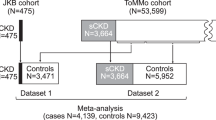
Abstract
Chronic kidney disease (CKD) is a common complex condition associated with high morbidity and mortality. Polygenic prediction could enhance CKD screening and prevention; however, this approach has not been optimized for ancestrally diverse populations. By combining APOL1 risk genotypes with genome-wide association studies (GWAS) of kidney function, we designed, optimized and validated a genome-wide polygenic score (GPS) for CKD. The new GPS was tested in 15 independent cohorts, including 3 cohorts of European ancestry (n = 97,050), 6 cohorts of African ancestry (n = 14,544), 4 cohorts of Asian ancestry (n = 8,625) and 2 admixed Latinx cohorts (n = 3,625). We demonstrated score transferability with reproducible performance across all tested cohorts. The top 2% of the GPS was associated with nearly threefold increased risk of CKD across ancestries. In African ancestry cohorts, the APOL1 risk genotype and polygenic component of the GPS had additive effects on the risk of CKD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The final formulation of the GPS for CKD along with the standardized metrics of performance have been deposited in the PGS catalog at https://www.pgscatalog.org/publication/PGP000269/. The UKBB genotype and phenotype data are available through the UKBB web portal at https://www.ukbiobank.ac.uk/. The eMERGE-III imputed genotype and phenotype data are available through the database of Genotypes and Phenotypes (dbGAP) under accession no. phs001584.v2.p2. The BioMe genotype datasets used in this study were generated by Regeneron and are not publicly available. However, the data will be made available for the purposes of replicating the results by contacting the corresponding author and through appropriate collaboration and/or data sharing agreements. The WPC and REGARDS imputed genotype and phenotype data are available through dbGAP under accession nos. phs000708.v1.p1 and phs002719.v1.p1, respectively. The GenHAT cohort is also available on dbGAP under accession no. phs002716.v1.p1. The HyperGEN cohort has been sequenced by the TOPMed consortium; WGS data along with phenotype data are available through dbGAP under accession no. phs001293.v3.p1. Minimum testing datasets with the GPS, CKD outcome, and a set of essential clinical covariates for each cohort are also available when consistent with the consent given by the participants and can be requested directly from the corresponding author with a 2–4-week response time frame. Because these datasets contain clinical data, access to them may require a data use agreement.
Code availability
The CKD phenotype software is available from the Phenotype Knowledge Database at https://phekb.org/phenotype/chronic-kidney-disease. The CKD GPS score equation is available through the PGS catalog at https://www.pgscatalog.org/publication/PGP000269/ and through our laboratory website at http://www.columbiamedicine.org/divisions/kiryluk/study_GPS_CKD.php.
References
Coresh, J. et al. Prevalence of chronic kidney disease in the United States. JAMA 298, 2038–2047 (2007).
Naghavi, M. et al. Global, regional, and national age-sex specific mortality for 264 causes of death, 1980–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet 390, 1151–1210 (2017).
Chronic Kidney Disease in the United States (Centers for Disease Control and Prevention, 2022); https://www.cdc.gov/kidneydisease/publications-resources/ckd-national-facts.html
Shang, N. et al. Medical records-based chronic kidney disease phenotype for clinical care and “big data” observational and genetic studies. NPJ Digit. Med. 4, 70 (2021).
Fox, C. S. et al. Genomewide linkage analysis to serum creatinine, GFR, and creatinine clearance in a community-based population: the Framingham Heart Study. J. Am. Soc. Nephrol. 15, 2457–2461 (2004).
Langefeld, C. D. et al. Heritability of GFR and albuminuria in Caucasians with type 2 diabetes mellitus. Am. J. Kidney Dis. 43, 796–800 (2004).
Satko, S. G. & Freedman, B. I. The familial clustering of renal disease and related phenotypes. Med. Clin. North. Am. 89, 447–456 (2005).
Groopman, E. E. et al. Diagnostic utility of exome sequencing for kidney disease. N. Engl. J. Med. 380, 142–151 (2019).
Lata, S. Whole-exome sequencing in adults with chronic kidney disease: a pilot study. Ann. Intern. Med. 168, 100–109 (2018).
Köttgen, A. et al. Multiple loci associated with indices of renal function and chronic kidney disease. Nat. Genet. 41, 712–717 (2009).
Wuttke, M. et al. A catalog of genetic loci associated with kidney function from analyses of a million individuals. Nat. Genet. 51, 957–972 (2019).
Genovese, G. et al. Association of trypanolytic ApoL1 variants with kidney disease in African Americans. Science 329, 841–845 (2010).
Parsa, A. et al. APOL1 risk variants, race, and progression of chronic kidney disease. N. Engl. J. Med. 369, 2183–2196 (2013).
Thomson, R. et al. Evolution of the primate trypanolytic factor APOL1. Proc. Natl Acad. Sci. USA 111, E2130–E2139 (2014).
Ko, W.-Y. et al. Identifying Darwinian selection acting on different human APOL1 variants among diverse African populations. Am. J. Hum. Genet. 93, 54–66 (2013).
Nadkarni, G. N. et al. Worldwide frequencies of APOL1 renal risk variants. N. Engl. J. Med. 379, 2571–2572 (2018).
Gladding, P. A., Legget, M., Fatkin, D., Larsen, P. & Doughty, R. Polygenic risk scores in coronary artery disease and atrial fibrillation. Heart Lung Circ. 29, 634–640 (2020).
Khera, A. V. et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 50, 1219–1224 (2018).
Läll, K., Mägi, R., Morris, A., Metspalu, A. & Fischer, K. Personalized risk prediction for type 2 diabetes: the potential of genetic risk scores. Genet. Med. 19, 322–329 (2017).
Hoffmann, T. J. et al. Genome-wide association analyses using electronic health records identify new loci influencing blood pressure variation. Nat. Genet. 49, 54–64 (2017).
Ehret, G. B. et al. The genetics of blood pressure regulation and its target organs from association studies in 342,415 individuals. Nat. Genet. 48, 1171–1184 (2016).
Khera, A. V. et al. Polygenic prediction of weight and obesity trajectories from birth to adulthood. Cell 177, 587–596.e9 (2019).
Weinberger, D. R. Polygenic risk scores in clinical schizophrenia research. Am. J. Psychiatry 176, 3–4 (2019).
Reginsson, G. W. et al. Polygenic risk scores for schizophrenia and bipolar disorder associate with addiction. Addict. Biol. 23, 485–492 (2018).
Power, R. A. et al. Polygenic risk scores for schizophrenia and bipolar disorder predict creativity. Nat. Neurosci. 18, 953–955 (2015).
Aly, M. et al. Polygenic risk score improves prostate cancer risk prediction: results from the Stockholm-1 cohort study. Eur. Urol. 60, 21–28 (2011).
Fritsche, L. G. et al. Association of polygenic risk scores for multiple cancers in a phenome-wide study: results from the Michigan Genomics Initiative. Am. J. Hum. Genet. 102, 1048–1061 (2018).
Jeon, J. et al. Determining risk of colorectal cancer and starting age of screening based on lifestyle, environmental, and genetic factors. Gastroenterology 154, 2152–2164.e19 (2018).
Huyghe, J. R. et al. Discovery of common and rare genetic risk variants for colorectal cancer. Nat. Genet. 51, 76–87 (2019).
Mavaddat, N. et al. Polygenic risk scores for prediction of breast cancer and breast cancer subtypes. Am. J. Hum. Genet. 104, 21–34 (2019).
Seibert, T. M. et al. Polygenic hazard score to guide screening for aggressive prostate cancer: development and validation in large scale cohorts. BMJ 360, j5757 (2018).
Martin, A. R. et al. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat. Genet. 51, 584–591 (2019).
Wand, H. et al. Improving reporting standards for polygenic scores in risk prediction studies. Nature 591, 211–219 (2021).
Zhang, J., Thio, C. H. L., Gansevoort, R. T. & Snieder, H. Familial aggregation of CKD and heritability of kidney biomarkers in the general population: the Lifelines Cohort Study. Am. J. Kidney Dis. 77, 869–878 (2021).
Inker, L. A. et al. New creatinine- and cystatin C-based equations to estimate GFR without race. N. Engl. J. Med. 385, 1737–1749 (2021).
Yu, Z. et al. Polygenic risk scores for kidney function and their associations with circulating proteome, and incident kidney diseases. J. Am. Soc. Nephrol. 32, 3161–3173 (2021).
Polubriaginof, F., Tatonetti, N. P. & Vawdrey, D. K. An assessment of family history information captured in an electronic health record. AMIA Annu. Symp. Proc. 2015, 2035–2042 (2015).
Tada, H. et al. Risk prediction by genetic risk scores for coronary heart disease is independent of self-reported family history. Eur. Heart J. 37, 561–567 (2016).
Timmerman, N. et al. Family history and polygenic risk of cardiovascular disease: independent factors associated with secondary cardiovascular events in patients undergoing carotid endarterectomy. Atherosclerosis 307, 121–129 (2020).
Hindy, G. et al. Genome-wide polygenic score, clinical risk factors, and long-term trajectories of coronary artery disease. Arterioscler. Thromb. Vasc. Biol. 40, 2738–2746 (2020).
Inouye, M. et al. Genomic risk prediction of coronary artery disease in 480,000 adults: implications for primary prevention. J. Am. Coll. Cardiol. 72, 1883–1893 (2018).
Lee, A. et al. BOADICEA: a comprehensive breast cancer risk prediction model incorporating genetic and nongenetic risk factors. Genet. Med. 21, 1708–1718 (2019).
Orlando, L. A. et al. Development and validation of a primary care-based family health history and decision support program (MeTree). N. C. Med. J. 74, 287–296 (2013).
Fahed, A. C. et al. Polygenic background modifies penetrance of monogenic variants for tier 1 genomic conditions. Nat. Commun. 11, 3635 (2020).
Zanoni, F. & Kiryluk, K. Genetic background and transplantation outcomes: insights from genome-wide association studies. Curr. Opin. Organ Transpl. 25, 35–41 (2020).
Hellwege, J. N. et al. Mapping eGFR loci to the renal transcriptome and phenome in the VA Million Veteran Program. Nat. Commun. 10, 3842 (2019).
Sheng, X. et al. Mapping the genetic architecture of human traits to cell types in the kidney identifies mechanisms of disease and potential treatments. Nat. Genet. 53, 1322–1333 (2021).
Neugut, Y. D., Mohan, S., Gharavi, A. G. & Kiryluk, K. Cases in precision medicine: APOL1 and genetic testing in the evaluation of chronic kidney disease and potential transplant. Ann. Intern. Med. 171, 659–664 (2019).
Sirugo, G., Williams, S. M. & Tishkoff, S. A. The missing diversity in human genetic studies. Cell 177, 26–31 (2019).
Delanaye, P. et al. CKD: a call for an age-adapted definition. J. Am. Soc. Nephrol. 30, 1785–1805 (2019).
Teumer, A. et al. Genome-wide association meta-analyses and fine-mapping elucidate pathways influencing albuminuria. Nat. Commun. 10, 4130 (2019).
Kiryluk, K. et al. Discovery of new risk loci for IgA nephropathy implicates genes involved in immunity against intestinal pathogens. Nat. Genet. 46, 1187–1196 (2014).
Xie, J. et al. The genetic architecture of membranous nephropathy and its potential to improve non-invasive diagnosis. Nat. Commun. 11, 1600 (2020).
Sohail, M. et al. Polygenic adaptation on height is overestimated due to uncorrected stratification in genome-wide association studies. Elife 8, e39702 (2019).
Vyas, D. A., Eisenstein, L. G. & Jones, D. S. Hidden in plain sight—reconsidering the use of race correction in clinical algorithms. N. Engl. J. Med. 383, 874–882 (2020).
Delgado, C. et al. A unifying approach for GFR estimation: recommendations of the NKF-ASN Task Force on reassessing the inclusion of race in diagnosing kidney disease. J. Am. Soc. Nephrol. 32, 2994–3015 (2021).
Khan, A. et al. Medical records-based genetic studies of the complement system. J. Am. Soc. Nephrol. 32, 2031–2047 (2021).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Abraham, G. & Inouye, M. Fast principal component analysis of large-scale genome-wide data. PLoS ONE 9, e93766 (2014).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Belbin, G. M. et al. Toward a fine-scale population health monitoring system. Cell 184, 2068–2083.e11 (2021).
Howard, V. J. et al. The reasons for geographic and racial differences in stroke study: objectives and design. Neuroepidemiology 25, 135–143 (2005).
Williams, R. R. et al. NHLBI family blood pressure program: methodology and recruitment in the HyperGEN network. Ann. Epidemiol. 10, 389–400 (2000).
Limdi, N. A. et al. Influence of kidney function on risk of supratherapeutic international normalized ratio-related hemorrhage in warfarin users: a prospective cohort study. Am. J. Kidney Dis. 65, 701–709 (2015).
Arnett, D. K. et al. Pharmacogenetic approaches to hypertension therapy: design and rationale for the Genetics of Hypertension Associated Treatment (GenHAT) study. Pharmacogenomics J. 2, 309–317 (2002).
Furberg, C. D. et al. Major cardiovascular events in hypertensive patients randomized to doxazosin vs chlorthalidone: the antihypertensive and lipid-lowering treatment to prevent heart attack trial (ALLHAT). ALLHAT Collaborative Research Group. JAMA 283, 1967–1975 (2000).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Levey, A. S. & Stevens, L. A. Estimating GFR using the CKD Epidemiology Collaboration (CKD-EPI) creatinine equation: more accurate GFR estimates, lower CKD prevalence estimates, and better risk predictions. Am. J. Kidney Dis. 55, 622–627 (2010).
Kidney Disease: Improving Global Outcomes (KDIGO) Chronic Kidney Disease Work Group KDIGO 2012 clinical practice guideline for the evaluation and management of chronic kidney disease. Kidney Int. Suppl. 3, 1–150 (2013).
Vilhjálmsson, B. J. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592 (2015).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Khera, A. V. et al. Whole-genome sequencing to characterize monogenic and polygenic contributions in patients hospitalized with early-onset myocardial infarction. Circulation 139, 1593–1602 (2019).
Acknowledgements
This work was funded by the National Human Genome Research Institute eMERGE-IV grant nos. 2U01HG008680-05, 1U01HG011167-01 and 1U01HG011176-01. Additional sources of funding included grant nos. UG3DK114926 (K.K.), RC2DK116690 (K.K.), R01LM013061 (C.W., K.K.), K25DK128563 (A.K.), UL1TR001873 (A.K., K.K.), R01HL151855 (J.B.M.) and UM1DK078616 (J.B.M.). The parent REGARDS study was supported by cooperative agreement no. U01NS041588 cofunded by the National Institute of Neurological Disorders and Stroke and the National Institute on Aging, the National Institutes of Health (NIH) and the Department of Health and Human Services. The HyperGEN (R01HL055673), GenHAT (R01HL123782) and WPC (R01HL092173, K24HL133373) studies were all supported by the NHLBI. Parts of this study were conducted using the UKBB resource under UKBB project no. 41849. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
A.K. and K.K. conceptualized the study. A.K., K.K., M.C.T., A.P., V.S., R.N., A.C.J., E.M., C.K., N.L., I.I.L., T.G., M.R.I., H.K.T. and E.E.K. devised the methodology and carried out the genetic data analysis. N.S., C.Li., G.H., C.W. and G.N. carried out the e-phenotyping. O.D., I.J.K., D.J.S., E.K., J.B.M., J.W.S., C.La., D.R.C., G.P.J., P.K.B., J.N.H., P.C., L.R.T., A.G.G., W.K.C., G.H. and C.W. provided the eMERGE-III data contributions. G.N., J.H.C., N.S.A-H. and E.E.K. provided the BioMe data contributions. M.R.I., H.K.T. and N.A.L. provided the UAB data contributions. A.K. and A.F. managed the project. K.K. provided overall supervision. A.K. and K.K. wrote the original manuscript draft. A.K., N.S., E.M., A.G.G., T.G., J.B.M., D.R.C., J.N.H., I.I.L., G.H., M.R.I., H.K.T., E.E.K., N.A.L. and K.K. reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Charles Rotimi, Andrew Mallett and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Anna Maria Ranzoni, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
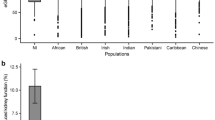
Extended Data Fig. 1 Distribution of risk allele frequencies (RAF) and their effect sizes for the variants included in the GPS.
Distribution of risk allele frequencies (RAF) and their effect sizes for the variants included in the GPS (a) comparison of RAF distributions for the risk variants included in the CKD GPS demonstrates higher frequency of rare (RAF < 0.01) and common (RAF > 0.99) risk alleles in African compared to European genomes (based on 1000 G reference populations); this may be explained by the exclusion of variants with MAF < 0.01 in European discovery GWAS; (b) highly skewed effect size (weight) distribution for the variants included in the GPS for CKD; (c) Distribution of RAF difference (AFR-EUR) demonstrating higher average frequency of risk alleles in African genomes (mean RAF difference = 0.002) and a slight rightward shift of the RAF difference distribution from the expected mean of 0; (d) Mean RAF difference (AFR-EUR) as a function of effect size binned into three categories (high, intermediate, and low) based on the observed distribution of effects sizes in panel b, demonstrating that the risk alleles with larger effect size have higher average frequency in African compared to European genomes. EUR: European (N = 503) and AFR: African (N = 661). The bars represent 95% confidence intervals around the mean RAF difference estimate for each bin; two-sided P-values were calculated using t-test.
Extended Data Fig. 2 Risk score distributions in eMERGE-III (N = 22,453) and UKBB (N = 77,584) validation datasets.
Risk score distributions in eMERGE-III (N = 22,453) and UKBB (N = 77,584) validation datasets: (a) the distribution of raw polygenic score without APOL1 in UKBB by ancestry; (b) the distribution of ancestry-adjusted polygenic score (method 1: mean-adjusted) in UKBB by ancestry; (c) the distribution of ancestry-adjusted polygenic score (method 2: mean and variance-adjusted) in UKBB by ancestry. Panels (d), (e) and (f) show the same analyses for the eMERGE-III dataset, respectively.
Extended Data Fig. 3 Final GPS calibration analysis in eMERGE-III cohorts combined (N = 22,453).
Final GPS calibration analysis in eMERGE-III cohorts combined (N = 22,453): predicted risk (X-axis) as a function of the observed risk (Y-axis) in the multiethnic eMERGE-III dataset after ancestry adjustment with (a) method 1 and (b) method 2. The bars represent 95% confidence intervals.
Extended Data Fig. 4 Distributions of the raw (non-standardized) genome-wide polygenic score (GPS) by Yu et al. in the eMERGE-III validation datasets by ancestry.
Distributions of the raw (non-standardized) genome-wide polygenic score (GPS) by Yu et al. in the eMERGE-III validation datasets by ancestry.
Extended Data Fig. 5 PCA projections of the study participants from the UKBB (top) and eMERGE-III (bottom) against the 1000 G reference populations.
PCA projections of the study participants from the UKBB (top) and eMERGE-III (bottom) against the 1000 G reference populations: (a) UKBB (N = 77,584) and (b) eMERGE-III (N = 22,453) participants plotted against the reference 1000 G populations (N = 2,504); (b, e) plotted by self-reported race/ethnicity; and (c, f) plotted by final ancestry group assignment. X-axis: PC1; Y-axis: PC2; AFR: African; AMR: Admixed American; EAS: East Asian; EUR: European; and SAS: South Asian.
Supplementary information
Supplementary Information
Supplementary Tables 1–11.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khan, A., Turchin, M.C., Patki, A. et al. Genome-wide polygenic score to predict chronic kidney disease across ancestries. Nat Med 28, 1412–1420 (2022). https://doi.org/10.1038/s41591-022-01869-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-022-01869-1
This article is cited by
-
Recent advances in polygenic scores: translation, equitability, methods and FAIR tools
Genome Medicine (2024)
-
Principles and methods for transferring polygenic risk scores across global populations
Nature Reviews Genetics (2024)
-
A new era in the science and care of kidney diseases
Nature Reviews Nephrology (2024)
-
Diet quality in relation to kidney function and its potential interaction with genetic risk of kidney disease among Dutch post-myocardial infarction patients
European Journal of Nutrition (2024)
-
Selection, optimization and validation of ten chronic disease polygenic risk scores for clinical implementation in diverse US populations
Nature Medicine (2024)