Abstract
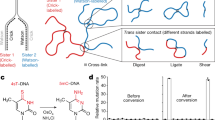
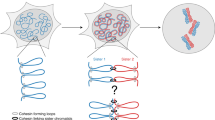
Chromosome segregation requires both compaction and disentanglement of sister chromatids. We describe SisterC, a chromosome conformation capture assay that distinguishes interactions between and along identical sister chromatids. SisterC employs 5-bromo-2′-deoxyuridine (BrdU) incorporation during S-phase to label newly replicated strands, followed by Hi-C and then the destruction of 5-bromodeoxyuridine-containing strands via Hoechst/ultraviolet treatment. After sequencing of the remaining intact strands, this allows assignment of Hi-C products as inter- and intra-sister interactions based on the strands that reads are mapped to. We performed SisterC on mitotic Saccharomyces cerevisiae cells. We find precise alignment of sister chromatids at centromeres. Along arms, sister chromatids are less precisely aligned, with inter-sister connections every ~35 kilobase (kb). Inter-sister interactions occur between cohesin binding sites that are often offset by 5 to 25 kb. Along sister chromatids, cohesin results in the formation of loops of up to 50 kb. SisterC allows study of the complex interplay between sister chromatid compaction and their segregation during mitosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All genomic data generated for this study are publicly available on the NCBI Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) under accession number GSE145695.
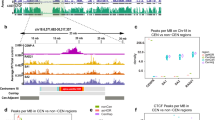
In Fig. 1, Scc1-calibrated ChIP–seq tracks from Hu et al.22 were used for the cohesin pile-up SisterC heatmaps and ChIP–seq tracks. This dataset is available on GEO under accession number GSM1712309. The peaks were called on this dataset using MACS2. The pairwise cohesin interactions were compiled by listing all possible pairwise combinations of cohesin peak sites in the same chromosome, followed by separation based on distance between cohesin pairs (smaller than 10 kb, 10 to 20 kb, 20 to 35 kb and 35 to 50 kb). The cohesin sites in a 50-kb window around the centromeres and on all of chrXII and chrIV were removed from the dataset. Additionally, to investigate distance decay, Hi-C samples from cdc45 mutant cells were used from Schalbetter et al.7. This dataset is available on GEO under accession number GSM2327664. This data was processed identically to the Hi-C libraries produced for this study. The sites of origin of replication were downloaded from OriDB (http://www.oridb.org/)44. Source data are provided with this paper.
Code availability
The Hi-C mapping pipeline distiller-nf is available at https://github.com/mirnylab/distiller-nf. The downstream analysis tools, pairtools and cooltools are available at https://github.com/mirnylab/pairtools and https://github.com/mirnylab/cooltools.
References
Yatskevich, S., Rhodes, J. & Nasmyth, K. Organization of chromosomal DNA by SMC complexes. Annu Rev. Genet. 53, 445–482 (2019).
Nagasaka, K., Hossain, M. J., Roberti, M. J., Ellenberg, J. & Hirota, T. Sister chromatid resolution is an intrinsic part of chromosome organization in prophase. Nat. Cell Biol. https://doi.org/10.1038/ncb3353 (2016).
Falconer, E. et al. Identification of sister chromatids by DNA template strand sequences. Nature 463, 93–97 (2010).
Espinosa, E., Paly, E. & Barre, F.-X. High-resolution whole-genome analysis of sister-chromatid contacts. Mol. Cell https://doi.org/10.1016/j.molcel.2020.06.033 (2020).
Haering, C. H., Farcas, A. M., Arumugam, P., Metson, J. & Nasmyth, K. The cohesin ring concatenates sister DNA molecules. Nature 454, 297–301 (2008).
Tanaka, T., Cosma, M. P., Wirth, K. & Nasmyth, K. Identification of cohesin association sites at centromeres and along chromosome arms. Cell 98, 847–858 (1999).
Schalbetter, S. A. et al. SMC complexes differentially compact mitotic chromosomes according to genomic context. Nat. Cell Biol. 19, 1071–1080 (2017).
Lazar‐Stefanita, L. et al. Cohesins and condensins orchestrate the 4D dynamics of yeast chromosomes during the cell cycle. EMBO J. 36, 2684–2697 (2017).
Kim, Y., Shi, Z., Zhang, H., Finkelstein, I. J. & Yu, H. Human cohesin compacts DNA by loop extrusion. Science 366, 1345–1349 (2019).
Ganji, A. M. et al. Real-time imaging DNA loop extrusion condensin.Science 360, 102–105 (2018).
Golfier, S., Quail, T., Kimura, H. & Brugues, J. Cohesin and condensin extrude DNA loops in a cell cycle-dependent manner. eLife 9, e53885 (2020).
Alves, P. & Jonasson, J. New staining method for the detection of sister-chromatid exchanges in BrdU-labelled chromosomes. J. Cell Sci. 32, 185–195 (1978).
Falconer, E. et al. DNA template strand sequencing of single-cells maps genomic rearrangements at high resolution. Nat. Methods 9, 1107–1112 (2012).
Claussin, C. et al. Genome-wide mapping of sister chromatid exchange events in single yeast cells using strand-seq. eLife 6, e30560 (2017).
Van Wietmarschen, N. & Lansdorp, P. M. Bromodeoxyuridine does not contribute to sister chromatid exchange events in normal or Bloom syndrome cells. Nucleic Acids Res. 44, 6787–6793 (2016).
Chaisson, M. J. P. et al. Multi-platform discovery of haplotype-resolved structural variation in human genomes. Nat. Commun. 10, 1784 (2019).
Porubsky, D. et al. Dense and accurate whole-chromosome haplotyping of individual genomes. Nat. Commun. 8, 1293 (2017).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Belaghzal, H., Dekker, J. & Gibcus, J. H. Hi-C 2.0: an optimized Hi-C procedure for high-resolution genome-wide mapping of chromosome conformation. Methods 123, 56–65 (2017).
Vernis, L., Piskur, J. & Diffley, J. F. X. Reconstitution of an efficient thymidine salvage pathway in Saccharomyces cerevisiae. Nucleic Acids Res. 31, 120e–120e (2003).
Lajoie, B. R., Dekker, J. & Kaplan, N. The Hitchhiker’s guide to Hi-C analysis: practical guidelines. Methods 72, 65–75 (2015).
Hu, B. et al. Biological chromodynamics: a general method for measuring protein occupancy across the genome by calibrating ChIP-seq. Nucleic Acids Res. 43, e132 (2015).
Fudenberg, G., Abdennur, N., Imakaev, M., Goloborodko, A. & Mirny, L. A. Emerging evidence of chromosome folding by loop extrusion. Cold Spring Harb. Symp. Quant. Biol. LXXXII, 034710 (2018).
Lengronne, A. et al. Cohesin relocation from sites of chromosomal loading to places of convergent transcription. Nature 430, 573–578 (2004).
Schalbetter, S. A., Neale, M. J., Fudenberg, G., Baxter, J. & Pollard, K. S. Principles of meiotic chromosome assembly revealed in S. cerevisiae. Nat. Commun. https://doi.org/10.1038/s41467-019-12629-0 (2019).
Muller, H. et al. Characterizing meiotic chromosomes’ structure and pairing using a designer sequence optimized for Hi‐C. Mol. Syst. Biol. 14, e8293 (2018).
AlHaj Abed, J. et al. Highly structured homolog pairing reflects functional organization of the Drosophila genome. Nat. Commun. 10, 4485 (2019).
Zheng, G., Kanchwala, M., Xing, C. & Yu, H. MCM2-7-dependent cohesin loading during S phase promotes sister-chromatid cohesion. eLife 7, e33920 (2018).
Lengronne, A. et al. Establishment of sister chromatid cohesion at the S. cerevisiae replication fork. Mol. Cell https://doi.org/10.1016/j.molcel.2006.08.018 (2006).
Moldovan, G. L., Pfander, B. & Jentsch, S. PCNA controls establishment of sister chromatid cohesion during S phase. Mol. Cell https://doi.org/10.1016/j.molcel.2006.07.007 (2006).
Borrie, M. S., Campor, J. S., Joshi, H. & Gartenberg, M. R. Binding, sliding, and function of cohesin during transcriptional activation. Proc. Natl Acad. Sci. USA 114, E1062–E1071 (2017).
Bausch, C. et al. Transcription alters chromosomal locations of cohesin in Saccharomyces cerevisiae. Mol. Cell. Biol. 27, 8522–8532 (2007).
Makrantoni, V. & Marston, A. L. Cohesin and chromosome segregation. Curr. Biol. 28, R688–R693 (2018).
Kikuchi, S., Borek, D. M., Otwinowski, Z., Tomchick, D. R. & Yu, H. Crystal structure of the cohesin loader Scc2 and insight into cohesinopathy. Proc. Natl Acad. Sci. USA 113, 12444–12449 (2016).
Lee, B. G. et al. Crystal structure of the cohesin gatekeeper Pds5 and in complex with Kleisin Scc1. Cell Rep. 14, 2108–2115 (2016).
Tóth, A. et al. Yeast cohesin complex requires a conserved protein, Eco1p(Ctf7), to establish cohesion between sister chromatids during DNA replication. Genes Dev. 13, 320–333 (1999).
Skibbens, R. V., Corson, L. B., Koshland, D. & Hieter, P. Ctf7p is essential for sister chromatid cohesion and links mitotic chromosome structure to the DNA replication machinery. Genes Dev. 13, 307–319 (1999).
Mitter, M. et al. Sister-chromatid-sensitive Hi-C reveals the conformation of replicated human chromosomes. Nature https://doi.org/10.1038/s41586-020-2744-4 (2020).
Gibcus, J. H. et al. A pathway for mitotic chromosome formation. Science 359, eaao6135 (2018).
Falconer, E. & Lansdorp, P. M. Strand-seq: a unifying tool for studies of chromosome segregation. Semin. Cell Dev. Biol. 24, 643–652 (2013).
Andrus, A. & Kuimelis, R. G. Base composition analysis of nucleosides using HPLC. Curr. Protoc. Nucleic Acid Chem. 1, 10.6.1–10.6.6 (2000).
Abdennur, N. & Mirny, L. A. Cooler: scalable storage for Hi-C data and other genomically labeled arrays. Bioinformatics 36, 311–316 (2019).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Siow, C. C., Nieduszynska, S. R., Müller, C. A. & Nieduszynski, C. A. OriDB, the DNA replication origin database updated and extended. Nucleic Acids Res. 40, 682–686 (2012).
Acknowledgments
We thank all current and former Dekker lab members for helpful discussions, in particular S. V. Venev for advice on data analysis. We thank J. Benanti, H. Arsenault and N. Rhind for advice on culturing and synchronizing yeast and providing the yeast strain used in this study. We thank S. Schalbetter, N. Minchell and M. Naele for providing their yeast Hi-C protocol and suggestions. We also thank B. van Steensel for advice and suggestions. This work was supported by grants from the National Institutes of Health (no. HG003143 to J.D. and no. NS111990 to J.K.W.). J.D. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
M.E.O. and J.D. conceived and designed the project. M.E.O. cultured yeast, generated and analyzed the SisterC and Hi-C datasets and performed the flow cytometry experiments. A.K.H. and J.K.W. designed the HPLC and mass spectrometry experiments, A.K.H. performed and analyzed the HPLC and mass spectrometry experiments. M.E.O. and J.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Lei Tang was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Outline of SisterC yeast culture conditions and flow cytometry analysis.
a, Asynchronous yeast cultures are cultured and synchronized in late G1 using alpha factor. Cells are released in media containing BrdU or Thymidine, followed by an arrest in mitosis using nocodazole or G1 using a second alpha factor arrest. Cells are harvested and prepared for Hi-C or SisterC library production or processed for BrdU detection using HPLC. b, Flow cytometry analysis of cell cycle profile and BrdU incorporation of harvested yeast cultures for preparation of SisterC libraries.
Extended Data Fig. 2 Depletion of BrdU containing DNA molecules by PCR.
DNA fragments were amplified in presence of 0%, 10%, 50%, 90% and 100% BrdU to allow for incorporation in both strands (first 5 lanes). This was followed by treatment of UV only (second 5 lanes), Hoechst only (third 5 lanes) or treatment with both UV and Hoechst (last 5 lanes). Fragments containing more than 10% BrdU did not get amplified after UV/Hoechst treatment. This experiment was repeated twice with highly similar results.
Extended Data Fig. 3 Distance decay plots of all SisterC replicates.
Distance decay plots of all SisterC mitotic libraries: DpnII replicate 1 a, DpnII replicate 2 b, DpnII replicate 3 c, and HindIII replicate 1 d.
Extended Data Fig. 4 Distance decay plots of all SisterC control experiments.
Distance decay plots of all SisterC control libraries: a-b, G1 (a) and mitotic (b) arrested cells grown in BrdU and treated with UV/Hoechst. c-d, G1 (c) and mitotic (d) arrested cells grown in BrdU, not treated with UV/Hoechst. e-f, G1 (e) and mitotic (f) arrested cells grown in Thymidine, treated with UV/Hoechst. g-h, G1 (g) and mitotic (h) arrested cells grown in Thymidine, not treated with UV/Hoechst.
Extended Data Fig. 5 Observed/expected interaction heatmap of chrXIII:450,000–550,000.
Log2 observed over expected interaction frequency on region ChrXIII:450,000–550,000 (as shown in main Fig. 2) for all interactions a, inter-sister interactions b, and intra-sister interactions c.
Extended Data Fig. 6 Centromeric region of chrXII.
a-c, Interaction heatmaps of 50 kb window around the centromeric region of chrXII, for all interactions (a), inter-sister interactions (b) and intra-sister interactions (c). d-f, Log2 observed over expected interaction heatmaps of all interactions (d), inter-sister interactions (e) and intra-sister interactions (f) on the centromeric region on chrXII.
Extended Data Fig. 7 Anchor plot on ChrXIII:469000.
Interaction frequency of inter-sister and intra-sister interactions anchored on chrXIII:469,000 shows higher frequency of inter-sister interactions at 25 kb distance from the anchor (highlighted in green). Annotated below the plot are the positions of Scc1 peaks.
Extended Data Fig. 8 Ranking of SisterC signal on pairs of cohesin binding sites at different genomic distances.
a, b, Inter-sister interaction (a) and intra-sister interaction (b) frequency of pairs of cohesin binding sites separated by 10 to 20 kb were ranked by inter-sister interaction intensity for 10 kb window around the cohesin binding site. c, Interaction pile up plots of inter-sister and intra-sister interactions of the top 10 percent sites that were ranked in (a). d-e, Inter-sister interaction (d) and intra-sister interaction (e) frequency of pairs of cohesin binding sites separated by 35 to 50 kb were ranked by intra-sister interaction intensity for 10 kb window around the cohesin binding site. f, Interaction pile up plots of inter-sister and intra-sister interactions of the top 10 percent sites that were ranked in (e). g, 284 cohesin sites that anchor the top 10% inter-sister interacting cohesin pairs at 10 to 20 kb distance were identified as well as again 284 cohesin sites that anchor the top 10% intra-sister interacting cohesin pairs at 35 to 50 kb distance. 65 cohesin sites are found to mediate both top 10% inter-sister cohesin-cohesin interactions and top 10% intra-sister cohesin-cohesin interactions.
Extended Data Fig. 9 SisterC pile up plots of origins of replication.
a, b, Mitotic SisterC inter-sister interactions (a) and intra-sister interactions (b) plotted on replication origins. c, Pile up plot of Hi-C data of G1 alpha arrested cells on replication origins.
Extended Data Fig. 10 Efficiency of SisterC inter- and intra-sister interaction assignment.
a, b, Normalized frequency SisterC G1 (a) and mitotic (b) inter-sister and intra-sister interactions as a function of AT percentage. c, d, Normalized frequency SisterC G1 (c) and mitotic (d) inter-sister and intra-sister interactions as a function of distance to DpnII digestion site. e, f, Normalized frequency SisterC G1 (e) and mitotic (f) inter-sister and intra-sister interactions as a function of distance to origins of replication.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4.
Supplementary Table
Supplementary Table 1. SisterC and Hi-C mapping statistics across all DpnII and HindIII experiments
Source data
Source Data Fig. 1
HPLC datapoints.
Source Data Fig. 2
Datapoints of scaling plots and heatmaps
Source Data Fig. 3
Datapoints of scaling plots and heatmaps.
Source Data Fig. 4
Datapoints of heatmaps and anchorplot.
Source Data Extended Data Fig. 3
Datapoints of scaling plots.
Source Data Extended Data Fig. 4
Datapoints of scaling plots.
Source Data Extended Data Fig. 5
Datapoints of heatmaps.
Source Data Extended Data Fig. 6
Datapoints of heatmaps.
Source Data Extended Data Fig. 7
Datapoints of anchor plot.
Source Data Extended Data Fig. 8
Datapoints of heatmaps.
Source Data Extended Data Fig. 9
Datapoints of heatmaps.
Source Data Extended Data Fig. 10
Datapoints of frequency plots.
Rights and permissions
About this article
Cite this article
Oomen, M.E., Hedger, A.K., Watts, J.K. et al. Detecting chromatin interactions between and along sister chromatids with SisterC. Nat Methods 17, 1002–1009 (2020). https://doi.org/10.1038/s41592-020-0930-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0930-9
This article is cited by
-
A spatial genome aligner for resolving chromatin architectures from multiplexed DNA FISH
Nature Biotechnology (2023)
-
Sister chromatid–sensitive Hi-C to map the conformation of replicated genomes
Nature Protocols (2022)
-
Understanding 3D genome organization by multidisciplinary methods
Nature Reviews Molecular Cell Biology (2021)