Odstrcil, E. A., Tu, B. P. & McKnight, S. L. Restriction of DNA replication to the reductive phase of the metabolic cycle protects genome integrity. Science 316, 1916–1919 (2007).
Klevecz, R. R., Bolen, J., Forrest, G. & Murray, D. B. A genomewide oscillation in transcription gates DNA replication and cell cycle. Proc. Natl Acad. Sci. USA 101, 1200–1205 (2004).
Lloyd, D., Lemar, K. M., Salgado, L. E., Gould, T. M. & Murray, D. B. Respiratory oscillations in yeast: mitochondrial reactive oxygen species, apoptosis and time; a hypothesis. FEMS Yeast Res. 3, 333–339 (2003).
Zeida, A. et al. Catalysis of peroxide reduction by fast reacting protein thiols. Chem. Rev. 119, 10829–10855 (2019).
Perkins, A., Nelson, K. J., Parsonage, D., Poole, L. B. & Karplus, P. A. Peroxiredoxins: guardians against oxidative stress and modulators of peroxide signaling. Trends Biochem. Sci. 40, 435–445 (2015).
Veal, E. A., Underwood, Z. E., Tomalin, L. E., Morgan, B. A. & Pillay, C. S. Hyperoxidation of peroxiredoxins: gain or loss of function? Antioxid. Redox. Signal. 28, 574–590 (2018).
Morgan, B. et al. Real-time monitoring of basal H2O2 levels with peroxiredoxin-based probes. Nat. Chem. Biol. 12, 437–443 (2016).
Nijkamp, J. F. et al. De novo sequencing, assembly and analysis of the genome of the laboratory strain Saccharomyces cerevisiae CEN.PK113-7D, a model for modern industrial biotechnology. Microb. Cell Fact. 11, 36 (2012).
Murray, D. B. et al. in Systems Biology of Metabolic and Signaling Networks (eds Aon, M. et al.) 323–349 (Springer, 2014).
Morgan, B., Sobotta, M. C. & Dick, T. P. Measuring E(GSH) and H2O2 with roGFP2-based redox probes. Free Radic. Biol. Med. 51, 1943–1951 (2011).
Staudacher, V. et al. Redox-sensitive GFP fusions for monitoring the catalytic mechanism and inactivation of peroxiredoxins in living cells. Redox Biol. 14, 549–556 (2018).
Fomenko, D. E. et al. Thiol peroxidases mediate specific genome-wide regulation of gene expression in response to hydrogen peroxide. Proc. Natl Acad. Sci. USA 108, 2729–2734 (2011).
Mulleder, M., Campbell, K., Matsarskaia, O., Eckerstorfer, F. & Ralser, M. Saccharomyces cerevisiae single-copy plasmids for auxotrophy compensation, multiple marker selection, and for designing metabolically cooperating communities. F1000Res. 5, 2351 (2016).
Ewald, J. C. How yeast coordinates metabolism, growth and division. Curr. Opin. Microbiol. 45, 1–7 (2018).
Zhao, G., Chen, Y., Carey, L. & Futcher, B. Cyclin-dependent kinase co-ordinates carbohydrate metabolism and cell cycle in S. cerevisiae. Mol. Cell 62, 546–557 (2016).
Slavov, N., Macinskas, J., Caudy, A. & Botstein, D. Metabolic cycling without cell division cycling in respiring yeast. Proc. Natl Acad. Sci. USA 108, 19090–19095 (2011).
Williamson, D. H. The timing of deoxyribonucleic acid synthesis in the cell cycle of Saccharomyces cerevisiae. J. Cell Biol. 25, 517–528 (1965).
Schneider, B. L., Yang, Q. H. & Futcher, A. B. Linkage of replication to start by the Cdk inhibitor Sic1. Science 272, 560–562 (1996).
Brown, J. D. et al. A peroxiredoxin promotes H2O2 signaling and oxidative stress resistance by oxidizing a thioredoxin family protein. Cell Rep. 5, 1425–1435 (2013).
Calabrese, G. et al. Hyperoxidation of mitochondrial peroxiredoxin limits H2O2-induced cell death in yeast. EMBO J. 38, e101552 (2019).
Stöcker, S., Maurer, M., Ruppert, T. & Dick, T. P. A role for 2-Cys peroxiredoxins in facilitating cytosolic protein thiol oxidation. Nat. Chem. Biol. 14, 148–155 (2018).
Sobotta, M. C. et al. Peroxiredoxin-2 and STAT3 form a redox relay for H2O2 signaling. Nat. Chem. Biol. 11, 64–70 (2015).
Jarvis, R. M., Hughes, S. M. & Ledgerwood, E. C. Peroxiredoxin 1 functions as a signal peroxidase to receive, transduce, and transmit peroxide signals in mammalian cells. Free Radic. Biol. Med. 53, 1522–1530 (2012).
Delaunay, A., Pflieger, D., Barrault, M. B., Vinh, J. & Toledano, M. B. A thiol peroxidase is an H2O2 receptor and redox-transducer in gene activation. Cell 111, 471–481 (2002).
Roger, F. et al. Peroxiredoxin promotes longevity and H2O2-resistance in yeast through redox-modulation of protein kinase A. eLife 9, e60346 (2020).
Kuang, Z., Pinglay, S., Ji, H. & Boeke, J. D. Msn2/4 regulate expression of glycolytic enzymes and control transition from quiescence to growth. eLife 6, 29938 (2017).
Silverman, S. J. et al. Metabolic cycling in single yeast cells from unsynchronized steady-state populations limited on glucose or phosphate. Proc. Natl Acad. Sci. USA 107, 6946–6951 (2010).
Haase, S. B. & Reed, S. I. Evidence that a free-running oscillator drives G1 events in the budding yeast cell cycle. Nature 401, 394–397 (1999).
Orlando, D. A. et al. Global control of cell-cycle transcription by coupled CDK and network oscillators. Nature 453, 944–947 (2008).
Simmons Kovacs, L. A. et al. Cyclin-dependent kinases are regulators and effectors of oscillations driven by a transcription factor network. Mol. Cell 45, 669–679 (2012).
Iraqui, I. et al. Peroxiredoxin Tsa1 is the key peroxidase suppressing genome instability and protecting against cell death in Saccharomyces cerevisiae. PLoS Genet. 5, e1000524 (2009).
Nystrom, T., Yang, J. & Molin, M. Peroxiredoxins, gerontogenes linking aging to genome instability and cancer. Genes Dev. 26, 2001–2008 (2012).
Morgan, D. O. Principles of CDK regulation. Nature 374, 131–134 (1995).
Meyer, A. J. & Dick, T. P. Fluorescent protein-based redox probes. Antioxid. Redox Signal. 13, 621–650 (2010).
Acknowledgements
B.M. acknowledges generous financial support from the Deutsche Forschungsgemeinschaft in the framework of the SPP1710 (MO 2774/2-1) and IRTG1830 programmes, as well as funding from the Technische Universität Kaiserslautern Nachwuchsring and the Forschungsinitiative Rhineland-Pfalz BioComp. G.Y.M. is funded by the Georg Forster Research Fellowship, awarded from the Alexander von Humboldt Foundation. We thank W. Zachariae (Max Planck Institute of Biochemistry) for providing the Clb2 antibody and B. Luke (Institute of Molecular Biology) for providing the Sic1 antibody. We thank T. Dick, J. Herrmann, J. Riemer, L. Prates Roma and F. Hannemann for invaluable discussions and for helpful and insightful comments on the manuscript. We thank V. Nehr for valuable technical assistance.
Author information
Authors and Affiliations
Contributions
B.M., P.S.A. and Z.S. designed all experiments and wrote the manuscript. P.S.A., J.Z., M.M. and S.M. performed metabolic cycle and online roGFP2-Tsa2ΔCR-based measurements, as well as experiments to assess the impact of redox compounds on oxygen and roGFP2-Tsa2ΔCR cycling. They also performed the experiments to assess the impact of genetic and chemical perturbation of the YMC on cell division using flow cytometry-based analysis of DNA content. P.S.A. performed tetrad dissection and the experiments associated with auxin degron-based regulation of peroxiredoxin level. G.Y. performed all budding index experiments and western blot analyses of cell cycle markers. P.S.A., T.M. and B.M. performed the correlation analyses and statistical analyses of all datasets. All authors contributed to data interpretation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Oxygen consumption and cytosolic H2O2 levels oscillate in phase.
a–c, The media oxygen saturation and roGFP2-Tsa2ΔCR oxidation was monitored for three complete cycles in three independent YMC-synchronized cultures of wild-type cells (datasets presented in Fig. 1c and Supplementary Fig. 2b). Autocorrelation analysis of the oxygen and roGFP2-Tsa2ΔCR oxidation revealed robust and in-phase periodicity of the two signals. d, Plot showing time points of the oxygen saturation and roGFP2-Tsa2ΔCR oxidation minima and maxima. Data were fitted by linear regression and significance, as tested by a two-sided ANOVA, revealed p ≈ 0 (4.9 × 10−324).
Extended Data Fig. 2 LOC to HOC switching correlates with probe reduction after diamide treatment.
a–c, RoGFP2-Tsa2ΔCR probe oxidation following the addition of 2 mM diamide towards the end of LOC phase in three independent YMC-synchronized cultures.
Extended Data Fig. 3 Peroxiredoxin deletion perturbs the YMC.
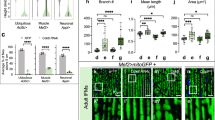
a–d, Oxygen traces to show YMC cycling in either wild-type cells or cells deleted for the genes encoding the indicated peroxiredoxins. Each trace is derived from a completely independent YMC-synchronized culture. e, Cartoon illustrating the subcellular localization of the peroxiredoxins presented in a–d. f, Graph showing the average YMC periods determined from the datasets presented in a–d. n = 3 independent YMC-synchronized cultures. Error bars, mean ± s.d. P values are derived from an unpaired two-tailed Student’s t-test.
Extended Data Fig. 4 Combined deletion of AHP1 and TSA1 is lethal in CEN.PK yeast.
a, Scheme illustrating the mating, sporulation and tetrad dissection procedure. b, Images of tetrad dissection plates for all 33 tetrads dissected. c, Images showing growth of cells from all recovered viable spores on media containing the indicated antibiotics to assess for the presence of the antibiotic resistance cassettes used for gene deletion. d, Table showing the eight possible genotypes and the number of spores recovered with each genotype.
Extended Data Fig. 5 A prolonged switch to HOC phase leads to accumulation of cells with two buds.
a, Representative microscopy images of DAPI stained cells with 2 buds, isolated from YMC-synchronized cultures ~10 hours after addition of 5 mM DTT, the scale bar represents 2 µM. n = 3 independent experimental repeats.
Supplementary information
Supplementary Information
Supplementary Figs. 1–16 and Table 1.
Supplementary Data 1
Statistical source data for all relevant Supplementary figures.
Source data
Source Data Fig. 1
Statistical source data Fig. 1.
Source Data Fig. 2
Statistical source data Fig. 2.
Source Data Fig. 2b
Statistical source data Fig. 2b.
Source Data Fig. 3
Statistical source data Fig. 3.
Source Data Extended Data Fig. 2
Statistical source data Extended Fig. 2.
Source Data Extended Data Fig. 3
Statistical source data Extended Fig. 3.
Rights and permissions
About this article
Cite this article
Amponsah, P.S., Yahya, G., Zimmermann, J. et al. Peroxiredoxins couple metabolism and cell division in an ultradian cycle. Nat Chem Biol 17, 477–484 (2021). https://doi.org/10.1038/s41589-020-00728-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00728-9