Abstract
General anesthesia (GA) can produce analgesia (loss of pain) independent of inducing loss of consciousness, but the underlying mechanisms remain unclear. We hypothesized that GA suppresses pain in part by activating supraspinal analgesic circuits. We discovered a distinct population of GABAergic neurons activated by GA in the mouse central amygdala (CeAGA neurons). In vivo calcium imaging revealed that different GA drugs activate a shared ensemble of CeAGA neurons. CeAGA neurons also possess basal activity that mostly reflects animals’ internal state rather than external stimuli. Optogenetic activation of CeAGA potently suppressed both pain-elicited reflexive and self-recuperating behaviors across sensory modalities and abolished neuropathic pain-induced mechanical (hyper-)sensitivity. Conversely, inhibition of CeAGA activity exacerbated pain, produced strong aversion and canceled the analgesic effect of low-dose ketamine. CeAGA neurons have widespread inhibitory projections to many affective pain-processing centers. Our study points to CeAGA as a potential powerful therapeutic target for alleviating chronic pain.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All raw data described in this study are available from the corresponding authors upon reasonable request. All codes described in this study are available from the corresponding authors upon reasonable request. The MIN1PIPE code for processing miniscope imaging (correspondence should be addressed to Jinghao Lu, jinghao.lu@duke.edu) is available at: https://github.com/JinghaoLu/MIN1PIPE.
References
Aranake, A. et al. Increased risk of intraoperative awareness in patients with a history of awareness. Anesthesiology 119, 1275–1283 (2013).
Sebel, P. S. et al. The incidence of awareness during anesthesia: a multicenter United States study. Anesth. Analg. 99, 833–839 (2004).
Bergman, S. A. Ketamine: review of its pharmacology and its use in pediatric anesthesia. Anesth. Prog. 46, 10–20 (1999).
Sadove, M. S., Shulman, M., Hatano, S. & Fevold, N. Analgesic effects of ketamine administered in subdissociative doses. Anesth. Analg. 50, 452–457 (1971).
Gorlin, A. W., Rosenfeld, D. M. & Ramakrishna, H. Intravenous sub-anesthetic ketamine for perioperative analgesia. J. Anaesthesiol. Clin. Pharmacol. 32, 160–167 (2016).
Brown, E. N., Purdon, P. L. & Van Dort, C. J. General anesthesia and altered states of arousal: a systems neuroscience analysis. Annu. Rev. Neurosci. 34, 601–628 (2011).
Jiang-Xie, L.-F. et al. A common neuroendocrine substrate for diverse general anesthetics and sleep. Neuron 102, 1053–1065 (2019).
Haubensak, W. et al. Genetic dissection of an amygdala microcircuit that gates conditioned fear. Nature 468, 270–276 (2010).
Kim, J., Zhang, X., Muralidhar, S., LeBlanc, S. A. & Tonegawa, S. Basolateral to central amygdala neural circuits for appetitive behaviors. Neuron 93, 1464–1479 (2017).
Choi, H. M. T. et al. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. Development 145, dev165753 (2018).
Sakurai, K. et al. Capturing and manipulating activated neuronal ensembles with cane delineates a hypothalamic social-fear circuit. Neuron 92, 739–753 (2016).
Rodriguez, E. et al. A craniofacial-specific monosynaptic circuit enables heightened affective pain. Nat. Neurosci. 20, 1734–1743 (2017).
McCullough, K. M., Morrison, F. G., Hartmann, J., Carlezon, W. A. & Ressler, K. J. Quantified coexpression analysis of central amygdala subpopulations. eNeuro 5, ENEURO.0010-18.2018 (2018).
Sadler, K. E. et al. Divergent functions of the left and right central amygdala in visceral nociception. Pain 158, 747–759 (2017).
Ji, G. & Neugebauer, V. Hemispheric lateralization of pain processing by amygdala neurons. J. Neurophysiol. 102, 2253–2264 (2009).
Baas, D., Aleman, A. & Kahn, R. S. Lateralization of amygdala activation: a systematic review of functional neuroimaging studies. Brain Res. Brain Res. Rev. 45, 96–103 (2004).
Lu, J. et al. MIN1PIPE: A miniscope 1-photon-based calcium imaging signal extraction pipeline. Cell Rep. 23, 3673–3684 (2018).
Sheintuch, L. et al. Tracking the same neurons across multiple days in Ca2+ imaging data. Cell Rep. 21, 1102–1115 (2017).
Boyden, E. S., Zhang, F., Bamberg, E., Nagel, G. & Deisseroth, K. Millisecond-timescale, genetically targeted optical control of neural activity. Nat. Neurosci. 8, 1263–1268 (2005).
Chow, B. Y. et al. High-performance genetically targetable optical neural silencing by light-driven proton pumps. Nature 463, 98–102 (2010).
Huang, T. et al. Identifying the pathways required for coping behaviours associated with sustained pain. Nature 565, 86–90 (2019).
Hunskaar, S. & Hole, K. The formalin test in mice: dissociation between inflammatory and non-inflammatory pain. Pain 30, 103–114 (1986).
Basbaum, A. I., Bautista, D. M., Scherrer, G. & Julius, D. Cellular and molecular mechanisms of pain. Cell 139, 267–284 (2009).
Ding, W. et al. An improved rodent model of trigeminal neuropathic pain by unilateral chronic constriction injury of distal infraorbital nerve. J. Pain 18, 899–907 (2017).
Giglio, J. A. & Gregg, J. M. Development of mirror pain following trigeminal nerve injury: a case report and review of neuropathic mechanisms. Gen. Dent. 66, 27–32 (2018).
Milligan, E. D. et al. Spinal glia and proinflammatory cytokines mediate mirror-image neuropathic pain in rats. J. Neurosci. 23, 1026–1040 (2003).
Cai, Y.-Q., Wang, W., Paulucci-Holthauzen, A. & Pan, Z. Z. Brain circuits mediating opposing effects on emotion and pain. J. Neurosci. 38, 6340–6349 (2018).
Han, S., Soleiman, M. T., Soden, M. E., Zweifel, L. S. & Palmiter, R. D. Elucidating an affective pain circuit that creates a threat memory. Cell 162, 363–374 (2015).
Wang, D. et al. Functional divergence of Δ and μ opioid receptor organization in CNS pain circuits. Neuron 98, 90–108.e5 (2018).
Corder, G. et al. An amygdalar neural ensemble that encodes the unpleasantness of pain. Science 363, 276–281 (2019).
Levin-Arama, M. et al. Subcutaneous compared with intraperitoneal ketamine–xylazine for anesthesia of mice. J. Am. Assoc. Lab. Anim. Sci. 55, 794–800 (2016).
Beecher, H. K. Pain in men wounded in battle. Bull. US Army Med. Dep. 5, 445–454 (1946).
Petrovic, P., Kalso, E., Petersson, K. M. & Ingvar, M. Placebo and opioid analgesia-imaging a shared neuronal network. Science 295, 1737–1740 (2002).
Benedetti, F. Placebo and endogenous mechanisms of analgesia. Handb. Exp. Pharmacol. 393–413 (2007).
Eippert, F. et al. Activation of the opioidergic descending pain control system underlies placebo analgesia. Neuron 63, 533–543 (2009).
Basbaum, A. I. & Fields, H. L. Endogenous pain control systems: brainstem spinal pathways and endorphin circuitry. Annu. Rev. Neurosci. 7, 309–338 (1984).
Ossipov, M. H., Dussor, G. O. & Porreca, F. Central modulation of pain. J. Clin. Invest. 120, 3779–3787 (2010).
Lau, B. K. & Vaughan, C. W. Descending modulation of pain: the GABA disinhibition hypothesis of analgesia. Curr. Opin. Neurobiol. 29, 159–164 (2014).
Butler, R. K. & Finn, D. P. Stress-induced analgesia. Prog. Neurobiol. 88, 184–202 (2009).
Wager, T. D., Scott, D. J. & Zubieta, J.-K. Placebo effects on human μ-opioid activity during pain. Proc. Natl Acad. Sci. USA 104, 11056–11061 (2007).
Werka, T. & Marek, P. Post-stress analgesia after lesions to the central nucleus of the amygdala in rats. Acta Neurobiol. Exp. 50, 13–22 (1990).
Duvarci, S. & Pare, D. Amygdala microcircuits controlling learned fear. Neuron 82, 966–980 (2014).
Herry, C. & Johansen, J. P. Encoding of fear learning and memory in distributed neuronal circuits. Nat. Neurosci. 17, 1644–1654 (2014).
Neugebauer, V. Amygdala pain mechanisms. Handb. Exp. Pharmacol. 227, 261–284 (2015).
Parikh, D. et al. Stress-induced analgesia and endogenous opioid peptides: the importance of stress duration. Eur. J. Pharmacol. 650, 563–567 (2011).
Wilson, T. D. et al. Dual and opposing functions of the central amygdala in the modulation of pain. Cell Rep. 29, 332–346 (2019).
Yu, K. et al. The central amygdala controls learning in the lateral amygdala. Nat. Neurosci. 20, 1680–1685 (2017).
Cui, Y. et al. A central amygdala-substantia innominata neural circuitry encodes aversive reinforcement signals. Cell Rep. 21, 1770–1782 (2017).
Cai, H., Haubensak, W., Anthony, T. E. & Anderson, D. J. Central amygdala PKC-δ+ neurons mediate the influence of multiple anorexigenic signals. Nat. Neurosci. 17, 1240–1248 (2014).
Venza, M. et al. The overriding of TRAIL resistance by the histone deacetylase inhibitor MS-275 involves c-myc up-regulation in cutaneous, uveal, and mucosal melanoma. Int. Immunopharmacol. 28, 313–321 (2015).
Pang, D. et al. Polyphyllin II inhibits liver cancer cell proliferation, migration and invasion through downregulated cofilin activity and the AKT/NF-κB pathway. Biol. Open 9, bio046854 (2020).
Faget, L. et al. Opponent control of behavioral reinforcement by inhibitory and excitatory projections from the ventral pallidum. Nat. Commun. 9, 849 (2018).
Zamparo, I. et al. Axonal odorant receptors mediate axon targeting. Cell Rep. 29, 4334–4348 (2019).
Hara, Y. et al. Anti-tumor effects of an antagonistic mAb against the ASCT2 amino acid transporter on KRAS-mutated human colorectal cancer cells. Cancer Med. 9, 302–312 (2020).
Simon, C. M. et al. Stasimon contributes to the loss of sensory synapses and motor neuron death in a mouse model of spinal muscular atrophy. Cell Rep. 29, 3885–3901.e5 (2019).
Van Segbroeck, M., Knoll, A. T., Levitt, P. & Narayanan, S. MUPET-Mouse ultrasonic profile extraction: a signal processing tool for rapid and unsupervised analysis of ultrasonic vocalizations. Neuron 94, 465–485.e5 (2017).
Zhang, Y., Chen, Y., Liedtke, W. & Wang, F. Lack of evidence for ectopic sprouting of genetically labeled Aβ touch afferents in inflammatory and neuropathic trigeminal pain. Mol. Pain 11, 18 (2015).
Zhang, Y. et al. Identifying local and descending inputs for primary sensory neurons. J. Clin. Invest. 125, 3782–3794 (2015).
Acknowledgements
We thank J. Takatoh for helping with optimizing image compression that retains good resolution and for designing the color scheme for projection data. We also thank V. Prevosto and P. Thompson for helping with statistics and R. Mooney, R.-R. Ji and members of the Wang lab for critical reading of this manuscript. We thank the Janelia GENIE project for making GCaMP6m sensor available to the research community. Correspondence for calcium imaging data processing codes should be addressed to Jinghao Lu (jinghao.lu@duke.edu). T.H. is supported by the NSF GRFP scholarship. Y.C. is supported by R01 DE027454. This work is supported by NIH DP1MH103908, a Holland-Trice Scholar award, a W. M. Keck Foundation grant, NIH R01 NS109947 and NIH R01 DE029342.
Author information
Authors and Affiliations
Contributions
F.W. conceived the initial idea. F.W., T.H. and B.C. designed experiments with help from K.S. T.H. was responsible for immunohistochemical analysis, HCR in situ hybridization, all optogenetic manipulations during all types of sensory and affective behaviors and for generating and analyzing axon projection data. B.C. was responsible for all miniscope GCaMP6 imaging experiments under different conditions. J.L. was responsible for analysis of all imaging data and EEG data (blinded to experimental conditions). D.L. performed stress responses, mating experiments and EEG recordings and analyzed some of the behavioral data. K.S. performed some initial experiments. J.K. did some of the quantifications of histology and behavior results. S.Z. produced all CANE-LV viruses. S.Z. and D.L. performed some of the two-color molecular marker analysis. B.-X.H. took care of mouse husbandry and genotyping. L.Y. performed some slice recording experiments (data not shown). Y.C. performed chronic ligation surgeries. F.W., T.H. and B.C. wrote the manuscript with help from J.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Neuroscience thanks Gregory Corder, Nora McCall, Katelyn Sadler, Jessica Wojick and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Isoflurane general anesthesia activates different neuronal ensembles and molecular marker analysis of CeAGA neurons.
General anesthesia activated the a, central amygdala (CeA), b, bed nucleus of stria terminalis (BNST), and c, super optic nucleus (SON). Representative images two-color experiments examining the expression of various markers in Fos+ CeAGA neurons (induced by isoflurane). d, CeAGA neurons are a subset of GABAergic (vGAT) neurons in the central amygdala and all CeAGA express vGAT-GFP. e–i, CeAGA neurons have minimal to no overlap with e, somatostatin (SST), f, prodynorphin (Pdyn), g, neurotensin (NTS), and h, i, calcitonin gene-related peptide receptor (CGRPR) expressing neurons in CeA.
Extended Data Fig. 2 Representative post hoc histology and session distributions of activity of CANE-captured isoflurane-activated CeAGA neurons in response to isoflurane or ketamine.
a, Representative images of CANE-GCaMP6m+ neurons (green) and isoflurane-activated Fos+ neurons (red) and their overlap. Dotted box showing the placement of the GRIN lens in CeA. b, Quantification of the percent colocalization of Iso:Fos&GCaMP6m:GFP/all GCaMP6m (75.45 ± 5.04%) and Iso:Fos&GCaMP6m:GFP/all Fos (50.1 ± 8.70%) (n = 6 animals). Data are mean ± s.e.m. c, Three consecutive representative images of post hoc histology showing optic fiber tract into the CeA from bregma −1.00 mm to −1.16 mm. Optic fiber diameter is 200 µm. d, Session-wise percentage distribution of iso.-sustained and iso.-transient neurons from Fig. 2d. e, Session-wise percentage distribution of ketamine-sustained and ketamine-transient neurons from Fig. 2f. f, g, Session-wise percentage distribution of the isoflurane-activated neurons (f), or the ketamine-activated neurons (g), calculated using four methods calculated based on the ratio between the post- and pre-stimulation (that is anesthetic administration) activity. Cyan, Eff./Eff., mean activity of (post-stim) effective time / mean activity of (pre-stim) effective time. Orange, Mean/Mean, mean activity of all the (post-stim) time /mean activity of all the (pre-stim) time. Yellow, Eff./Mean, mean activity of (post-stim) effective time / mean activity of all the (pre-stim) time. Purple, Eff. time, total (post-stim) effective time. “Effective time” refers to the time points of a neural trace whose intensity is two median absolute deviation above its mean. Only these time points were considered to compute the effective mean. See Methods for details of the 4 methods. Pre-, pre-stimuli, awake state (−4–0 min). Post-, post-stimuli, isoflurane or ketamine (0–20 min).
Extended Data Fig. 3 CANE-captured CeAGA neurons are mostly inhibited by stress.
a, Schematic of CANE captured isoflurane-activated CeAGA neurons followed by a second exposure to restraint stress (left, for Fos expression, right, for calcium imaging). b, Left, representative images of CANEISO-tdTomato neurons (red) and stress-activated Fos+ neurons (green) and their overlap. (n = 3 animals). Right, representative images of isoflurane-only activated Fos+ neurons (green, repeated experiments n = 5 biologically independent samples.) and compared to stress-only activated Fos+ neurons (green, repeated experiments n = 3 biologically independent samples.) in the CeA. c, Heatmap, activity patterns of CANEISO-GCaMP6m captured CeAGA neurons in stress experiment sorted by the average activity during stress period (-8–0 min, pre-stress; 0–8 min, restraint stress; 8 -16 min, post-stress. 282 neurons from 5 mice × 2 trials). A small number of neurons (in the bottom of the heatmap) were activated by stress. d, Example traces of CANEISO-GCaMP6m captured CeAGA neurons in stress experiment showing both stress-inhibited and stress-activated neurons. Norm. intensity: normalized calcium signals rescaled to 0–1. e, Scatter plots of the tracked same neurons based on isoflurane and stress related activity patterns. Each dot is calculated based on effectiveness corrected activity ratio between the post- and pre-stimulus periods for isoflurane-stimulus and stress-stimulus, separately. Corrected ratio, mean activity of (post-) effective time / mean activity of (pre-) effective time. Single dots represent individual neurons in the logarithmic scale coordinates, and the circled dots represent robustly firing neurons, with the maximum intensity of each neuron in the whole duration exceeding a threshold. f, Same plots with isoflurane responses calculated by actively firing time exceeding a threshold (1 min). Active duration, total (post-) effective time of effective moment >1 min. “Effective time” refers to the time points of a neural trace whose intensity is two median absolute deviation above its mean. Only these time points were considered to compute the mean. g, Neuron count summary of e. Left, Neuron count distribution of activity of isoflurane-suppressed neurons during stress. Right, Neural count distribution of activity of isoflurane-activated neurons during stress. h, Neural count distribution of f. Left, marginal count distribution of activity of CANEISO-GCaMP6m captured CeAGA neurons during stress. Right, marginal count distribution of activity of CANEISO-GCaMP6m captured CeAGA neurons during isoflurane GA.
Extended Data Fig. 4 Manipulation of CeAGA neurons did not induce anxiety-like or fear-like behavior or change the gross brain state.
a, Schematics of the Elevated Plus Maze (left) and Open Field (right) apparatus. b, Quantification of total time spent in the inner (GFP: 15.67 ± 5.97 s (baseline), 14.18 ± 7.24 s (stim), 18.07 ± 6.38 s (post); ChR2: 22.59 ± 4.92 s (baseline), 35.90 ± 10.22 s (stim), 29.73 ± 6.49 s (post); eArch: 5.69 ± 2.32 s (baseline), 11.01 ± 6.61 s (stim), 8.54 ± 5.20 s (post)) and outer perimeter (GFP: 284.32 ± 6.01 s (baseline), 285.70 ± 7.25 s (stim), 281.80 ± 6.39 s (post); ChR2: 277.31 ± 4.92 s (baseline), 264.00 ± 10.22 s (stim), 270.17 ± 6.47 s (post); eArch: 294.31 ± 2.32 s (baseline), 288.99 ± 6.61 s (stim), 291.46 ± 5.20 s (post)) of the Open Field Test (control, n = 8 animals, ChR2, n = 8 animals, eArch, n = 6 animals; two-way repeated measures ANOVA; P-value was above 0.05, no significance; F4,42 = 0.8743 (inner), F4,42 = 0.8633 (outer)). Data are mean ± s.e.m. c, Quantification of total distance travelled (GFP: 4.27 ± 0.73 m (baseline), 4.36 ± 0.72 m (stim), 3.36 ± 0.51 m (post); ChR2: 6.92 ± 0.87 m (baseline), 5.91 ± 0.88 m (stim), 5.85 ± 0.95 m (post); eArch: 4.59 ± 0.56 m (baseline), 5.45 ± 0.99 m (stim), 3.91 ± 0.59 m (post)), and d, total time spent in the open (GFP: 38.53 ± 12.26 s (baseline), 19.19 ± 6.65 s (stim), 11.03 ± 3.44 s (post); ChR2: 28.61 ± 9.69 s (baseline), 21.21 ± 6.31 s (stim), 27.91 ± 5.76 s (post); eArch (24.33 ± 4.70 s (baseline), 24.85 ± 4.98 s (stim), 14.35 ± 6.56 s (post)) and closed arms (GFP: 239.16 ± 17.28 s (baseline), 270.86 ± 8.54 s (stim), 280.76 ± 4.61 s (post); ChR2: 248.58 ± 11.66 s (baseline), 258.80 ± 9.36 s (stim), 261.08 ± 7.60 s (post); eArch (261.22 ± 4.07 s (baseline), 264.45 ± 5.50 s (stim), 272.48 ± 10.19 s (post)) of the Elevated Plus Maze (control, n = 8 animals, ChR2, n = 8 animals, eArch, n = 8 animals; two-way repeated measures ANOVA; P-value was above 0.05, no significance; F4,38 = 1.083 (distance), F4,38 = 1.402 (open arms), F4,38 = 1.355 (closed arms)). Data are mean ± s.e.m. e, Power spectrum of EEG signals in the frontal and parietal cortex. Left, laser on, right, laser off. f, Overlap of power spectrum of EEG signals in the frontal and parietal cortex from e. The mean spectrum in each condition was calculated from the average across 9 sessions (n = 3 mice, 2 min laser on / 2 min laser off, 3 repetitions in each mouse). The error bar represents the standard error. The power spectrum was normalized.
Extended Data Fig. 5 Manipulations of CeAGA neurons modulated reflexive withdrawal threshold to von Frey filaments and activation of CeAGA neurons did not alter courtship behaviors.
a, b, Quantifications of the withdrawal threshold to von Frey filaments applied to the whisker pad in a, naïve (control, n = 9 animals (0.44 ± 0.0g5 (ipsi-off), 0.39 ± 0.05g (ipsi-on), 0.39 ± 0.06g (contra-off), 0.36 ± 0.06g (contra-on)), ChR2, n = 8 animals (0.34 ± 0.06g (ipsi-off), 0.95 ± 0.05g (ipsi-on), 0.52 ± 0.12g (contra-off), 0.93 ± 0.08g (contra-on)), eArch, n = 7 animals (0.21 ± 0.07g (ipsi-off), 0.06 ± 0.01g (ipsi-on), 0.35 ± 0.07g (contra-off), 0.22 ± 0.05g (contra-on)); two-way ANOVA; ****P < 0.0001, **P = 0.0042; F6,84 = 10.80) and b, IoN-CCI mice (control, n = 6 animals (0.28 ± 0.09g (off), 0.39 ± 0.14g (on)), ChR2, n = 6 animals (0.16 ± 0g (off), 0.87 ± 0.08g (on)); two-way ANOVA; **P = 0.0032; F1,20 = 10.54). c-d, Quantification of light-illumination induced changes in total syllable number of ultrasonic vocalizations (GFP: 23.25 ± 35.65; ChR2: 21.17 ± 44.17), or total duration of anogenital sniffing and mounting behavior (GFP: 16.25 ± 4.31 s; ChR2: 18.17 ± 4.53 s) in control CeAGA-GFP and CeAGA-ChR2 mice (with light - without light) during 2 min of social interactions (control, n = 4 animals, ChR2, n = 6 animals; unpaired t-test, two-tailed; P = 0.974, F5,3 = 2.303 (syllable number), P = 0.7791, F5,3 = 1.656 (anogenital sniffing)). Data are mean ± s.e.m.
Extended Data Fig. 6 Activation of the left CeAGA neurons modulated pain-related behaviors in naïve mice and acute pain models.
a, Quantification of effects of optogenetic activation of the left CeAGA neurons on the paw withdrawal frequency to six graded von Frey filaments ranging from 0.4 to 4.0 grams applied to the ipsilateral (Off: 0 ± 0 (0.40g), 2.50 ± 0.56 (0.60g), 5.00 ± 0.63 (1.0g), 6.33 ± 0.95 (1.40g), 8.33 ± 1.09 (2.0g), 10 ± 0 (4.0g); On: 0 ± 0 (0.40g), 0.33 ± 0.21 (0.60g), 2.50 ± 0.50 (1.0g), 3.67 ± 0.80 (1.40g), 5.33 ± 1.38 (2.0g), 9.33 ± 0.49 (4.0g)) or contralateral paw (Off: 0 ± 0 (0.40g), 2.00 ± 0.82 (0.60g), 6.83 ± 1.11 (1.0g), 7.83 ± 0.17 (1.40g), 8.83 ± 0.65 (2.0g), 9.83 ± 0.17 (4.0g); On: 0 ± 0 (0.40g), 0.67 ± 0.33 (0.60g), 3.00 ± 0.45 (1.0g),5.0 ± 0.37 (1.40g), 6.0 ± 1.15 (2.0g), 9.67 ± 0.21 (4.0g)) to the left CeA. (Ipsilateral and contralateral, ChR2, n = 6 animals; two-way ANOVA; *P = 0.0500 (2.0g), *P = 0.0217 (1.4g), ****P < 0.0001, **P = .0023 (1.4g), **P = .0029 (2.0g); F1,60 = 20.51 (ipsi), F1,60 = 28.81 (contra)). b, Quantification of optogenetic activation of the left CeAGA neurons showed that this manipulation did not induce any change in the head withdrawal frequency to eight von Frey filaments ranging from 0.008 to 1.0 gram applied to either the ipsilateral (Off: 0 ± 0 (0.008g), 0 ± 0 (0.02g), 0 ± 0 (0.04g), 1.0 ± 0.52 (0.07g), 4.33 ± 0.61 (0.16g), 6.50 ± 0.85 (0.40g), 9.67 ± 0.21 (0.60g), 10.0 ± 0 (1.0g); On: 0 ± 0 (0.008g), 0 ± 0 (0.02g), 0 ± 0 (0.04g), 0.67 ± 0.49 (0.07g), 4.17 ± 0.54 (016g), 7.00 ± 0.89 (0.40g), 9.83 ± 0.17 (0.60g), 10.0 ± 0 (1.0g)) or the contralateral whisker pad (Off: 0 ± 0 (0.008g), 0 ± 0 (0.02g), 0.50 ± 0.34 (0.04g), 2.50 ± 0.50 (0.07g), 4.67 ± 0.49 (0.16g), 8.0 ± 0.68 (0.40g), 10.0 ± 0 (0.60g), 10.0 ± 0 (1.0g); On: 0 ± 0 (0.008g), 0 ± 0 (0.02g), 0.17 ± 0.17 (0.04g), 2.33 ± 0.80 (0.07g), 5.17 ± 0.17 (016g), 8.17 ± 0.79 (0.40g), 9.83 ± 0.17 (0.60g), 10.0 ± 0 (1.0g)) to the left CeA. (Ipsilateral and contralateral, ChR2, n = 6 animals; two-way ANOVA; not significant P > 0.05; F1,80 = 0.9205 (ipsi), F1,40 = 0.000 (contra)). c, Quantification of the optogenetics induced changes in withdrawal latency (sec) to dry ice (2.29 ± 0.33 s (off-left), 6.86 ± 3.70 s (on-left), 2.33 ± 0.25 s (off-right), 5.81 ± 2.75 s (on-right)) (ChR2, n = 7 animals; one-way ANOVA; **P = 0.0045, *P = 0.0311; F3,24 = 6.241) and heat (6.61 ± 0.93 s (off-left), 12.94 ± 1.89 s (on-left), 6.09 ± 0.65 s (off-right), 15.04 ± 1.80 s (on-right)) (ChR2, n = 7 animals; one-way ANOVA; ****P < 0.0001; F3,24 = 59.93). d, Quantification of total licking and face wiping latency (sec) from left CeAGA neurons optogenetic activation after formalin injection during the second phase of inflammatory pain (134.33 ± 22.88 s (paw licking-off), 10.33 ± 8.24 s (paw licking-on), 151.67 ± 35.99 s (face wiping-off), 12.00 ± 8.74 s (face wiping-on)). (ChR2, n = 6 animals; one-way ANOVA; ****P < 0.0001; F3,20 = 59.54).
Extended Data Fig. 7 CeAGA activities are not correlated to the onsets of sensory stimuli.
Neuronal activity patterns during a, cold, b, heat, c, paw von Frey, and d, face von Frey tests sorted by neurons peak responses timing: from -10 to +10 sec, with 0 as the onset of stimulus application. Top of each heatmap, averaged population activity from 10 seconds before to 10 seconds after each stimulus onset. Thick lines indicated mean and shaded areas indicated s.e.m.
Extended Data Fig. 8 CeAGA neurons are distinct from pain-activated neurons in CeA and high magnification image of CeAGA neurons projection into the ipsilateral BLA.
a, CeAGA neurons have minimal co-localization with formalin-induced Fos+ cells. Formalin-activated cells primarily locate in the capsular division of CeA outside the lateral division where CeAGA locate. Insert i)-v), Example of five consecutive slices of the lateral division of CeA showing minimal co-localization with formalin-induced Fos+ cells with CeAGA neurons, and the quantification of fraction of co-colocalization between CANE-captured CeAGA cells and formalin-induced Fos+ cells (n = 5 biologically independent samples for each condition) (0.78 ± 0.06 (Iso/Cane-Iso); 0.22 ± 0.04 (Form/Cane-Iso)). b, Coronal schematic next to example coronal slice of low magnification (high exposure) of CANE-GFP labeled CeAGA neurons and their axons with a box around the ipsilateral BLA. Insert i)-ii), High mag images show projections in ipsilateral BLA with top panel showing that isoflurane did not induce Fos+ cells, and bottom panel showing that formalin-pain induced robust Fos+ expression in BLA. (n = 3 biologically independent samples).
Extended Data Fig. 9 Consistent axonal projections from CeAGA neurons.
In sequential order: a, frontal cortex, b, nucleus accumbens (NAc), c, striatum, d, insular, e, bed nucleus stria terminalis (BNST), f, intralaminar, g, temporal association cortex (TeA) and ectorhinal cortex (Ect), h, subthalamic nucelus (SubTh), i, periaqueductal grey (PAG), j, parabrachial nucleus (PBN), k, solitary tract (SolT), and l, reticular formation (RT). m, Quantification of the mean intensity value (artificial units) of the axonal projections from each region of interest (ROI) listed above (a-l) (n = 3 biologically independent samples) (59.40 ± 15.68 (FC), 74.38 ± 14.18 (NAc), 46.59 ± 3.21 (Striatum), 81.07 ± 8.86 (Ins), 112.09 ± 10.85 (BNST), 106.15 ± 15.69 (Intra), 115.35 ± 34.52 (TeA/Ect), 82.99 ± 11.06 (SubTh), 66.32 ± 9.84 (PAG), 116.29 ± 22.67 (PBN), 92.70 ± 6.27 (SolT), 53.26 ± 3.63 (RT)).
Extended Data Fig. 10 CeAGA neurons are also activated by low dose anesthetics.
a, Heatmaps, activity patterns of the same neurons tracked in isoflurane (1.5%) and low isoflurane (0.5%) experiments, aligned by isoflurane (1.5%) neural patterns. 106 tracked same neurons from 5 mice × 1 trial. b, Left, mean and difference traces of the population normalized activity in isoflurane and low isoflurane experiments. Right, intensity distribution of the traces. c, Heatmaps, activity patterns of the same neurons tracked in isoflurane (1.5%), ketamine (100 mg/kg) and low ketamine (12 mg/kg) experiments, aligned by isoflurane neural patterns. 69 tracked same neurons from 5 mice × 1 trial. d, Left, mean trace of the population normalized activity in isoflurane, ketamine and low ketamine experiments. Right, intensity distribution of the traces.
Supplementary information
Supplementary Information
Supplementary Table 1
Supplementary Video 1
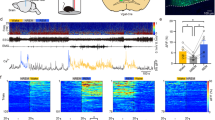
Video shows in vivo calcium imaging of the CeAGA neurons in freely moving mice during isoflurane-induced GA. Most imaged neurons showed calcium activities during isoflurane-induced GA. Twenty-two sessions from 12 mice were imaged for 1.5% isoflurane (ten mice × two trials and two mice × one trial).
Supplementary Video 2
Video shows that the transition from laser off to optogenetically activate CeAGA neurons did not induce anxiety- or fear-like behaviors (such as freezing, fleeing or spending more time in the perimeter zone) in open field tests. Repeated independently n = 8 with similar results.
Supplementary Video 3
In normal conditions, the innocuous 0.0–4-g von Frey filament does not elicit any responses. Upon bilateral optogenetic silencing the CeAGA neurons, mice displayed both low- and high-intensity withdrawal response to 0.04-g stimulation. Repeated independently n = 7 with similar results.
Supplementary Video 4
Formalin was injected into the hind paw and, upon activation of CeAGA neurons, the CeAGA-ChR2 mouse stopped the ongoing paw-licking behavior. Repeated independently n = 9 with similar results.
Supplementary Video 5
Formalin was injected into the whisker pad and, upon activation of CeAGA neurons, CeAGA-ChR2 mice stopped the ongoing face-wiping behavior. Repeated independently n = 9 with similar results.
Supplementary Video 6
Formalin was injected into the hind paw, and silencing CeAGA neurons in CeAGA-eArch mice re-induced licking of the injured paw. Repeated independently n = 7 with similar results.
Supplementary Video 7
In a CeAGA-ChR2 CCI-IoN mouse, without illumination, the 1-g von Frey filament elicited a strong nocifensive response. ChR2 activation of CeAGA neurons in the same mouse dramatically abolished withdrawal response to this painful (1-g) mechanical stimulus. Repeated independently n = 7 with similar results.
Supplementary Video 8
In a CeAGA-GFP CCI-IoN mouse, light illumination of CeA did not change the strong nocifensive responses to 1-g von Frey stimulation. Repeated independently n = 8 with similar results.
Rights and permissions
About this article
Cite this article
Hua, T., Chen, B., Lu, D. et al. General anesthetics activate a potent central pain-suppression circuit in the amygdala. Nat Neurosci 23, 854–868 (2020). https://doi.org/10.1038/s41593-020-0632-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-0632-8
This article is cited by
-
BNST GABAergic neurons modulate wakefulness over sleep and anesthesia
Communications Biology (2024)
-
The parabrachial to central amygdala pathway is critical to injury-induced pain sensitization in mice
Neuropsychopharmacology (2024)
-
The secondary somatosensory cortex gates mechanical and heat sensitivity
Nature Communications (2024)
-
Optogenetic Approach in Trigeminal Neuralgia and Potential Concerns: Preclinical Insights
Molecular Neurobiology (2024)
-
A common neuronal ensemble in nucleus accumbens regulates pain-like behaviour and sleep
Nature Communications (2023)