Abstract
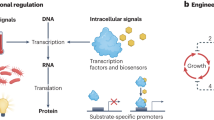
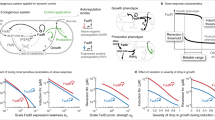
Dynamic regulation is a promising strategy for fine-tuning metabolic fluxes in microbial cell factories. However, few of these synthetic regulatory systems have been developed for central carbon metabolites. Here we created a set of programmable and bifunctional pyruvate-responsive genetic circuits for dynamic dual control (activation and inhibition) of central metabolism in Bacillus subtilis. We used these genetic circuits to design a feedback loop control system that relies on the intracellular concentration of pyruvate to fine-tune the target metabolic modules, leading to the glucaric acid titer increasing from 207 to 527 mg l−1. The designed logic gate-based circuits were enabled by the characterization of a new antisense transcription mechanism in B. subtilis. In addition, a further increase to 802 mg l−1 was achieved by blocking the formation of by-products. Here, the constructed pyruvate-responsive genetic circuits are presented as effective tools for the dynamic control of central metabolism of microbial cell factories.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data produced or analyzed for this study are included in the published article, supplementary information files and source data files, or are available from the corresponding authors upon reasonable request. Sequence data in this article can be found in NCBI under the following accession numbers: pdhR (GenBank ID: 944827), yeeZ (GenBank ID: 946538), udh (GenBank ID: SPD80607.1), gene ino1 (GenBank ID: CAA89448.1). Source data are provided with this paper.
Code availability
The Python code used in this study can be found in Supplementary Note 1.
References
Choi, K. R. et al. Systems metabolic engineering strategies: integrating systems and synthetic biology with metabolic engineering. Trends Biotechnol. 37, 817–837 (2019).
Gu, Y. et al. Advances and prospects of Bacillus subtilis cellular factories: from rational design to industrial applications. Metab. Eng. 50, 109–121 (2018).
Liu, D. et al. Construction, model-based analysis, and characterization of a promoter library for fine-tuned gene expression in Bacillus subtilis. ACS Synth. Biol. 7, 1785–1797 (2018).
Salis, H. M. The ribosome binding site calculator. Methods Enzymol. 498, 19–42 (2011).
Deng, C. et al. Synthetic repetitive extragenic palindromic (REP) sequence as an efficient mRNA stabilizer for protein production and metabolic engineering in prokaryotic cells. Biotechnol. Bioeng. 116, 5–18 (2019).
Trentini, D. B. et al. Arginine phosphorylation marks proteins for degradation by a Clp protease. Nature 539, 48–53 (2016).
Holtz, W. J. & Keasling, J. D. Engineering static and dynamic control of synthetic pathways. Cell 140, 19–23 (2010).
Phan, T. T., Tran, L. T., Schumann, W. & Nguyen, H. D. Development of Pgrac100-based expression vectors allowing high protein production levels in Bacillus subtilis and relatively low basal expression in Escherichia coli. Microb. Cell Fact. 14, 72 (2015).
Bhavsar, A. P., Zhao, X. & Brown, E. D. Development and characterization of a xylose-dependent system for expression of cloned genes in Bacillus subtilis: conditional complementation of a teichoic acid mutant. Appl. Environ. Microbiol. 67, 403–410 (2001).
Guzman, L. M., Belin, D., Carson, M. J. & Beckwith, J. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J. Bacteriol. 177, 4121–4130 (1995).
Xu, P. et al. Design and kinetic analysis of a hybrid promoter-regulator system for malonyl-CoA sensing in Escherichia coli. ACS Chem. Biol. 9, 451–458 (2014).
Peters, G. et al. Development of N-acetylneuraminic acid responsive biosensors based on the transcriptional regulator NanR. Biotechnol. Bioeng. 115, 1855–1865 (2018).
Lehning, C. E., Siedler, S., Ellabaan, M. M. H. & Sommer, M. O. A. Assessing glycolytic flux alterations resulting from genetic perturbations in E. coli using a biosensor. Metab. Eng. 42, 194–202 (2017).
Wang, J. et al. Developing a pyruvate-driven metabolic scenario for growth-coupled microbial production. Metab. Eng. 55, 191–200 (2019).
Bothfeld, W., Kapov, G. & Tyo, K. E. J. A glucose-sensing toggle switch for autonomous, high productivity genetic control. ACS Synth. Biol. 6, 1296–1304 (2017).
Yang, Y. et al. Sensor-regulator and RNAi based bifunctional dynamic control network for engineered microbial synthesis. Nat. Commun. 9, 3043 (2018).
Liang, M. et al. A CRISPR–Cas12a-derived biosensing platform for the highly sensitive detection of diverse small molecules. Nat. Commun. 10, 3672 (2019).
Pelechano, V. & Steinmetz, L. M. Gene regulation by antisense transcription. Nat. Rev. Genet. 14, 880–893 (2013).
Steensels, J. & Verstrepen, K. J. Stop that noise and turn up the antisense transcription. Cell Rep. 15, 2575–2576 (2016).
Quereda, J. J. & Cossart, P. Regulating bacterial virulence with RNA. Annu. Rev. Microbiol. 71, 263–280 (2017).
Brophy, J. A. & Voigt, C. A. Antisense transcription as a tool to tune gene expression. Mol. Syst. Biol. 12, 854 (2016).
Ogasawara, H., Ishida, Y., Yamada, K., Yamamoto, K. & Ishihama, A. PdhR (pyruvate dehydrogenase complex regulator) controls the respiratory electron transport system in Escherichia coli. J. Bacteriol. 189, 5534–5541 (2007).
Ma, W. et al. Metabolic engineering of carbon overflow metabolism of Bacillus subtilis for improved N-acetyl-glucosamine production. Bioresour. Technol. 250, 642–649 (2018).
Panahi, R., Vasheghani-Farahani, E., Shojaosadati, S. A. & Bambai, B. Auto-inducible expression system based on the SigB-dependent ohrB promoter in Bacillus subtilis. Mol. Biol. 48, 970–976 (2014).
Yang, S., Du, G., Chen, J. & Kang, Z. Characterization and application of endogenous phase-dependent promoters in Bacillus subtilis. Appl. Microbiol. Biotechnol. 101, 4151–4161 (2017).
Crampton, N., Bonass, W. A., Kirkham, J., Rivetti, C. & Thomson, N. H. Collision events between RNA polymerases in convergent transcription studied by atomic force microscopy. Nucleic Acids Res. 34, 5416–5425 (2006).
Sesto, N., Wurtzel, O., Archambaud, C., Sorek, R. & Cossart, P. The excludon: a new concept in bacterial antisense RNA-mediated gene regulation. Nat. Rev. Microbiol. 11, 75–82 (2013).
Sneppen, K. et al. A mathematical model for transcriptional interference by RNA polymerase traffic in Escherichia coli. J. Mol. Biol. 346, 399–409 (2005).
Palmer, A. C., Ahlgren-Berg, A., Egan, J. B., Dodd, I. B. & Shearwin, K. E. Potent transcriptional interference by pausing of RNA polymerases over a downstream promoter. Mol. Cell 34, 545–555 (2009).
Walaszek, Z., Szemraj, J., Hanausek, M., Adams, A. K. & Sherman, U. d-Glucaric acid content of various fruits and vegetables and cholesterol-lowering effects of dietary d-glucarate in the rat. Nutr. Res. 16, 673–681 (1996).
Werpy, T. & Peterson, G. (eds) Top Value Added Chemicals from Biomass. Volume 1—Results of Screening for Potential Candidates from Sugars and Synthesis Gas (US Department of Energy, 2004).
Doong, S. J., Gupta, A. & Prather, K. L. J. Layered dynamic regulation for improving metabolic pathway productivity in Escherichia coli. Proc. Natl Acad. Sci. USA 115, 2964–2969 (2018).
Gupta, A., Reizman, I. M., Reisch, C. R. & Prather, K. L. Dynamic regulation of metabolic flux in engineered bacteria using a pathway-independent quorum-sensing circuit. Nat. Biotechnol. 35, 273–279 (2017).
Gu, Y. et al. Synthetic redesign of central carbon and redox metabolism for high yield production of N-acetylglucosamine in Bacillus subtilis. Metab. Eng. 51, 59–69 (2019).
Ma, W. et al. Combinatorial pathway enzyme engineering and host engineering overcomes pyruvate overflow and enhances overproduction of N-acetylglucosamine in Bacillus subtilis. Microb. Cell Fact. 18, 1 (2019).
Tan, S. Z. & Prather, K. L. Dynamic pathway regulation: recent advances and methods of construction. Curr. Opin. Chem. Biol. 41, 28–35 (2017).
San Martin, A. et al. Imaging mitochondrial flux in single cells with a FRET sensor for pyruvate. PLoS ONE 9, e85780 (2014).
Meyer, A. J., Segall-Shapiro, T. H., Glassey, E., Zhang, J. & Voigt, C. A. Escherichia coli “marionette” strains with 12 highly optimized small-molecule sensors. Nat. Chem. Biol. 15, 196–204 (2019).
Mutalik, V. K., Qi, L., Guimaraes, J. C., Lucks, J. B. & Arkin, A. P. Rationally designed families of orthogonal RNA regulators of translation. Nat. Chem. Biol. 8, 447–454 (2012).
Stazic, D., Lindell, D. & Steglich, C. Antisense RNA protects mRNA from RNase E degradation by RNA-RNA duplex formation during phage infection. Nucleic Acids Res. 39, 4890–4899 (2011).
Leistra, A. N., Curtis, N. C. & Contreras, L. M. Regulatory non-coding sRNAs in bacterial metabolic pathway engineering. Metab. Eng. 52, 190–214 (2019).
Hao, N., Palmer, A. C., Ahlgren-Berg, A., Shearwin, K. E. & Dodd, I. B. The role of repressor kinetics in relief of transcriptional interference between convergent promoters. Nucleic Acids Res. 44, 6625–6638 (2016).
Camargo, S., Valladares, A., Flores, E. & Herrero, A. Transcription activation by NtcA in the absence of consensus NtcA-binding sites in an Anabaena heterocyst differentiation gene promoter. J. Bacteriol. 194, 2939–2948 (2012).
Tinberg, C. E. et al. Computational design of ligand-binding proteins with high affinity and selectivity. Nature 501, 212–216 (2013).
Taylor, N. D. et al. Engineering an allosteric transcription factor to respond to new ligands. Nat. Methods 13, 177–183 (2016).
Zhao, E. M. et al. Optogenetic regulation of engineered cellular metabolism for microbial chemical production. Nature 555, 683–687 (2018).
Xu, P., Li, L., Zhang, F., Stephanopoulos, G. & Koffas, M. Improving fatty acids production by engineering dynamic pathway regulation and metabolic control. Proc. Natl Acad. Sci. USA 111, 11299–11304 (2014).
Whiteley, M., Diggle, S. P. & Greenberg, E. P. Corrigendum: progress in and promise of bacterial quorum sensing research. Nature 555, 126 (2018).
Zong, Y. et al. Insulated transcriptional elements enable precise design of genetic circuits. Nat. Commun. 8, 52 (2017).
Moon, T. S., Lou, C., Tamsir, A., Stanton, B. C. & Voigt, C. A. Genetic programs constructed from layered logic gates in single cells. Nature 491, 249–253 (2012).
Zhang, X. Z., Cui, Z. L., Hong, Q. & Li, S. P. High-level expression and secretion of methyl parathion hydrolase in Bacillus subtilis WB800. Appl. Environ. Microbiol. 71, 4101–4103 (2005).
Zheng, S. et al. One-pot two-strain system based on glucaric acid biosensor for rapid screening of myo-inositol oxygenase mutations and glucaric acid production in recombinant cells. Metab. Eng. 49, 212–219 (2018).
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (grant nos 31870069, 31930085 and 21676119), the National Key Research and Development Program of China (grant nos 2018YFA0900300 and 2018YFA0900504), the Fundamental Research Funds for the Central Universities (grant no. JUSRP51713B) and by BBSRC (grant no. BB/R01602X/1).
Author information
Authors and Affiliations
Contributions
X.X., J.C. and L.L. conceived this project and designed the experiments. X.X. and Y.Z. performed experiments. X.L. performed model studies. X.X. wrote the manuscript. Y.L., J.L., G.D., R.L.-A. and L.L. edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–17, Tables 1–4 and Note 1.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Rights and permissions
About this article
Cite this article
Xu, X., Li, X., Liu, Y. et al. Pyruvate-responsive genetic circuits for dynamic control of central metabolism. Nat Chem Biol 16, 1261–1268 (2020). https://doi.org/10.1038/s41589-020-0637-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0637-3
This article is cited by
-
Advances in the optimization of central carbon metabolism in metabolic engineering
Microbial Cell Factories (2023)
-
Construction of the genetic switches in response to mannitol based on artificial MtlR box
Bioresources and Bioprocessing (2023)
-
Microbial cell factories based on filamentous bacteria, yeasts, and fungi
Microbial Cell Factories (2023)
-
Exploring the potential of Bacillus subtilis as cell factory for food ingredients and special chemicals
Microbial Cell Factories (2023)
-
Engineering yeast for bio-production of food ingredients
Systems Microbiology and Biomanufacturing (2023)