Abstract
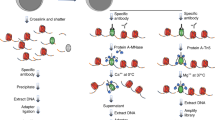
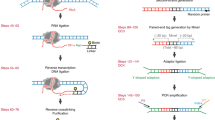
Cell identity is determined by the selective activation or silencing of specific genes via transcription factor binding and epigenetic modifications on the genome. Chromatin immunoprecipitation (ChIP) has been the standard technique for mapping the sites of transcription factor binding and histone modification. Recently, alternative methods to ChIP have been developed for addressing the increasing demands for low-input epigenomic profiling. Chromatin integration labeling (ChIL) followed by sequencing (ChIL-seq) has been demonstrated to be particularly useful for epigenomic profiling of low-input samples or even single cells because the technique amplifies the target genomic sequence before cell lysis. After labeling the target protein or modification in situ with an oligonucleotide-conjugated antibody (ChIL probe), the nearby genome sequence is amplified by Tn5 transposase-mediated transposition followed by T7 RNA polymerase-mediated transcription. ChIL-seq enables the detection of the antibody target localization under a fluorescence microscope and at the genomic level. Here we describe the detailed protocol of ChIL-seq with assessment methods for the key steps, including ChIL probe reaction, transposition, in situ transcription and sequencing library preparation. The protocol usually takes 3 d to prepare the sequencing library, including overnight incubations for the ChIL probe reaction and in situ transcription. The ChIL probe can be separately prepared and stored for several months, and its preparation and evaluation protocols are also documented in detail. An optional analysis for multiple targets (multitarget ChIL-seq) is also described. We anticipate that the protocol presented here will make the ChIL technique more widely accessible for analyzing precious samples and facilitate further applications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
ChIL-seq and mtChIL-seq data in this study have been deposited in the Gene Expression Omnibus (GEO) under the accession code GSE140659. Source data are provided with this paper.
Code availability
All performance metrics of low-input epigenomic profiling methods in Table 2 are the re-evaluation of our original ChIL-seq data analysis (except for the adapted information from previous reports, specified in the table legend). The main workhorse is the tableTPFP function in RScripts/myROC.R in https://github.com/kazumits/DataAnalysisForChILT. The use case of calculating ROC at TSS is according to the example described at https://github.com/kazumits/DataAnalysisForChILT/blob/master/ChILT_optimalResolution.md. The evaluation of genome-wide prediction performance (for example, precision, recall) against gold-standard ChIP-seq peaks is shown at https://github.com/kazumits/DataAnalysisForChILT/blob/master/ChILT_sensitivity-specificity.md. Source data are provided with this paper.
Change history
29 November 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41596-023-00940-6
References
Bannister, A. J. & Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 21, 381–395 (2011).
Kimura, H. Histone modifications for human epigenome analysis. J. Hum. Genet. 58, 439–445 (2013).
Harada, A. et al. A chromatin integration labelling method enables epigenomic profiling with lower input. Nat. Cell Biol. 21, 287–296 (2019).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Carter, B. et al. Mapping histone modifications in low cell number and single cells using antibody-guided chromatin tagmentation (ACT-seq). Nat. Commun. 10, 3747 (2019).
Wang, Q. et al. CoBATCH for high-throughput single-cell epigenomic profiling. Mol. Cell 76, 206–216.e207 (2019).
Skene, P. J., Henikoff, J. G. & Henikoff, S. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat. Protoc. 13, 1006–1019 (2018).
Ku, W. L. et al. Single-cell chromatin immunocleavage sequencing (scChIC-seq) to profile histone modification. Nat. Methods 16, 323–325 (2019).
Ai, S. et al. Profiling chromatin states using single-cell itChIP-seq. Nat. Cell Biol. 21, 1164–1172 (2019).
Zhu, B. et al. MOWChIP-seq for low-input and multiplexed profiling of genome-wide histone modifications. Nat. Protoc. 14, 3366–3394 (2019).
Lara-Astiaso, D. et al. Immunogenetics. Chromatin state dynamics during blood formation. Science 345, 943–949 (2014).
Dahl, J. A. et al. Broad histone H3K4me3 domains in mouse oocytes modulate maternal-to-zygotic transition. Nature 537, 548–552 (2016).
Brind’Amour, J. et al. An ultra-low-input native ChIP-seq protocol for genome-wide profiling of rare cell populations. Nat. Commun. 6, 6033 (2015).
Zhang, B. et al. Allelic reprogramming of the histone modification H3K4me3 in early mammalian development. Nature 537, 553–557 (2016).
van Galen, P. et al. A multiplexed system for quantitative comparisons of chromatin landscapes. Mol. Cell 61, 170–180 (2016).
Cao, Z., Chen, C., He, B., Tan, K. & Lu, C. A microfluidic device for epigenomic profiling using 100 cells. Nat. Methods 12, 959–962 (2015).
Hainer, S. J., Boskovic, A., McCannell, K. N., Rando, O. J. & Fazzio, T. G. Profiling of pluripotency factors in single cells and early embryos. Cell 177, 1319–1329.e11 (2019).
Rotem, A. et al. Single-cell ChIP-seq reveals cell subpopulations defined by chromatin state. Nat. Biotechnol. 33, 1165–1172 (2015).
Ma, S., Hsieh, Y. P., Ma, J. & Lu, C. Low-input and multiplexed microfluidic assay reveals epigenomic variation across cerebellum and prefrontal cortex. Sci. Adv. 4, eaar8187 (2018).
Renaud, G., Stenzel, U., Maricic, T., Wiebe, V. & Kelso, J. deML: robust demultiplexing of Illumina sequences using a likelihood-based approach. Bioinformatics 31, 770–772 (2015).
Pombo, A. et al. Regional specialization in human nuclei: visualization of discrete sites of transcription by RNA polymerase III. EMBO J. 18, 2241–2253 (1999).
Kimura, H., Hayashi-Takanaka, Y., Goto, Y., Takizawa, N. & Nozaki, N. The organization of histone H3 modifications as revealed by a panel of specific monoclonal antibodies. Cell Struct. Funct. 33, 61–73 (2008).
Hayashi-Takanaka, Y. et al. Tracking epigenetic histone modifications in single cells using Fab-based live endogenous modification labeling. Nucleic Acids Res. 39, 6475–6488 (2011).
Chandra, T. et al. Independence of repressive histone marks and chromatin compaction during senescent heterochromatic layer formation. Mol. Cell 47, 203–214 (2012).
Stasevich, T. J. et al. Regulation of RNA polymerase II activation by histone acetylation in single living cells. Nature 516, 272–275 (2014).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Sato, S. et al. Biochemical analysis of nucleosome targeting by Tn5 transposase. Open Biol. 9, 190116 (2019).
Acknowledgements
We are grateful to the staff of Ohkawa Lab. The computation was carried out using the computer resource offered under the category of Intensively Promoted Projects by Research Institute for Information Technology, Kyushu University. This work was in part supported by MEXT/JSPS KAKENHI (JP17K17719 to T.H.; JP18K19432, JP19H03211, JP19H05425 and JP20H05368 to A.H.; JP18H04904, JP19H04970, JP19H03158 and JP20H05393 to K.M.; JP18H05534 to H.Ku.; JP18H04802, JP18H05527, JP19H05244, JP17H03608, JP20H00456, JP20H04846 and JP20K21398 to Y.O.; and JP18H05527 and JP17H01417 to H.Ki), JST CREST (JPMJCR16G1 to Y.O., H.Ku. and H.Ki.), JST PRESTO JPMJPR19K7 to A.H, AMED JP20ek0109489h0001 to Y.O. and AMED BINDS (JP19am0101076 to H.Ku. and JP19am0101105 to H.Ki.).
Author information
Authors and Affiliations
Contributions
T.H., A.H., Y.O. and H. Ki. designed the experiments. T.H. and A.H. performed the ChIL experiments with deep-sequencing analyses. K.M. performed the bioinformatics and statistical analyses. S.S. and H.Ku. produced the in-house Tn5 transposase. M.N. and N.G. contributed to the assay for in situ transcription. T.H., A.H., K.M., Y.O. and H.Ki. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests except T.H., A.H., H. Ku., Y.O. and H.Ki, who are involved in a pending patent related to ChIL.
Additional information
Peer review information Nature Protocols thanks Chang Lu, Sinem K. Saka and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol:
Harada, A. et al. Nat. Cell Biol. 21, 287−296 (2019): https://doi.org/10.1038/s41556-018-0248-3
Supplementary information
Supplementary Information
Supplementary Tables 1–5 and Figs. 1–9.
Source data
Source Data Fig. 2
Unprocessed gels.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Handa, T., Harada, A., Maehara, K. et al. Chromatin integration labeling for mapping DNA-binding proteins and modifications with low input. Nat Protoc 15, 3334–3360 (2020). https://doi.org/10.1038/s41596-020-0375-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-0375-8
This article is cited by
-
Combinatorial single-cell profiling of major chromatin types with MAbID
Nature Methods (2024)
-
Multifactorial epigenomic profiling of six chromatin states in single cells
Nature Methods (2024)
-
Chromatin accessibility profiling by ATAC-seq
Nature Protocols (2022)
-
Concurrent mapping of multiple epigenetic marks and co-occupancy using ACT2-seq
Cell & Bioscience (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.