Abstract
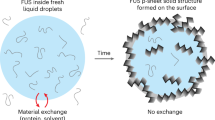
Many RNA-binding proteins undergo liquid–liquid phase separation, which underlies the formation of membraneless organelles, such as stress granules and P-bodies. Studies of the molecular mechanism of phase separation in vitro are hampered by the coalescence and sedimentation of organelle-sized droplets interacting with glass surfaces. Here, we demonstrate that liquid droplets of fused in sarcoma (FUS)—a protein found in cytoplasmic aggregates of amyotrophic lateral sclerosis and frontotemporal dementia patients—can be stabilized in vitro using an agarose hydrogel that acts as a cytoskeleton mimic. This allows their spectroscopic characterization by liquid-phase NMR and electron paramagnetic resonance spectroscopy. Protein signals from both dispersed and condensed phases can be observed simultaneously, and their respective proportions can be quantified precisely. Furthermore, the agarose hydrogel acts as a cryoprotectant during shock-freezing, which facilitates pulsed electron paramagnetic resonance measurements at cryogenic temperatures. Surprisingly, double electron–electron resonance measurements revealed a compaction of FUS in the condensed phase.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data supporting the findings of this study are available within the paper and its Supplementary Information files.
Code availability
MATLAB code used for analysis of the DEER data and ImageJ script used for analysis of the fluorescence microscopy experiments are available from the authors upon reasonable request.
References
Guo, M. et al. Cell volume change through water efflux impacts cell stiffness and stem cell fate. Proc. Natl Acad. Sci. USA 114, E8618–E8627 (2017).
Milo, R. What is the total number of protein molecules per cell volume? A call to rethink some published values. Bioessays 35, 1050–1055 (2013).
Abbondanzieri, E. A. et al. More than just a phase: the search for membraneless organelles in the bacterial cytoplasm. Curr. Genet. 65, 691–694 (2019).
Gomes, E. et al. The molecular language of membraneless organelles. J. Biol. Chem. 294, 7115–7127 (2019).
Mitrea, D. M. et al. Phase separation in biology; functional organization of a higher order. Cell Commun. Signal. 14, 1 (2016).
Protter, D. S. W. et al. Principles and properties of stress granules. Trends Cell Biol. 26, 668–679 (2016).
Bosco, D. A. et al. Mutant FUS proteins that cause amyotrophic lateral sclerosis incorporate into stress granules. Hum. Mol. Genet. 19, 4160–4175 (2010).
Bentmann, E. et al. Requirements for stress granule recruitment of fused in sarcoma (FUS) and TAR DNA-binding protein of 43 kDa (TDP-43). J. Biol. Chem. 287, 23079–23094 (2012).
Lin, Y. et al. Intrinsically disordered sequences enable modulation of protein phase separation through distributed tyrosine motifs. J. Biol. Chem. 292, 19110–19120 (2017).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 174, 688–699 (2018).
Wheeler, J. R. et al. Isolation of yeast and mammalian stress granule cores. Methods 126, 12–17 (2017).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Burke, K. A. et al. Residue-by-residue view of in vitro FUS granules that bind the C-terminal domain of RNA polymerase II. Mol. Cell 60, 231–241 (2015).
Murakami, T. et al. ALS/FTD mutation-induced phase transition of FUS liquid droplets and reversible hydrogels into irreversible hydrogels impairs RNP granule function. Neuron 88, 678–690 (2015).
Shang, Y. et al. Mechanisms of FUS mutations in familial amyotrophic lateral sclerosis. Brain Res. 1647, 65–78 (2016).
Murthy, A. C. et al. Molecular interactions underlying liquid–liquid phase separation of the FUS low-complexity domain. Nat. Struct. Mol. Biol. 26, 637–648 (2019).
Boeynaems, S. et al. Protein phase separation: a new phase in cell biology. Trends Cell Biol. 28, 420–435 (2018).
Brady, J. P. et al. Structural and hydrodynamic properties of an intrinsically disordered region of a germ cell-specific protein on phase separation. Proc. Natl Acad. Sci. USA 114, E8194–E8203 (2017).
Feric, M. & Brangwynne, C. P. A nuclear F-actin scaffold stabilizes ribonucleoprotein droplets against gravity in large cells. Nat. Cell Biol. 15, 1253–1259 (2013).
Pastore, A. et al. Peptides and proteins in a confined environment: NMR spectra at natural isotopic abundance. J. Pept. Sci. 13, 342–347 (2007).
Lameiras, P. et al. Polar mixture analysis by NMR under spin diffusion conditions in viscous sucrose solution and agarose gel. Faraday Discuss. 218, 233–246 (2019).
Mitrev, Y. et al. NMR analysis of weak molecular interactions using slice-selective experiments via study of concentration gradients in agar gels. Chem. Commun. 52, 5418–5420 (2016).
McSwiggen, D. T. et al. Evaluating phase separation in live cells: diagnosis, caveats and functional consequences. Genes Dev. 33, 1619–1634 (2019).
Tripathi, V. et al. SRSF1 regulates the assembly of pre-mRNA processing factors in nuclear speckles. Mol. Biol. Cell 23, 3694–3706 (2012).
Li, P. et al. Phase transitions in the assembly of multivalent signalling proteins. Nature 483, 336–340 (2012).
Nott, T. J. et al. Phase transition of a disordered nuage protein generates environmentally responsive membraneless organelles. Mol. Cell 57, 936–947 (2015).
Elbaum-Garfinkle, S. et al. The disordered P granule protein LAF-1 drives phase separation into droplets with tunable viscosity and dynamics. Proc. Natl Acad. Sci. USA 112, 7189–7194 (2015).
Klare, J. P. et al. Spin labeling EPR. Photosynth. Res. 102, 377–390 (2009).
Baust, J. G. et al. Advances in Biopreservation, Ch. 8 (CRC Press, 2006).
Jeschke, G. DEER distance measurements on proteins. Annu. Rev. Phys. Chem. 63, 419–446 (2012).
Borbat, P. P. et al. Improved sensitivity for long-distance measurements in biomolecules: five-pulse double electron–electron resonance. J. Phys. Chem. Lett. 4, 170–175 (2013).
Breitgoff, F. D. et al. Artefact suppression in 5-pulse double electron electron resonance for distance distribution measurements. Phys. Chem. Chem. Phys. 19, 15766–15779 (2017).
Fábregas Ibáñez, L. et al. DeerLab: a comprehensive toolbox for analyzing dipolar EPR spectroscopy data. Magn. Reson. Discuss. 1, 209–224 (2020).
Murray, D. T. et al. Structure of FUS protein fibrils and its relevance to self-assembly and phase separation of low-complexity domains. Cell 171, 615–627 (2017).
Hughes, M. P. et al. Atomic structures of low-complexity protein segments reveal kinked β sheets that assemble networks. Science 359, 698–701 (2018).
Luo, F. et al. Atomic structures of FUS LC domain segments reveal bases for reversible amyloid fibril formation. Nat. Struct. Mol. Biol. 25, 341–346 (2018).
Reichheld, S. E. et al. Direct observation of structure and dynamics during phase separation of an elastomeric protein. Proc. Natl Acad. Sci. USA 114, E4408–E4415 (2017).
Kim, T. H. et al. Phospho-dependent phase separation of FMRP and CAPRIN1 recapitulates regulation of translation and deadenylation. Science 365, 825–829 (2019).
Tsang, B. et al. Phosphoregulated FMRP phase separation models activity-dependent translation through bidirectional control of mRNA granule formation. Proc. Natl Acad. Sci. USA 116, 4218–4227 (2019).
Alberti, S. et al. A user’s guide for phase separation assays with purified proteins. J. Mol. Biol. 430, 4806–4820 (2018).
Lin, Y. et al. Formation and maturation of phase-separated liquid droplets by RNA-binding proteins. Mol. Cell 60, 208–219 (2015).
Murthy, A. C. et al. The (un)structural biology of biomolecular liquid–liquid phase separation using NMR spectroscopy. J. Biol. Chem. 295, 2375–2384 (2020).
Lucas, L. H. et al. Measuring ligand–protein binding using NMR diffusion experiments. Concepts Magn. Reson. A 20, 24–41 (2004).
Souquere, S. et al. Unravelling the ultrastructure of stress granules and associated P-bodies in human cells. J. Cell Sci. 122, 3619–3626 (2009).
Dorn, G. et al. Structural modeling of protein–RNA complexes using crosslinking of segmentally isotope-labeled RNA and MS/MS. Nat. Methods 14, 487–490 (2017).
Bolte, S. & Cordelières, F. P. A guided tour into subcellular colocalization analysis in light microscopy. J. Microsc. 224, 213–232 (2006).
Gromov, I. et al. A Q-band pulse EPR/ENDOR spectrometer and the implementation of advanced one- and two-dimensional pulse EPR methodology. J. Magn. Reson. 149, 196–203 (2001).
Tschaggelar, R. et al. Cryogenic 35-GHz pulse ENDOR probe head accommodating large sample sizes: performance and applications. J. Magn. Reson. 200, 81–87 (2009).
Jeschke, G., Sajid, M., Schulte, M. & Godt, A. Three-spin correlations in double electron–electron resonance. Phys. Chem. Chem. Phys. 11, 6580–6591 (2009).
Acknowledgements
This work was supported by the Swiss National Science Foundation with Sinergia grant no. CR-SII5_170976, NCCR RNA & Disease and an EMBO long-term postdoctoral fellowship. E.K. acknowledges support from Volkswagen Foundation Experiment! grant no. 95664. We thank K. Weis and M. Hondele for sharing microscope instruments and expertise, and A. Gossert and I. Ritsch for valuable discussions.
Author information
Authors and Affiliations
Contributions
L.E. and L.E.-H. designed the project. L.E. performed the NMR, turbidity and microscopy experiments. L.E.-H. performed the EPR and microscopy experiments. F.F.D. and M.Y. assisted with the experimental design for NMR and EPR, respectively. T.d.V. and C.K.X.N. provided protein samples. L.F.I. assisted with DEER data analysis. S.M. and E.K. performed fluorescence microscopy and analysis. G.J. and F.H.-T.A. provided infrastructure, financial support and overall supervision of the project. L.E. and L.E.-H. wrote the manuscript with support from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 FUS NTD droplets.
Light microscope image of FUS NTD droplets formed upon dilution of urea. Representative image from four independent experiments. Scale bar: 10 μm.
Extended Data Fig. 2 FUS NTD droplets in presence and absence of agarose.
a,b) Upfield region of 1H 1D and 1H-15N HSQC NMR spectra of FUS NTD in absence and presence of 0.5% agarose hydrogel showing comparable linewidths. c) Unattenuated DOSY signal of FUS NTD biphasic sample at different agarose concentrations.
Extended Data Fig. 3 FUS NTD droplet fraction quantification correlates with turbidity.
a) Aliphatic proton region of 1H 1D spectra in the presence and in absence of urea. Color scale (magenta → blue → red) corresponds to spectra recorded with increasing diffusion gradient strength. b) Integrals of the spectral region in (a) as a function of diffusion gradient strength normalized to the integral at the lowest gradient strength. The fraction of the slowly diffusing component obtained by averaging data points obtained at >30 G/cm is 19%. c) Droplet size comparison among different total protein concentrations determined by fluorescence microscopy. d) Correlation of final percentage of unattenuated signal from the DOSY experiments with sample turbidity as function of protein concentration. Error bars indicate standard deviation and dots the mean from three independent experiments.
Extended Data Fig. 4 Protein phase exchange during NMR DOSY experiment.
Estimation of protein molecules remaining in the same phase during the diffusion time of our diffusion NMR experiments (0.08 s). In vitro half time, defined as ln(2)/(exchange rate), of FUS NTD in the droplet phase, as measured by FRAP experiments14, is marked with a yellow dashed line. The range of accessible experimental diffusion times in NMR experiments is highlighted with diagonal stripes.
Extended Data Fig. 5 Effect of agarose hydrogel on stability of liquid droplet of FUS, DDX4, PTBP1 and SRSF1.
Brightfield microscope images of liquid droplets from different proteins hours post preparation in the presence and absence of agarose hydrogel. Representative images from three independent experiments. (Scale bar: 10 μm).
Extended Data Fig. 6 DDX4 droplets in 0.5% agarose hydrogel.
a) Time progression of sample turbidity in the absence (black) and presence (red) of agarose hydrogel. Error bars indicate standard deviation and dots the mean from three independent experiments. b) Overlay of 1H 1D DOSY spectra of dispersed (top) and biphasic (bottom) sample in agarose. Increasing gradient strength is visualized by a color gradient magenta → blue → red. c) Integral of the spectral region shown in (b) normalized to the integral at the lowest gradient strength as a function of the gradient strength. Black and red data denote dispersed and biphasic sample in agarose, respectively. d) Integrals of 1H 1D DOSY spectra as function of gradient strength at different DDX4 concentrations. e) Correlation of final percentage of unattenuated signal from the DOSY experiments shown in (e) with sample turbidity as function of protein concentration. Error bars indicate standard deviation and dots the mean from three independent experiments.
Extended Data Fig. 7 Diffusion NMR experiments on SRSF1 and PTBP1.
a, b) Integrated normalized aliphatic spectral region plotted vs. gradient strength for dispersed (black) and biphasic (red) PTBP1 and SRSF1 respectively. c, d) Amide and aliphatic regions of 1H 1D NMR spectra of biphasic PTBP1 and SRSF1 in agarose. e, f) 1H-15N HSQC spectrum of biphasic PTBP1 and SRSF1 in agarose.
Extended Data Fig. 8 Brightfield microscope images of FUS NTD.
FUS NTD in a) 50% glycerol and in b) 50% PEG shows no phase separation under buffer conditions where liquid phase separation is otherwise observed, as reflected in c). Representative images from three independent experiments. Scale bar: 30 μm.
Extended Data Fig. 9 DEER experiments on the NTD of FUS.
Primary DEER data and corresponding distance distributions using a model-free fit with Tikhonov regularization (black) and a Gaussian distribution (red) of A10C S29C R1 and A105C G128C R1 in a,b) the dispersed state with 3 M urea, in c, d) the dispersed state with 0.6 M urea, and e, f) in the bulk phase. The experimental data are displayed as black dots, the fit as a solid line and the 95% confidence interval obtained via 1000 bootstrap samples as shaded area. The Tikhonov regularization and fit with unimodal Gaussian function with variable mean and width lead to very similar distribution shapes. Total modulation depth Δ as a function of nominal inversion efficiency λnominal for A10C S29C R1 in g) the biphasic state and h) the bulk phase, and A105C G128C R1 in i) the biphasic state and j) the bulk phase. Good fits of the experimental data (black dots) are obtained using a model for two spins (solid red line), which confirms that the spin dilution employed is sufficient to avoid intermolecular distance contributions in the DEER experiment.
Extended Data Fig. 10 Schematics of the fitting algorithm employed for the analysis of the biphasic DEER measurements.
Red corresponds to spin-labeled protein in the dispersed phase, blue is spin-labeled protein in the condensed phase, and gray represents unlabeled, and therefore EPR-silent, protein. The total signal of the biphasic sample is displayed in black.
Supplementary information
Supplementary Information
Supplementary Table 1 and Fig. 1.
Rights and permissions
About this article
Cite this article
Emmanouilidis, L., Esteban-Hofer, L., Damberger, F.F. et al. NMR and EPR reveal a compaction of the RNA-binding protein FUS upon droplet formation. Nat Chem Biol 17, 608–614 (2021). https://doi.org/10.1038/s41589-021-00752-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00752-3
This article is cited by
-
An aberrant fused in sarcoma liquid droplet of amyotrophic lateral sclerosis pathological variant, R495X, accelerates liquid–solid phase transition
Scientific Reports (2024)
-
A solid beta-sheet structure is formed at the surface of FUS droplets during aging
Nature Chemical Biology (2024)
-
Large dynamics of a phase separating arginine-glycine-rich domain revealed via nuclear and electron spins
Nature Communications (2024)
-
Advances in mass spectrometry to unravel the structure and function of protein condensates
Nature Protocols (2023)
-
Single-molecule FRET unmasks structural subpopulations and crucial molecular events during FUS low-complexity domain phase separation
Nature Communications (2023)