Abstract
The AAA+ ATPase spastin remodels microtubule arrays through severing and its mutation is the most common cause of hereditary spastic paraplegias (HSP). Polyglutamylation of the tubulin C-terminal tail recruits spastin to microtubules and modulates severing activity. Here, we present a ~3.2 Å resolution cryo-EM structure of the Drosophila melanogaster spastin hexamer with a polyglutamate peptide bound in its central pore. Two electropositive loops arranged in a double-helical staircase coordinate the substrate sidechains. The structure reveals how concurrent nucleotide and substrate binding organizes the conserved spastin pore loops into an ordered network that is allosterically coupled to oligomerization, and suggests how tubulin tail engagement activates spastin for microtubule disassembly. This allosteric coupling may apply generally in organizing AAA+ protein translocases into their active conformations. We show that this allosteric network is essential for severing and is a hotspot for HSP mutations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Electron microscopy map and the top scoring model of five atomic models obtained from an EM multi-model pipeline have been deposited at the Electron Microscopy Data Bank and Protein Data Bank under accession numbers EMD-20226 and PDB 6P07, respectively. All data used in this study are available from the corresponding authors upon reasonable request.
Change history
19 March 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
McnallyF. J., & Roll-MecakA. Microtubule-severing enzymes: from cellular functions to molecular mechanism. J. Cell Biol. 217, 4057 (2018).
Salinas, S. et al. Human spastin has multiple microtubule-related functions. J. Neurochem. 95, 1411–1420 (2005).
Eckert, T. et al. Spastin’s microtubule-bnding properties and comparison to katanin. PLoS ONE 7, 1–16 (2012).
Wen, M. & Wang, C. The nucleotide cycle of spastin correlates with its microtubule-binding properties. FEBS J. 280, 3868–3877 (2013).
Evans, K. J., Gomes, E. R., Reisenweber, S. M., Gundersen, G. G. & Lauring, B. P. Linking axonal degeneration to microtubule remodeling by Spastin-mediated microtubule severing. J. Cell Biol. 168, 599–606 (2005).
Roll-Mecak, A. & Vale, R. D. Structural basis of microtubule severing by the hereditary spastic paraplegia protein spastin. Nature 451, 363–367 (2008).
Roll-Mecak, A. & Vale, R. D. The Drosophila homologue of the hereditary spastic paraplegia protein, spastin, severs and disassembles microtubules. Curr. Biol. 15, 650–655 (2005).
Hazan, J. et al. Spastin, a new AAA protein, is altered in the most frequent form of autosomal dominant spastic paraplegia. Nat. Genet. 23, 296–303 (1999).
Blackstone, C., O’Kane, C. J. & Reid, E. Hereditary spastic paraplegias: membrane traffic and the motor pathway. Nat. Rev. Neurosci. 12, 31–42 (2011).
Solowska, J. M. & Baas, P. W. Hereditary spastic paraplegia SPG4: what is known and not known about the disease. Brain 138, 2471–2484 (2015).
Stone, M. C. et al. Normal spastin gene dosage Is specifically required for axon regeneration. Cell Rep. 2, 1340–1350 (2012).
Havlicek, S. et al. Gene dosage-dependent rescue of HSP neurite defects in SPG4 patients’ neurons. Hum. Mol. Genet. 23, 2527–2541 (2014).
White, S. R., Evans, K. J., Lary, J., Cole, J. L. & Lauring, B. Recognition of C-terminal amino acids in tubulin by pore loops in Spastin is important for microtubule severing. J. Cell Biol. 176, 995–1005 (2007).
Zehr, E. et al. Katanin spiral and ring structures shed light on power stroke for microtubule severing. Nat. Struct. Mol. Biol. 24, 717–725 (2017).
Hartman, J. J. & Vale, R. D. Microtubule disassembly by ATP-dependent oligomerization of the AAA enzyme katanin. Science 286, 782–785 (1999).
Eckert, T. et al. Subunit interactions and cooperativity in the microtubule-severing AAA ATPase spastin. J. Biol. Chem. 287, 26278–26290 (2012).
Cummings, C. M., Bentley, C. A., Perdue, S. A., Baas, P. W. & Singer, J. D. The Cul3/Klhdc5 E3 ligase regulates p60/katanin and is required for normal mitosis in mammalian cells. J. Biol. Chem. 284, 11663–11675 (2009).
Lu, C., Srayko, M. & Mains, P. E. The Caenorhabditis elegans microtubule-severing complex MEI-1/MEI-2 katanin interactsdifferently with two superficially redundant β-tubulin isotypes. Mol. Biol. Cell 15, 142–150 (2004).
Sherwood, N. T., Sun, Q., Xue, M., Zhang, B. & Zinn, K. Drosophila spastin regulates synaptic microtubule networks and is required for normal motor function. PLoS Biol. 2, e429 (2004).
Valenstein, M. L. & Roll-Mecak, A. Graded control of microtubule severing by tubulin glutamylation. Cell 164, 911–921 (2016).
Vemu, A. et al. Severing enzymes amplify microtubule arrays through lattice GTP-tubulin incorporation. Science 361, eaau1504 (2018).
Itzhak, D. N., Tyanova, S., Cox, J. & Borner, G. H. H. Global, quantitative and dynamic mapping of protein subcellular localization. eLife 5, 1–36 (2016).
Geimer, S., Teltenkötter, A., Plessmann, U., Weber, K. & Lechtreck, K. F. Purification and characterization of basal apparatuses from a flagellate green alga. Cell Motil. Cytoskelet. 37, 72–85 (1997).
Schneider, A., Plessmann, U., Felleisen, R. & Weber, K. Posttranslational modifications of trichomonad tubulins; identification of multiple glutamylation sites. FEBS Lett. 429, 399–402 (1998).
Abid Ali, F. et al. Cryo-EM structures of the eukaryotic replicative helicase bound to a translocation substrate. Nat. Commun. 7, 10708 (2016).
Skordalakes, E. & Berger, J. M. Structure of the Rho transcription terminator: mechanism of mRNA recognition and helicase loading. Cell 114, 135–146 (2003).
Taylor, J. L., White, S. R., Lauring, B. & Kull, F. J. Crystal structure of the human spastin AAA domain. J. Struct. Biol. 179, 133–137 (2012).
Han, H. et al. binds ESCRT-III substrates through a repeating array of dipeptide-binding pockets. eLife 6, 1–15 (2017).
Gates, S. N. et al. Ratchet-like polypeptide translocation mechanism of the AAA+disaggregase Hsp104. Science 357, 273–279 (2017).
De la Peña, A. H., Goodall, E. A., Gates, S. N., Lander, G. C. & Martin, A. Substrate-engaged 26S proteasome structures reveal mechanisms for ATP-hydrolysis–driven translocation. Science 362, eaav0725 (2018).
Augustyniak, R. & Kay, L. E. Cotranslocational processing of the protein substrate calmodulin by an AAA+unfoldase occurs via unfolding and refolding intermediates. Proc. Natl Acad. Sci. USA 115, E4786–E4795 (2018).
Schlieker, C. et al. Substrate recognition by the AAA+chaperone ClpB. Nat. Struct. Mol. Biol. 11, 607–615 (2004).
Puchades, C. et al. Structure of the mitochondrial inner membrane AAA+protease YME1 gives insight into substrate processing. Science 358, eaao0464 (2017).
Ripstein, Z. A., Huang, R., Augustyniak, R., Kay, L. E. & Rubinstein, J. L. Structure of a AAA+unfoldase in the process of unfolding substrate. eLife 6, 1–14 (2017).
Alfieri, C., Chang, L. & Barford, D. Mechanism for remodelling of the cell cycle checkpoint protein MAD2 by the ATPase TRIP13. Nature 559, 274–278 (2018).
Scott, A. et al. Structural and mechanistic studies of VPS4 proteins. EMBO J. 24, 3658–3669 (2005).
Roll-Mecak, A. Intrinsically disordered tubulin tails: complex tuners of microtubule functions? Semin. Cell Dev. Biol. 37, 11–19 (2015).
Hinnerwisch, J., Fenton, W. A., Furtak, K. J., Farr, G. W. & Horwich, A. L. Loops in the central channel of ClpA chaperone mediate protein binding, unfolding, and translocation. Cell 121, 1029–1041 (2005).
Lee, J. et al. Structural determinants for protein unfolding and translocation by the Hsp104 protein disaggregase. Biosci. Rep. 37, BSR20171399 (2017).
Charvin, D. et al. Mutations of SPG4 are responsible for a loss of function of spastin, an abundant neuronal protein localized in the nucleus. Hum. Mol. Genet. 12, 71–78 (2003).
Mészárosová, A. U. et al. SPAST mutation spectrum and familial occurrence among Czech patients with pure hereditary spastic paraplegia. J. Hum. Genet. 61, 845–850 (2016).
Ishiura, H. et al. Molecular epidemiology and clinical spectrum of hereditary spastic paraplegia in the Japanese population based on comprehensive mutational analyses. J. Hum. Genet. 59, 163–172 (2014).
Depienne, C. et al. Spastin mutations are frequent in sporadic spastic paraparesis and their spectrum is different from that observed in familial cases. J. Med. Genet. 43, 259–265 (2006).
Tang, B. S. et al. Clinical features of hereditary spastic paraplegia with thin corpus callosum: report of 5 Chinese cases. Chin. Med. J. (Engl.) 117, 1002–1005 (2004).
Hentati, A. et al. Novel mutations in spastin gene and absence of correlation with age at onset of symptoms. Neurology 55, 1388–1390 (2000).
Meijer, I. A., Hand, C. K., Cossette, P., Figlewicz, D. A. & Rouleau, G. A. Spectrum of SPG4 mutations in a large collection of North American families with hereditary spastic paraplegia. Arch. Neurol. 59, 281–286 (2002).
Patrono, C. et al. Autosomal dominant hereditary spastic paraplegia: DHPLC-based mutation analysis of SPG4 reveals eleven novel mutations. Hum. Mutat. 25, 506 (2005).
Balicza, P. et al. Genetic background of the hereditary spastic paraplegia phenotypes in Hungary - An analysis of 58 probands. J. Neurol. Sci. 364, 116–121 (2016).
Bürger, J. et al. Hereditary spastic paraplegia caused by mutations in the SPG4 gene. Eur. J. Hum. Genet. 8, 771–776 (2000).
Elert-Dobkowska, E. et al. Molecular spectrum of the SPAST, ATL1 and REEP1 gene mutations associated with the most common hereditary spastic paraplegias in a group of Polish patients. J. Neurol. Sci. 359, 35–39 (2015).
Lu, X. et al. Genetic analysis of SPG4 and SPG3A genes in a cohort of Chinese patients with hereditary spastic paraplegia. J. Neurol. Sci. 347, 368–371 (2014).
Park, H. et al. Mutational spectrum of the SPAST and ATL1 genes in Korean patients with hereditary spastic paraplegia. J. Neurol. Sci. 357, 167–172 (2015).
Polymeris, A. A. et al. A series of Greek children with pure hereditary spastic paraplegia: clinical features and genetic findings. J. Neurol. 263, 1604–1611 (2016).
Aulitzky, A. et al. A complex form of hereditary spastic paraplegia in three siblings due to somatic mosaicism for a novel SPAST mutation in the mother. J. Neurol. Sci. 347, 352–355 (2014).
McDermott, C. J. et al. Clinical features of hereditary spastic paraplegia due to spastin mutation. Neurology 67, 45–51 (2006).
Yabe, I., Sasaki, H. & Tashiro, K. Spastin gene mutation in Japanese with hereditary spastic paraplegia. J. Med. 39, 14–15 (2002).
Dong, E. L. et al. Clinical spectrum and genetic landscape for hereditary spastic paraplegias in China. Mol. Neurodegener. 13, 1–14 (2018).
Luo, Y. et al. A diagnostic gene chip for hereditary spastic paraplegias. Brain Res. Bull. 97, 112–118 (2013).
Shoukier, M. et al. Expansion of mutation spectrum, determination of mutation cluster regions and predictive structural classification of SPAST mutations in hereditary spastic paraplegia. Eur. J. Hum. Genet. 17, 187–194 (2009).
Fonknechten, N. et al. Spectrum of SPG4 mutations in autosomal dominant spastic paraplegia. Hum. Mol. Genet. 9, 637–644 (2000).
Proukakis, C., Moore, D., Labrum, R., Wood, N. W. & Houlden, H. Detection of novel mutations and review of published data suggests that hereditary spastic paraplegia caused by spastin (SPAST) mutations is found more often in males. J. Neurol. Sci. 306, 62–65 (2011).
Solowska, J. M. et al. Pathogenic Mutation of Spastin Has Gain-of-Function Effects on Microtubule Dynamics. J. Neurosci. 34, 1856–1867 (2014).
Qiang, L. et al. Hereditary spastic paraplegia: gain-of-function mechanisms revealed by new transgenic mouse. Hum. Mol. Genet. 28, 1136–1152 (2018).
Errico, A., Ballabio, A. & Rugarli, E. I. Spastin, the protein mutated in autosomal dominant hereditary spastic paraplegia, is involved in microtubule dynamics. Hum. Mol. Genet. 11, 153–163 (2002).
Crippa, F. et al. Eight novel mutations in SPG4 in a large sample of patients with hereditary spastic paraplegia. Arch. Neurol. 63, 750–755 (2006).
França, M. C. et al. SPG4-related hereditary spastic paraplegia: frequency and mutation spectrum in Brazil. Clin. Genet. 86, 194–196 (2014).
Kim, T.-H. et al. Mutation analysis of SPAST, ATL1, and REEP1 in Korean patients with hereditary spastic paraplegia. J. Clin. Neurol. 10, 257–261 (2014).
Gillespie, M. K., Humphreys, P., McMillan, H. J. & Boycott, K. M. Association of early-onset spasticity and risk for cognitive impairment with mutations at amino acid 499 in SPAST. J. Child Neurol. 33, 329–332 (2018).
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Ziolkowska, N. & Roll-Mecak, A. In vitro microtubule severing assays. Neurochem. Int. 37, 399–400 (2013).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and Lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Brown, P. H., Balbo, A. & Schuck, P. Using prior knowledge in the determination of macromolecular size-distributions by analytical ultracentrifugation. Biomacromolecules 8, 2011–2024 (2007).
Carragher, B. et al. Leginon: an automated system for acquisition of images from vitreous ice specimens. J. Struct. Biol. 132, 33–45 (2000).
Voss, N. R., Yoshioka, C. K., Radermacher, M., Potter, C. S. & Carragher, B. DoG Picker and TiltPicker: software tools to facilitate particle selection in single particle electron microscopy. J. Struct. Biol. 166, 205–213 (2009).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Scheres, S. H. W. RELION: Implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Herzik, M. A., Fraser, J. S. & Lander, G. C. A Multi-model approach to assessing local and global cryo-EM map quality. Structure 27, 344–358 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Gell, C. et al. in Microtubule Dynamics. Methods in Molecular Biology (Methods and Protocols) Vol. 777 (ed. Straube, A.) (Humana Press, 2011).
Edelstein, A. D. et al. Advance methods of microscope control using microManager software. J. Biol. Methods 1, e10 (2014).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Cardone, G., Heymann, J. B. & Steven, A. C. One number does not fit all: mapping local variations in resolution in cryo-EM reconstructions. J. Struct. Biol. 184, 226–236 (2013).
Meng, E. C., Pettersen, E. F., Couch, G. S., Huang, C. C. & Ferrin, T. E. Tools for integrated sequence-structure analysis with UCSF Chimera. BMC Bioinformatics 7, 1–10 (2006).
Acknowledgements
We thank J.C. Ducom at The Scripps Research Institute High Performance Computing for computational support and B. Anderson at The Scripps Research Institute electron microscopy facility for microscope support. We thank M. Herzik and A. Hernandes for help with atomic modeling, E. Szczesna for help with microtubule severing assays, G. Piszcek from the Biophysics Core of the National Heart, Lung and Blood Institute (NHLBI) for help with AUC experiments, S. Chowdhury, C. Puchades and M. Wu for helpful discussion. C.R.S. was supported by a National Science Foundation predoctoral fellowship. G.C.L. was supported as a Searle Scholar, a Pew Scholar, an Amgen Young Investigator and by the National Institutes of Health (NIH) grant no. DP2EB020402. Computational analyses of EM data were performed using shared instrumentation funded by NIH grant no. S10OD021634 to G.C.L. A.R.M. was supported by the intramural programs of the National Institute of Neurological Disorders and Stroke (NINDS) and the NHLBI.
Author information
Authors and Affiliations
Contributions
C.R.S. froze EM grids, collected and processed EM data and built atomic models. A.S. purified all proteins, performed AUC and ATPase assays. E.A.Z. performed severing assays. C.R.S., G.C.L. and A.R.M. interpreted structural models and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Inês Chen was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Analytical ultracentrifugation and ATPase assays.
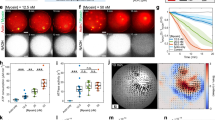
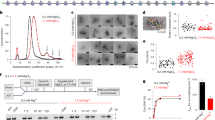
(a) AUC experimental distribution for the spastin Walker B mutant in the presence of ATP (Methods). The expected molecular weight of the spastin hexamer is 320 kDa. (b) Spastin ATPase stimulation by poly-glutamate (0.75–5 KDa) used for our structural determination; n=4; lines indicate mean and S.D. (c) AUC experimental distribution for the spastin Walker B mutant in the presence of ATP and Atto-488 labeled poly-glutamate peptide showing co-migration of the spastin hexamer and peptide. AUC experimental distribution for the spastin Walker B mutant alone at 6μM in the presence of ATP (top panel), Atto488-labeled VGSEEEEEEEEEE peptide at 16.7 μM in the presence of ATP (middle panel), spastin Walker B mutant at 10 μM (1.67 μM for the hexamer) and Atto-labeled peptide at 16.7 μM in the presence of ATP (lower panel).
Supplementary Figure 2 Structural comparison of the final reconstructions of the spastin hexamer with poly-Glu substrate and the 3.8 Å resolution map of the spastin hexamer obtained without the addition of substrate.
(a) EM density of the earlier reconstruction (yellow) adjacent to our final reconstruction with poly-Glu added (grey). The sample preparation for this structure did not include incubation in the presence of substrate. Despite this, an unknown density was found docked within the pore of the hexamer. Cryo-EM data pertaining to this map was processed in a similar fashion to our 3.2 Å map, except using Relion 2.1 instead of cryoSPARC for the final 3D reconstruction. Incubation of our sample with poly-Glu increased the number of intact hexameric particles used in the reconstruction and allowed us to acquire a map at higher resolution. (b) Comparison of raw micrograph images selected from both datasets. (c) Comparison of 2D classes selected from both datasets. (d) Comparison of substrate density for the final reconstruction (grey, poly-Glu substrate is light green) and unknown substrate density (yellow, unknown density is purple). (e) Euler distribution plot for the final reconstruction with the poly-Glu substrate. More populated views are shown in red.
Supplementary Figure 3 Image processing of cryo-EM data and local and global resolution of the final cryo-EM map.
(a) Flowchart for image processing of cryo-EM data. (b) Local resolution estimation of the final cryo-EM map using BSOFT85. (c) Gold standard Fourier shell correlation versus spatial frequency plot of final map refinement in cryoSPARC74. Blue horizontal line depicts the gold standard FSC (GSFSC) value equal to 0.143. (d) Multi-model validation79: per-residue Cα RMSD values were calculated from the top ten refined atomic models and plotted on the histogram with the mean per-residue value denoted by a black vertical bar. Inset is the atomic model in worm representation colored by per-residue Cα RMSD, as in the histogram.
Supplementary Figure 4 EM map quality at selected secondary structural elements and EM map and atomic model of the nucleotide binding pocket of each protomer.
(a) Quality of the EM map at selected secondary structural elements. The entirety of protomer C is shown in ribbon representation and colored according to Fig. 1 with the corresponding EM density shown as a transparent surface. Selected structural elements are labelled and shown in stick representation with EM density depicted in grey mesh. (b) EM map and atomic model of the nucleotide binding pocket of each protomer. Sidechains and nucleotides are depicted in stick representation, while the backbone is in ribbon and the EM map is shown as grey mesh. Each chain is colored and labeled according to its respective protomer (see Fig. 1b). For the ATP in protomers A through E, clear distinctive density was observed consistent with a coordinating magnesium ion. The nucleotide density for protomer F is less well resolved and may contain a mixture of ADP and ATP states.
Supplementary Figure 5 Structural alignment of spastin protomers within the hexamer and comparison to apo spastin monomer.
(a) Structural alignment of each protomer using Chimera’s Match Maker86 with a Cα-RMSD of 0.685 angstroms. (b) Superposition of the apo spastin monomer (light grey; PDB: 3B9P6) and the ATP-bound protomer C from our hexameric spastin structure (dark grey) with pore loops 1, 2 and 3 highlighted in blue, yellow and magenta, respectively. The pore loops in the apo structure are shown in lighter hues of the same colors. The NBD undergoes large structural rearrangements upon ATP and peptide substrate binding, resulting in repositioning of pore loop 1, a disorder-to-order transition for pore loop 2 and ~ 1 Å movement of pore loop 3. Additionally, the HBD of the ATP and substrate-bound protomer undergoes a rotation of 9 degrees away from the NBD relative to the apo protomer.
Supplementary Figure 6 Oligomerization interactions between spastin protomers.
(a) The linker (purple) and helix α1 (blue) participate in oligomerization interactions. (b) The C-terminus of spastin is stabilized in the oligomer and together with helix α11 (orange) forms a belt around the hexamer. (c) Invariant Y753 is part of the oligomerization interface with the α10-α11 loop. Protomer D is depicted in blue, protomer E in green. The cryo-EM density is shown as a semi-transparent surface. (d) R601 is within H-bonding distance to the polyglutamate substrate and S599 of the adjacent lower protomer, likely coupling oligomerization to substrate engagement. Protomers colored as in Fig. 1.
Supplementary Figure 7 Modeled substrate within the cryo-EM density.
(a). Poly-glutamate substrate shown in two opposing orientations with the EM density shown as a mesh. (b) Stereo view of the poly-glutamate fit into the EM density.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7 and Table 1
Rights and permissions
About this article
Cite this article
Sandate, C.R., Szyk, A., Zehr, E.A. et al. An allosteric network in spastin couples multiple activities required for microtubule severing. Nat Struct Mol Biol 26, 671–678 (2019). https://doi.org/10.1038/s41594-019-0257-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0257-3
This article is cited by
-
Structure of the human ATAD2 AAA+ histone chaperone reveals mechanism of regulation and inter-subunit communication
Communications Biology (2023)
-
Regulators of tubulin polyglutamylation control nuclear shape and cilium disassembly by balancing microtubule and actin assembly
Cell Research (2022)
-
Active conformation of the p97-p47 unfoldase complex
Nature Communications (2022)
-
The force required to remove tubulin from the microtubule lattice by pulling on its α-tubulin C-terminal tail
Nature Communications (2022)
-
AAA+ protease-adaptor structures reveal altered conformations and ring specialization
Nature Structural & Molecular Biology (2022)