Abstract
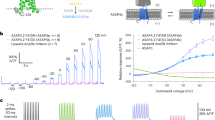
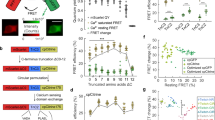
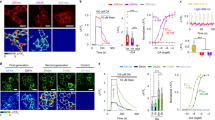
Current techniques for monitoring GABA (γ-aminobutyric acid), the primary inhibitory neurotransmitter in vertebrates, cannot follow transients in intact neural circuits. To develop a GABA sensor, we applied the design principles used to create the fluorescent glutamate receptor iGluSnFR. We used a protein derived from a previously unsequenced Pseudomonas fluorescens strain and performed structure-guided mutagenesis and library screening to obtain intensity-based GABA sensing fluorescence reporter (iGABASnFR) variants. iGABASnFR is genetically encoded, detects GABA release evoked by electric stimulation of afferent fibers in acute brain slices and produces readily detectable fluorescence increases in vivo in mice and zebrafish. We applied iGABASnFR to track mitochondrial GABA content and its modulation by an anticonvulsant, swimming-evoked, GABA-mediated transmission in zebrafish cerebellum, GABA release events during interictal spikes and seizures in awake mice, and found that GABA-mediated tone decreases during isoflurane anesthesia.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Code availability
All analysis code used in this study is available upon request.
References
Fritschy, J.-M. & Panzanelli, P. GABAA receptors and plasticity of inhibitory neurotransmission in the central nervous system. Eur. J. Neurosci. 39, 1845–1865 (2014).
Padgett, C. L. & Slesinger, P. A. GABAB receptor coupling to G-proteins and ion channels. Adv. Pharm 58, 123–147 (2010).
Sodickson, D. L. & Bean, B. P. GABAB receptor-activated inwardly rectifying potassium current in dissociated hippocampal CA3 neurons. J. Neurosci. 16, 6374–6385 (1996).
Macdonald, R. L. & Olsen, R. W. GABAA receptor channels. Annu. Rev. Neurosci. 17, 569–602 (1994).
van der Zeyden, M., Oldenziel, W. H., Rea, K., Cremers, T. I. & Westerink, B. H. Microdialysis of GABA and glutamate: analysis, interpretation and comparison with microsensors. Pharmacol. Biochem. Behav. 90, 135–147 (2008).
Dana, H. et al. Sensitive red protein calcium indicators for imaging neural activity. eLife 5, 413 (2016).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Dana, H. et al. High-performance GFP-based calcium indicators for imaging activity in neuronal populations and microcompartments. Nat. Methods https://doi.org/10.1038/s41592-019-0435-6 (2019).
Marvin, J. S. et al. Stability, affinity, and chromatic variants of the glutamate sensor iGluSnFR. Nat. Methods 15, 936–939 (2018).
Masharina, A., Reymond, L., Maurel, D., Umezawa, K. & Johnsson, K. A fluorescent sensor for GABA and synthetic GABAB receptor ligands. J. Am. Chem. Soc. 134, 19026–19034 (2012).
Grimley, J. S. et al. Visualization of synaptic inhibition with an optogenetic sensor developed by cell-free protein engineering automation. J. Neurosci. 33, 16297–16309 (2013).
Jensen, T. P., Zheng, K., Tyurikova, O., Reynolds, J. P. & Rusakov, D. A. Monitoring single-synapse glutamate release and presynaptic calcium concentration in organised brain tissue. Cell Calcium 64, 102–108 (2017).
Jensen, T. P. et al. Multiplex imaging relates quantal glutamate release to presynaptic Ca2+ homeostasis at multiple synapses in situ. Nat. Commun. 10, 1414–1414 (2019).
Marvin, J. S., Schreiter, E. R., Echevarría, I. M. & Looger, L. L. A genetically encoded, high-signal-to-noise maltose sensor. Proteins 79, 3025–3036 (2011).
Geng, Y. et al. Structure and functional interaction of the extracellular domain of human GABA(B) receptor GBR2. Nat. Neurosci. 15, 970–978 (2012).
Planamente, S. et al. A conserved mechanism of GABA binding and antagonism is revealed by structure-function analysis of the periplasmic binding protein Atu2422 in Agrobacterium tumefaciens. J. Biol. Chem. 285, 30294–30303 (2010).
Planamente, S. et al. Structural basis for selective GABA binding in bacterial pathogens. Mol. Microbiol. 86, 1085–1099 (2012).
Guthrie, G. D. & Nicholson-Guthrie, C. S. Gamma-Aminobutyric acid uptake by a bacterial system with neurotransmitter binding characteristics. Proc. Natl Acad. Sci. USA 86, 7378–7381 (1989).
Marvin, J. S. et al. The rational design of allosteric interactions in a monomeric protein and its applications to the construction of biosensors. Proc. Natl Acad. Sci. USA 94, 4366–4371 (1997).
Grimm, J. B. et al. A general method to fine-tune fluorophores for live-cell and in vivo imaging. Nat. Methods 14, 987–994 (2017).
Marvin, J. S. & Hellinga, H. W. Manipulation of ligand binding affinity by exploitation of conformational coupling. Nat. Struct. Biol. 8, 795–798 (2001).
Engel, D. et al. Plasticity of rat central inhibitory synapses through GABA metabolism. J. Physiol. 535, 473–482 (2001).
Brunton, L., Chabner, B. & Knollman, B. Goodman and Gilman’s The Pharmacological Basis of Therapeutics 12th edn (McGraw Hill Professional, 2011).
Putluri, N. et al. Metabolomic profiling reveals a role for androgen in activating amino acid metabolism and methylation in prostate cancer cells. PLoS ONE 6, e21417 (2011).
Benson, D. L., Watkins, F. H., Steward, O. & Banker, G. Characterization of GABAergic neurons in hippocampal cell cultures. J. Neurocytol. 23, 279–295 (1994).
Dodge, F. A. & Rahamimoff, R. Co-operative action of calcium ions in transmitter release at the neuromuscular junction. J. Physiol. 193, 419–432 (1967).
Adesnik, H., Bruns, W., Taniguchi, H., Huang, Z. J. & Scanziani, M. A neural circuit for spatial summation in visual cortex. Nature 490, 226–231 (2012).
Zhang, S. et al. Selective attention. Long-range and local circuits for top-down modulation of visual cortex processing. Science 345, 660–665 (2014).
Nelson, L. E. et al. The sedative component of anesthesia is mediated by GABA(A) receptors in an endogenous sleep pathway. Nat. Neurosci. 5, 979–984 (2002).
Westphalen, R. I. & Hemmings, H. C. Selective depression by general anesthetics of glutamate versus GABA release from isolated cortical nerve terminals. J. Pharmacol. Exp. Ther. 304, 1188–1196 (2003).
Farrant, M. & Nusser, Z. Variations on an inhibitory theme: phasic and tonic activation of GABA(A) receptors. Nat. Rev. Neurosci. 6, 215–229 (2005).
Cohen, I., Navarro, V., Clemenceau, S., Baulac, M. & Miles, R. On the origin of interictal activity in human temporal lobe epilepsy in vitro. Science 298, 1418–1421 (2002).
Trevelyan, A. J., Sussillo, D. & Yuste, R. Feedforward inhibition contributes to the control of epileptiform propagation speed. J. Neurosci. 27, 3383–3387 (2007).
Schevon, C. A. et al. Evidence of an inhibitory restraint of seizure activity in humans. Nat. Commun. 3, 1060 (2012).
Magloire, V., Mercier, M. S., Kullmann, D. M. & Pavlov, I. GABAergic interneurons in seizures: investigating causality with optogenetics. Neuroscientist https://doi.org/10.1177/1073858418805002 (2018).
Kätzel, D., Nicholson, E., Schorge, S., Walker, M. C. & Kullmann, D. M. Chemical-genetic attenuation of focal neocortical seizures. Nat. Commun. 5, 3847 (2014).
Ziemann, A. E. et al. Seizure termination by acidosis depends on ASIC1a. Nat. Neurosci. 11, 816–822 (2008).
Raimondo, J. V. et al. Tight coupling of astrocyte pH dynamics to epileptiform activity revealed by genetically encoded pH sensors. J. Neurosci. 36, 7002–7013 (2016).
Ahrens, M. B. et al. Brain-wide neuronal dynamics during motor adaptation in zebrafish. Nature 485, 471–477 (2012).
Scalise, K., Shimizu, T., Hibi, M. & Sawtell, N. B. Responses of cerebellar Purkinje cells during fictive optomotor behavior in larval zebrafish. J. Neurophysiol. 116, 2067–2080 (2016).
Tanabe, K. et al. Atypical protein kinase C regulates primary dendrite specification of cerebellar Purkinje cells by localizing Golgi apparatus. J. Neurosci. 30, 16983–16992 (2010).
Wu, J. et al. Genetically encoded glutamate indicators with altered color and topology. ACS Chem. Biol. 13, 1832–1837 (2018).
Somogyi, P. & Klausberger, T. Defined types of cortical interneurone structure space and spike timing in the hippocampus. J. Physiol. 562, 9–26 (2005).
Ohno-Shosaku, T. et al. Presynaptic cannabinoid sensitivity is a major determinant of depolarization-induced retrograde suppression at hippocampal synapses. J. Neurosci. 22, 3864–3872 (2002).
Jo, S. et al. GABA from reactive astrocytes impairs memory in mouse models of Alzheimer’s disease. Nat. Med 20, 886–896 (2014).
Emir, U. E., Tuite, P. J. & Öz, G. Elevated pontine and putamenal GABA levels in mild-moderate Parkinson disease detected by 7 Tesla proton MRS. PLoS ONE 7, e30918 (2012).
Raymond, L. A. et al. Pathophysiology of Huntington’s disease: time-dependent alterations in synaptic and receptor function. Neuroscience 198, 252–273 (2011).
Simpson, M. D., Slater, P., Deakin, J. F., Royston, M. C. & Skan, W. J. Reduced GABA uptake sites in the temporal lobe in schizophrenia. Neurosci. Lett. 107, 211–215 (1989).
Chao, H.-T. et al. Dysfunction in GABA signalling mediates autism-like stereotypies and Rett syndrome phenotypes. Nature 468, 263–269 (2010).
Kunkel, T. A., Bebenek, K. & McClary, J. Efficient site-directed mutagenesis using uracil-containing DNA. Meth. Enzym. 204, 125–139 (1991).
Studier, F. W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Rama, S. Shift and mean algorithm for functional imaging with high spatio-temporal resolution. Front. Cell. Neurosci. 9, 446 (2015).
Vladimirov, N. et al. Light-sheet functional imaging in fictively behaving zebrafish. Nat. Methods 11, 883–884 (2014).
White, R. M. et al. Transparent adult zebrafish as a tool for in vivo transplantation analysis. Cell Stem Cell 2, 183–189 (2008).
Kawakami, K. et al. A transposon-mediated gene trap approach identifies developmentally regulated genes in zebrafish. Dev. Cell 7, 133–144 (2004).
Acknowledgements
We would like to thank Catherine S. Nicholson-Guthrie (Indiana University) for the gift of the Pseudomonas fluorescens strain CNG89; D. Stern, S. Picard, A. Lemire and D. Kao for helping sequence the genome of CNG89; D.Walpita for rat neuronal culture; J. Macklin and R. Patel for collecting two-photon spectra; and A. Abdelfattah and C.-L. Hsu for assistance with brain slice recordings. Y.S., M.L. and D.M.K. are supported by the Medical Research Council (grant no. MR/L01095X/1) and Wellcome Trust (grant nos. 095580/Z/11/Z and 212285/Z/18/Z). V.M. is supported by Epilepsy Research UK (grant no. P1702). T.P.J. and D.A.R. are supported by the Wellcome Trust Principal Fellowship (no. 212251/Z/18/Z) and European Research Council Advanced Grant (no. 323113). M.A. is supported by Simons Collaboration on the Global Brain Research Awards 325171 and 542943SPI. This work was supported by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
J.S.M. and L.L.L. were responsible for protein engineering and experimental design and led the project. T.P.J. and D.A.R. performed hippocampal slice imaging. I.K. was responsible for somatosensory slice electrophysiology. K.P. and O.N. performed visual cortex volume imaging. Y.S., V.M., M.L. and D.M.K. produced the mouse epilepsy model. E.L.K. and N.J.L performed mitochondrial experiments. T.K. and M.B.A. designed and performed the zebrafish experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nina Vogt was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–22
Supplementary Video 1
Light-sheet imaging of zebrafish cerebellum during fictive swimming. Companion video to Fig. 5. Fluorescence in the region indicated by the arrow changes during bouts of swimming.
Rights and permissions
About this article
Cite this article
Marvin, J.S., Shimoda, Y., Magloire, V. et al. A genetically encoded fluorescent sensor for in vivo imaging of GABA. Nat Methods 16, 763–770 (2019). https://doi.org/10.1038/s41592-019-0471-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0471-2
This article is cited by
-
Engineered serum markers for non-invasive monitoring of gene expression in the brain
Nature Biotechnology (2024)
-
High-resolution tracking of unconfined zebrafish behavior reveals stimulatory and anxiolytic effects of psilocybin
Molecular Psychiatry (2024)
-
Targeting vulnerable microcircuits in the ventral hippocampus of male transgenic mice to rescue Alzheimer-like social memory loss
Military Medical Research (2024)
-
Plasticity of neuronal dynamics in the lateral habenula for cue-punishment associative learning
Molecular Psychiatry (2023)
-
Cortico-cortical feedback engages active dendrites in visual cortex
Nature (2023)