Abstract
We develop an automatic method for synaptic partner identification in insect brains and use it to predict synaptic partners in a whole-brain electron microscopy dataset of the fruit fly. The predictions can be used to infer a connectivity graph with high accuracy, thus allowing fast identification of neural pathways. To facilitate circuit reconstruction using our results, we develop CIRCUITMAP, a user interface add-on for the circuit annotation tool CATMAID.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Code availability
The code used to train networks and predict synaptic partners is available in the synful repository (https://github.com/funkelab/synful)22. Code used to evaluate the results is available in the synful_fafb repository (https://github.com/funkelab/synful_fafb)20. A python library to query circuits in the FAFB volume using the predicted synaptic partners is available in the SynfulCircuit repository (https://github.com/funkelab/synfulcircuit)14. The CATMAID widget used to interactively query circuits is available in the CircuitMap repository (https://github.com/catmaid/circuitmap)15.
References
Denk, W., Briggman, K. L. & Helmstaedter, M. Structural neurobiology: missing link to a mechanistic understanding of neural computation. Nat. Rev. Neurosci. 13, 351–358 (2012).
Zheng, Z. et al. A complete electron microscopy volume of the brain of adult Drosophila melanogaster. Cell 174, 730–743 (2018).
Li, P. H. et al. Automated reconstruction of a serial-section EM Drosophila brain with flood-filling networks and local realignment. Microsc. Microanal. 25 (Suppl. 2), 1364–1365 (2019).
Dorkenwald, S. et al. Automated synaptic connectivity inference for volume electron microscopy. Nat. Methods 14, 435–442 (2017).
Motta, A. et al. Dense connectomic reconstruction in layer 4 of the somatosensory cortex. Science 366, eaay3134 (2019).
Heinrich, L., Funke, J., Pape, C., Nunez-Iglesias, J. & Saalfeld, S. Synaptic cleft segmentation in non-isotropic volume electron microscopy of the complete drosophila brain. In Medical Image Computing and Computer Assisted Intervention – MICCAI 2018 (Springer, 2018).
Huang, G. B., Scheffer, L. K. & Plaza, S. M. Fully-automatic synapse prediction and validation on a large data set. Front. Neural Circuits 12, 87 (2018).
Kreshuk, A., Funke, J., Cardona, A. & Hamprecht, F. A. Who is talking to whom: synaptic partner detection in anisotropic volumes of insect brain. In Int. Conf. Medical Image Computing and Computer-Assisted Intervention, 661–668 (Springer, 2015).
Turner, N. L. et al. Synaptic partner assignment using attentional voxel association networks. In 2020 IEEE International Symposium on Biomedical Imaging, 1209–1213 (IEEE Computer Society, 2020).
Parag, T. et al. Detecting synapse location and connectivity by signed proximity estimation and pruning with deep nets. In Proc. European Conference on Computer Vision (ECCV) Workshops (Springer, 2018).
Buhmann, J. et al. Synaptic partner prediction from point annotations in insect brains. In International Conference on Medical Image Computing and Computer-Assisted Intervention, 309–316 (Springer, 2018).
Xu, C. S. et al. A connectome of the adult Drosophila central brain. eLife 9, e57443 (2020).
Ronneberger, O., Fischer, P. & Brox, T. U-net: convolutional networks for biomedical image segmentation. In Int. Conf. Medical Image Computing and Computer-Assisted Intervention, 234–241 (Springer, 2015).
Buhmann, J. & Funke, J. funkelab/synfulcircuit v1.0 (2021); https://doi.org/10.5281/zenodo.4633048
Kazimiers, T., Gerhard, S. & Funke, J. catmaid/circuitmap (2021); https://doi.org/10.5281/zenodo.4633884
Schneider-Mizell, C. M. et al. Quantitative neuroanatomy for connectomics in drosophila. eLife 5, e12059 (2016).
Falk, T. et al. U-net: deep learning for cell counting, detection, and morphometry. Nat. Methods 16, 67 (2019).
Funke, J. et al. Large scale image segmentation with structured loss based deep learning for connectome reconstruction. IEEE Trans. Pattern Anal. Mach. Intell. 41, 1669–1680 (2018).
Wang, K., Dou, J., Kemao, Q., Di, J. & Zhao, J. Y-net: a one-to-two deep learning framework for digital holographic reconstruction. Opt. Lett. 44, 4765–4768 (2019).
Buhmann, J. & Funke, J. funkelab/synful_fafb v1.0 (2021); https://doi.org/10.5281/zenodo.4633135
Funke, J., Buhmann, J., Sheridan, A., Nguyen, T. M. & Malin-Mayor, C. synful_experiments (2021); https://doi.org/10.5281/zenodo.4635362
Buhmann, J., Funke, J. & Nguyen, T. M. funkelab/synful v1.0 (2021); https://doi.org/10.5281/zenodo.4633044
Acknowledgements
We thank S. Lauritzen (HHMI, Janelia Research Campus) for helping with data acquisition; M. Nichols (HHMI, Janelia Research Campus) for importing CATMAID annotations; W. Patton (HHMI, Janelia Research Campus) for code contributions; N. Eckstein (HHMI, Janelia Research Campus) for helpful discussions; J. Maitin-Shepard (Google) for adding synapse visualization features to neuroglancer; V. Jayaraman (HHMI, Janelia Research Campus) for providing evaluation data; and the HHMI Janelia Connectome Annotation Team (R. Parekh, A. Suleiman, T. Paterson) for evaluation data. We also thank Z. Zheng, F. Li, C. Fisher, N. Sharifi and S. Calle-Schuler (HHMI, Janelia) for access to prepublication data used for evaluation. This work was supported by the HHMI and the Swiss National Science Foundation (grant 205321L 160133).
Author information
Authors and Affiliations
Contributions
S.C.T. and J.F. conceived the work. W.-C.A.L., R.W., M.C. and J.F. acquired funding. J.B., A.S., C.M.-M., S.G., R.K., T.M.N., L.H., P.S., S.S. and J.F. developed software. J.B., C.M.-M. and J.F. carried out validation and evaluation. P.S., G.S.X.E.J. and D.D.B. generated evaluation data. J.B., T.K., S.G. and J.F. disseminated data. M.C. and J.K. supervised the work. J.B., A.S., C.M.-M., P.S. and J.F. produced visualizations. J.B., P.S. and J.F. drafted the manuscript. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.G. is the founder and CEO of UniDesign Solutions GmbH, which provides IT services.
Additional information
Peer review information Nature Methods thanks Chung-Chuan Lo, Stephan Sigrist and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Brain-wide prediction accuracy.
Precision and recall over 10 densely annotated cubes in different brain regions (dataset DenseCubes). Highlights show best f-score over the detection threshold. Top row: Black triangle marks show the inter-human variance between two annotations, with one annotation treated as ground-truth evaluated against the other one. Bottom right: Visualization of the cube locations within the FAFB dataset.
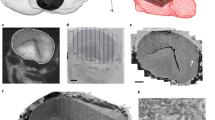
Extended Data Fig. 2 Polarity of olfactory projection neurons (PN).
a, Outline of axon-dendrite splitting procedure. b, Exemplary well segregated (left) and unsegregated (right) PN. Boxplot shows segregation index (SI) for all PNs separated into uni- and multiglomerular PNs. c, Impact of completeness of neuronal reconstruction on recovery of synapses.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3, Notes 1–3 and Discussion
Supplementary Data 1
Per-neuron prediction accuracy
Supplementary Video 1
CircuitMap instruction video
Supplementary Data 2
Source data for Supplementary Fig. 2.
Source data
Source Data Fig. 2
Source data for Fig. 2
Source Data Extended Data Fig. 1
Source data for Extended Data Fig. 1
Source Data Extended Data Fig. 2
Source data for Extended Data Fig. 2
Rights and permissions
About this article
Cite this article
Buhmann, J., Sheridan, A., Malin-Mayor, C. et al. Automatic detection of synaptic partners in a whole-brain Drosophila electron microscopy data set. Nat Methods 18, 771–774 (2021). https://doi.org/10.1038/s41592-021-01183-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-021-01183-7
This article is cited by
-
Modular segmentation, spatial analysis and visualization of volume electron microscopy datasets
Nature Protocols (2024)
-
Heterogeneity of synaptic connectivity in the fly visual system
Nature Communications (2024)
-
Multi-layered maps of neuropil with segmentation-guided contrastive learning
Nature Methods (2023)
-
Structured cerebellar connectivity supports resilient pattern separation
Nature (2023)
-
Local shape descriptors for neuron segmentation
Nature Methods (2023)