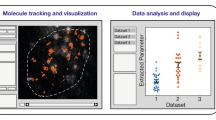
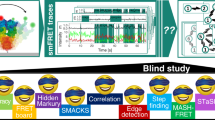
Abstract
In single-particle tracking, individual particles are localized and tracked over time to probe their diffusion and molecular interactions. Temporal crossing of trajectories, blinking particles, and false-positive localizations present computational challenges that have remained difficult to overcome. Here we introduce a robust, parameter-free alternative to single-particle tracking: temporal analysis of relative distances (TARDIS). In TARDIS, an all-to-all distance analysis between localizations is performed with increasing temporal shifts. These pairwise distances represent either intraparticle distances originating from the same particle, or interparticle distances originating from unrelated particles, and are fitted analytically to obtain quantitative measures on particle dynamics. We showcase that TARDIS outperforms tracking algorithms, benchmarked on simulated and experimental data of varying complexity. We further show that TARDIS performs accurately in complex conditions characterized by high particle density, strong emitter blinking or false-positive localizations, and is in fact limited by the capabilities of localization algorithms. TARDIS’ robustness enables fivefold shorter measurements without loss of information.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All data underlying this study is available at ref. 60. Source data are provided with this paper.
Code availability
The custom TARDIS software used in this manuscript is provided as supplementary data and can be accessed at ref. 39.
References
Elf, J. & Barkefors, I. Single-molecule kinetics in living cells. Annu. Rev. Biochem. 88, 635–659 (2019).
Shen, H. et al. Single particle tracking: from theory to biophysical applications. Chem. Rev. 117, 7331–7376 (2017).
Martens, K., van Duynhoven, J. & Hohlbein, J. Spatiotemporal heterogeneity of κ-carrageenan gels investigated via single-particle-tracking fluorescence microscopy. Langmuir 20, 5502–5509 (2020).
Prigent, S., Valades-Cruz, C. A., Leconte, L., Salamero, J. & Kervrann, C. STracking: a free and open-source Python library for particle tracking and analysis. Bioinformatics 38, 3671–3673 (2022).
Sbalzarini, I. F. & Koumoutsakos, P. Feature point tracking and trajectory analysis for video imaging in cell biology. J. Struct. Biol. 151, 182–195 (2005).
Manley, S. et al. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat. Methods 5, 155–157 (2008).
Ortega-Arroyo, J. & Kukura, P. Interferometric scattering microscopy (iSCAT): new frontiers in ultrafast and ultrasensitive optical microscopy. Phys. Chem. Chem. Phys. 14, 15625–15636 (2012).
Kukura, P. et al. High-speed nanoscopic tracking of the position and orientation of a single virus. Nat. Methods 6, 923–927 (2009).
Martens, K. J. A. et al. Visualisation of dCas9 target search in vivo using an open-microscopy framework. Nat. Commun. 10, 3552 (2019).
Ohana, R. F. et al. HaloTag7: a genetically engineered tag that enhances bacterial expression of soluble proteins and improves protein purification. Protein Expr. Purif. 68, 110–120 (2009).
Klein, T. et al. Live-cell dSTORM with SNAP-tag fusion proteins. Nat. Methods 8, 7–9 (2011).
Stallinga, S. & Rieger, B. Accuracy of the Gaussian point spread function model in 2D localization microscopy. Opt. Express 18, 24461–24476 (2010).
Li, Y. et al. Real-time 3D single-molecule localization using experimental point spread functions. Nat. Methods 15, 367–369 (2018).
Xu, F. et al. Three-dimensional nanoscopy of whole cells and tissues with in situ point spread function retrieval. Nat. Methods 17, 531–540 (2020).
Aristov, A., Lelandais, B., Rensen, E. & Zimmer, C. ZOLA-3D allows flexible 3D localization microscopy over an adjustable axial range. Nat. Commun. 9, 2409 (2018).
Nehme, E., Weiss, L. E., Michaeli, T. & Shechtman, Y. Deep-STORM: super-resolution single-molecule microscopy by deep learning. Optica 5, 458–464 (2018).
Speiser, A. et al. Deep learning enables fast and dense single-molecule localization with high accuracy. Nat. Methods 18, 1082–1090 (2021).
Nehme, E. et al. DeepSTORM3D: dense 3D localization microscopy and PSF design by deep learning. Nat. Methods 17.7, 734–740 (2020).
Li, N. et al. Photonic resonator interferometric scattering microscopy. Nat. Commun. 12, 1744 (2021).
Cnossen, J. et al. Localization microscopy at doubled precision with patterned illumination. Nat. Methods 17, 59–63 (2020).
Jouchet, P. et al. Nanometric axial localization of single fluorescent molecules with modulated excitation. Nat. Photonics https://doi.org/10.1038/s41566-020-00749-9 (2021).
Gu, L. et al. Molecular resolution imaging by repetitive optical selective exposure. Nat. Methods 16, 1114–1118 (2019).
Chenouard, N. et al. Objective comparison of particle tracking methods. Nat. Methods 11, 281–289 (2014).
Simon, F., Tinevez, J.-Y. & Teeffelen, S. van. ExTrack characterizes transition kinetics and diffusion in noisy single-particle tracks. J. Cell Biol. https://doi.org/10.1083/jcb.202208059; https://doi.org/10.1101/2022.07.13.499913 (2022).
Chenouard, N., Bloch, I. & Olivo-Marin, J.-C. Multiple hypothesis tracking for cluttered biological image sequences. IEEE Trans. Pattern Anal. Mach. Intell. 35, 2736–3750 (2013).
Floc’h, K. et al. Bacterial cell wall nanoimaging by autoblinking microscopy. Sci. Rep. 8, 14038 (2018).
Vink, J. N. A., Brouns, S. J. J. & Hohlbein, J. Extracting transition rates in particle tracking using analytical diffusion distribution analysis. Biophys. J. 119, 1970–1983 (2020).
Vink, J. N. A. et al. Direct visualization of native CRISPR target search in live bacteria reveals cascade DNA surveillance mechanism. Mol. Cell 77, 39–50.e10 (2020).
Karslake, J. D. et al. SMAUG: analyzing single-molecule tracks with nonparametric Bayesian statistics. Methods 193, 16–26 (2021).
Hansen, A. S. et al. Robust model-based analysis of single-particle tracking experiments with Spot-On. eLife 7, e33125 (2018).
Persson, F., Lindén, M., Unoson, C. & Elf, J. Extracting intracellular diffusive states and transition rates from single-molecule tracking data. Nat. Methods 10, 265–269 (2013).
Briane, V., Kervrann, C. & Vimond, M. Statistical analysis of particle trajectories in living cells. Phys. Rev. E 97, 062121 (2018).
Momboisse, F. et al. Tracking receptor motions at the plasma membrane reveals distinct effects of ligands on CCR5 dynamics depending on its dimerization status. eLife 11, e76281 (2022).
Granik, N. et al. Single-particle diffusion characterization by deep learning. Biophys. J. 117, 185–192 (2019).
Pécot, T., Zengzhen, L., Boulanger, J., Salamero, J. & Kervrann, C. A quantitative approach for analyzing the spatio-temporal distribution of 3D intracellular events in fluorescence microscopy. eLife 7, e32311 (2018).
Curd, A. P. et al. Nanoscale pattern extraction from relative positions of sparse 3D localizations. Nano Lett. 21, 1213–1220 (2021).
Endesfelder, U., Malkusch, S., Fricke, F. & Heilemann, M. A simple method to estimate the average localization precision of a single-molecule localization microscopy experiment. Histochem. Cell Biol. 141, 629–638 (2014).
Bohrer, C. H. et al. A pairwise distance distribution correction (DDC) algorithm to eliminate blinking-caused artifacts in SMLM. Nat. Methods 18, 669–677 (2021).
TARDIS-public. GitHub (accessed 2023); https://github.com/kjamartens/TARDIS-public
Jaqaman, K. et al. Robust single-particle tracking in live-cell time-lapse sequences. Nat. Methods 5, 695–702 (2008).
Tinevez, J.-Y. et al. TrackMate: an open and extensible platform for single-particle tracking. Methods 115, 80–90 (2017).
Roudot, P., Ding, L., Jaqaman, K., Kervrann, C. & Danuser, G. Piecewise-stationary motion modeling and iterative smoothing to track heterogeneous particle motions in dense environments. IEEE Trans. Image Process. 26, 5395–5410 (2017).
de Chaumont, F. et al. Icy: an open bioimage informatics platform for extended reproducible research. Nat. Methods 9, 690–696 (2012).
Wolf, A., Volz-Rakebrand, P., Balke, J. & Alexiev, U. Diffusion analysis of nanoscopic ensembles: a tracking-free diffusivity analysis for nanoscopic ensembles in biological samples and nanotechnology. Small 19, 2206722 (2023).
Bettridge, K., Harris, F. E., Yehya, N. & Xiao, J. RNAP promoter search and transcription kinetics in live E. coli cells. J. Phys. Chem. B 127, 3816–3828 (2023).
Endesfelder, U. et al. Multiscale spatial organization of RNA polymerase in Escherichia coli. Biophys. J. 105, 172–181 (2013).
Virant, D., Turkowyd, B., Balinovic, A. & Endesfelder, U. Combining primed photoconversion and UV-photoactivation for aberration-free, live-cell compliant multi-color single-molecule localization microscopy imaging. Int. J. Mol. Sci. 18, 1524 (2017).
Turkowyd, B. et al. A general mechanism of photoconversion of green‐to‐red fluorescent proteins based on blue and infrared light reduces phototoxicity in live‐cell single‐molecule imaging. Angew. Chem. Int. Ed. 56, 11634–11639 (2017).
Smal, I. et al. Multiple object tracking in molecular bioimaging by Rao-Blackwellized marginal particle filtering. Med. Image Anal. 12, 764–777 (2008).
Ritter, C. et al. Data fusion and smoothing for probabilistic tracking of viral structures in fluorescence microscopy images. Med. Image Anal. 73, 102168 (2021).
Elf, J., Li, G.-W. & Xie, X. S. Probing transcription factor dynamics at the single-molecule level in a living cell. Science 316, 1191–1194 (2007).
Berglund, A. J. Statistics of camera-based single-particle tracking. Phys. Rev. E 82, 011917 (2010).
Philip, J. The Probability Distribution of the Distance between Two Random Points in a Box. Tech. Rep. Number TRITA MAT 07 MA 10 (TRITA MAT, 1991).
Virant, D. et al. A peptide tag-specific nanobody enables high-quality labeling for dSTORM imaging. Nat. Commun. 9, 930 (2018).
Edelstein, A. D. et al. Advanced methods of microscope control using μManager software. J. Biol. Methods 1, e10 (2014).
Pinkard, H. et al. Pycro-Manager: open-source software for customized and reproducible microscope control. Nat. Methods https://doi.org/10.1038/s41592-021-01087-6 (2021).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Ovesny, M., Křížek, P., Borkovec, J., Švindrych, Z. & Hagen, G. M. ThunderSTORM: a comprehensive ImageJ plug-in for PALM and STORM data analysis and super-resolution imaging. Bioinformatics 30, 2389–2390 (2014).
Martens, K. J. A., Bader, A. N., Baas, S., Rieger, B. & Hohlbein, J. Phasor based single-molecule localization microscopy in 3D (pSMLM-3D): an algorithm for MHz localization rates using standard CPUs. J. Chem. Phys. 148, 123311 (2018).
Martens, K. J. A. et al. Data underlying the TARDIS manuscript. Zenodo https://doi.org/10.5281/zenodo.7900405 (2023).
Acknowledgements
This work was financially supported by funding from a VLAG PhD-fellowship (J.H.), start-up funds at Carnegie Mellon University (B.T., K.J.A.M. and U.E.), the NSF AI Institute: Physics of the Future (NSF PHY- 2020295) (U.E.), start-up funds at Bonn University (B.T., K.J.A.M. and U.E.), an Argelander Starter Kit at the University of Bonn (K.J.A.M.) and the Alexander von Humboldt Foundation (K.J.A.M.). We acknowledge the valuable input from group meetings from all members in the U.E. and J.H. laboratories. We thank M. Deserno (Carnegie Mellon University, Pittsburgh, USA) for helpful discussions on TARDIS’ mathematical foundation.
Author information
Authors and Affiliations
Contributions
Conceptualization: K.J.A.M. Data curation: K.J.A.M. Formal analysis: K.J.A.M. Funding acquisition: K.J.A.M., J.H. and U.E. Investigation: K.J.A.M. and B.T. Methodology: K.J.A.M., B.T., J.H. and U.E. Project administration: K.J.A.M., J.H. and U.E. Software: K.J.A.M. Supervision: K.J.A.M., J.H. and U.E. Visualization: K.J.A.M. Writing—original draft: K.J.A.M. Writing—review and editing: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Peer review
Peer review information
Nature Methods thanks J. Christof Gebhardt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Editor: Rita Strack, in collaboration with the Nature Methods team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Performance of TARDIS and spt tracking methods for a single diffusive population and two diffusive populations at increasing complexity, visualised on a linear x-axis.
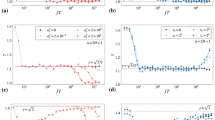
Performance of TARDIS is compared to the blind tracking algorithms uTrack-inspired PMMS (piecewise-stationary motion model and iterative smoothing)40,42, TrackMate40,41 and Nearest neighbour analysis, and to the prior-informed methods swift (Endesfelder et al., manuscript in prep.), and Multiple-Hypothesis Tracking (MHT)25 (MHT at the most complex dataset did not run to completion). This data is also presented in Figs. 1c and 2c. (c) Bhattacharyya distance of the distributions in (a) and (b) compared to the ground truth (GT) jump distance distribution, calculated as the negative natural logarithm of the sum of the square root of the product of the distribution value of a method and that of the jump distance ground truth.
Extended Data Fig. 2 The full dataset as presented Fig. 2a, analysed via TARDIS (blue), TARDIS-JD-extraction (light-blue), TrackMate-LAP40,41 (light green) and nearest-neighbour tracking (dark green).
(a) and (b) represent the same datasets, but visualised on a logarithmic (a) or linear (b) x-axis. Note the changing jump distance x-axis scaling in (b). The TARDIS fit data is also presented in Extended Data Fig. 3a.
Extended Data Fig. 3 Detailed information on diffusivity and bleach time obtained from TARDIS fitting.
(a) Boxplots showing the obtained diffusion coefficients of datasets presented in Fig. 2a, and (b) the obtained bleaching times of datasets presented in Fig. 2a, showing no bias in either over the complexity range. TARDIS is repeated 10 times on 20.000 simulated localizations for each condition. (c) Individual fit information of data presented in Fig. 2b. For every condition, TARDIS is repeated 10 times on 20.000 simulated localizations with random start positions in TARDIS. The obtained diffusion coefficients are visualised (scatter points represent individual measurements). Note the changing y-axis at 90% removed true positives in (c). Abbreviations used: TP: True Positives, FP: False Positives, fr: frame, locs: localizations. All boxplots show the median as the central mark, with the 25th and 75th percentile as lower and upper edges. Whiskers extend to non-outlier extreme points, and outlier points are plotted as plusses.
Extended Data Fig. 4 Detailed information on TARDIS fitting results of bleach time of single diffusive population with added noise and blinking chance.
Fit information of bleach characteristics corresponding to the data presented in Fig. 2a,b (main manuscript). For every condition, 10 repetitions were analysed with random start positions in TARDIS. (a,b) RMSE of the diffusion coefficient (a) and bleach half-time (b) for all conditions presented in Fig. 2a (c) The found diffusion coefficient presented in Fig. 2b is visualised (scatter points represent individual measurements). Note the changing y-axis at 90% removed true positives. TARDIS is repeated 10 times on 20.000 simulated localizations for each condition. All boxplots show the median as the central mark, with the 25th and 75th percentile as lower and upper edges. Whiskers extend to non-outlier extreme points, and outlier points are plotted as plusses. Abbreviations used: TP: True Positives, FP: False Positives, fr: frame, locs: localizations.
Extended Data Fig. 5 Effect of the fraction of inter-particle linkages on diffusion coefficient accuracy.
Analysis of 95% confidence interval (a, b), and fitted diffusion coefficient (c) as a measure of the inter-particle fraction. Dotted lines in (a) are added for clarity. The underlying analysed data is the same as shown in Fig. 2b. Reasons for increased fraction of inter-particle linkages are clarified via marker type (TP removal), marker colour (TP localization density), and marker darkness (FP introduction). Note that a and c have non-linear x-axis (a and b contain the same information, but with different x-axes).
Extended Data Fig. 6 Diffusion analysis of fluorescent beads at varying frame times.
The same information as presented in Fig. 2d, but with additional frame times in between those shown in the main manuscript. The excitation time on every frame is kept constant. The small decrease in obtained diffusion coefficient as a function of frame time is explained by particles having a higher chance to move outside the field-of-view with larger jump distances.
Extended Data Fig. 7 TARDIS-JD extraction from data of Chenouard et al.
Tracking data from Chenouard et al.23 has been deteriorated (removing 54% of localizations), analysed via the ‘extract JD’-function of TARDIS, and compared to the ground-truth (GT) data. Four different conditions are analysed: MICROTUBULE, RECEPTOR, VESICLE, and VIRUS, corresponding to [constant velocity], [tethered motion, switching, any direction], [Brownian motion, any direction], and [same direction dynamics, switching between Brownian and linear] dynamics, respectively. Densities are indicated in subplot titles, while the field-of-view is ∼50-by-50 µAU in size. In all scenarios, TARDIS accurately extracts the ground-truth data, and the level of noise is decreasing with decreasing localization removal. The following TARDIS settings were used: Δt bins of 1–3; maximum jump distance of 1e-05 AU; background frames starting at frame-shift of 35, using in total 50 frames; 300 BG bins starting at 3.5e-06 AU.
Extended Data Fig. 8 E. coli RNA polymerase jump distance analysis after nearest-neighbour tracking.
Jump distance analysis of RNA polymerase in E. coli with (red) and without (black) rifampicin, (the same data presented in Fig. 3b), via nearest-neighbour tracking. Notice the changing peak position and abundance as a function of localization density. The data is shown in a linear X-scale (left) and logarithmic X-scale (right).
Extended Data Fig. 9 Detailed information on kinetically state-changing particles.
Individual fit information of data presented in Fig. 3c, along with individual fit information for differing binding/unbinding kinetics (titles indicate kon / koff / Diffusion coefficient). For every condition 10.000 ‘true positive’ trajectories were simulated, and 10 repetitions of this were analysed with random start positions in TARDIS, and compared to analysing the same data with anaDDA on the ground-truth trajectory data. TP, Sp and me indicate true positive, spurious, and membrane localizations, respectively. All boxplots show the median as the central mark, with the 25th and 75th percentile as lower and upper edges. Whiskers extend to non-outlier extreme points, and outlier points are plotted as plusses.
Extended Data Fig. 10 Accuracy of software by Wolf et al44.
The same data analysed by TARDIS in Fig. 2b (a) and Extended Data Fig. 3b (b), analysed via the DANAE software44, which effectively only performs TARDIS-JD extraction. This is then fitted with a diffusive model afterwards. The accuracy, especially at high complexity scenarios, is worse compared to TARDIS. Additionally, DANAE shows bias towards too high values (right), which is caused by imperfect inter-particle distance distribution subtraction. DANAE is repeated 10 times on 20.000 simulated localizations for each condition. All boxplots show the median as the central mark, with the 25th and 75th percentile as lower and upper edges. Whiskers extend to non-outlier extreme points, and outlier points are plotted as plusses.
Supplementary information
Supplementary Information
Supplementary Notes 1–10 and Figs. 1–15.
Supplementary Software 1
TARDIS version 1.20, including Getting Started instructions, a manual, a program installation executable and MATLAB code.
Source data
All source data
Unprocessed video (including localization lists) and simulation data. Unprocessed TARDIS results data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Martens, K.J.A., Turkowyd, B., Hohlbein, J. et al. Temporal analysis of relative distances (TARDIS) is a robust, parameter-free alternative to single-particle tracking. Nat Methods (2024). https://doi.org/10.1038/s41592-023-02149-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41592-023-02149-7