Abstract
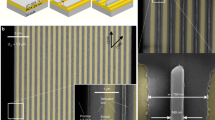
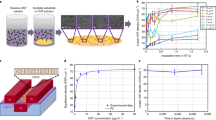
Structural DNA nanotechnology can be programmed into complex designer structures with molecular precision for directing a wide range of inorganic and biological materials. However, the use of DNA-templated approaches for the fabrication and performance requirements of ultra-scaled semiconductor electronics is limited by its assembly disorder and destructive interface composition. In this protocol, using carbon nanotubes (CNTs) as model semiconductors, we provide a stepwise process to build ultra-scaled, high-performance field-effect transistors (FETs) from micron-scale three-dimensional DNA templates. We apply the approach to assemble CNT arrays with uniform pitches scaled between 24.1 and 10.4 nm with yields of more than 95%, which exceeds the resolution limits of conventional lithography. To achieve highly clean CNT interfaces, we detail a rinsing-after-fixing step to remove residual DNA template and salt contaminations present around the contact and the channel regions, without modifying the alignment of the CNT arrays. The DNA-templated CNT FETs display both high on-state current (4–15 μA per CNT) and small subthreshold swing (60–100 mV per decade), which are superior to previous examples of biotemplated electronics and match the performance metrics of high-performance, silicon-based electronics. The scalable assembly of defect-free three-dimensional DNA templates requires 1 week and the CNT arrays can be synthesized within half a day. The interface engineering requires 1–2 d, while the fabrication of high-performance FET and logic gate circuits requires 2–4 d. The structural and performance characterizations of molecular-precise DNA self-assembly and high-performance electronics requires 1–2 d. The protocol is suited for users with expertise in DNA nanotechnology and semiconductor electronics.
Key points
-
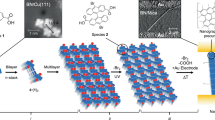
One-pot multistage isothermal annealing assembles DNA bricks into templates to which single-stranded DNAs/carbon nanotubes are hybridized. Source and drain electrodes and Y2O3 and HfO2 for the gate dielectrics of the field-effect transistors are layered via e-beam lithography, e-beam evaporation and atomic layer deposition.
-
Nanometer-sized DNA features fall below the resolution of conventional lithography. The resulting field-effect transistor channel pitches are smaller than typical silicon-based electronics, yet display high on-state current and small subthreshold swing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
References
Seeman, N. C. DNA in a material world. Nature 421, 427–431 (2003).
Seeman, N. C. Nucleic acid junctions and lattices. J. Theor. Biol. 99, 237–247 (1982).
Dey, S. et al. DNA origami. Nat. Rev. Methods Prim. 1, 13 (2021).
Dietz, H., Douglas, S. M. & Shih, W. M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725–730 (2009).
Aldaye, F. A., Palmer, A. L. & Sleiman, H. F. Assembling materials with DNA as the guide. Science 321, 1795–1799 (2008).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Ke, Y., Ong, L. L., Shih, W. M. & Yin, P. Three-dimensional structures self-assembled from DNA bricks. Science 338, 1177–1183 (2012).
Ke, Y. et al. DNA brick crystals with prescribed depths. Nat. Chem. 6, 994–1002 (2014).
Yan, H., Park, S. H., Finkelstein, G., Reif, J. H. & LaBean, T. H. DNA-templated self-assembly of protein arrays and highly conductive nanowires. Science 301, 1882–1884 (2003).
Yin, P. et al. Programming DNA tube circumferences. Science 321, 824–826 (2008).
Wagenbauer, K. F., Sigl, C. & Dietz, H. Gigadalton-scale shape-programmable DNA assemblies. Nature 552, 78–83 (2017).
Andersen, E. S. et al. Self-assembly of a nanoscale DNA box with a controllable lid. Nature 459, 73–76 (2009).
Ong, L. L. et al. Programmable self-assembly of three-dimensional nanostructures from 10,000 unique components. Nature 552, 72–77 (2017).
Seeman, N. C. & Sleiman, H. F. DNA nanotechnology. Nat. Rev. Mater. 3, 17068 (2017).
Dai, M., Jungmann, R. & Yin, P. Optical imaging of individual biomolecules in densely packed clusters. Nat. Nanotechnol. 11, 798–807 (2016).
Chen, Y. et al. A synthetic light-driven substrate channeling system for precise regulation of enzyme cascade activity based on DNA origami. J. Am. Chem. Soc. 140, 8990–8996 (2018).
Veneziano, R. et al. Role of nanoscale antigen organization on B-cell activation probed using DNA origami. Nat. Nanotechnol. 15, 716–723 (2020).
Sigl, C. et al. Programmable icosahedral shell system for virus trapping. Nat. Mater. 20, 1281–1289 (2021).
Li, S. et al. A DNA nanorobot functions as a cancer therapeutic in response to a molecular trigger in vivo. Nat. Biotechnol. 36, 258–264 (2018).
Liu, S. et al. A DNA nanodevice-based vaccine for cancer immunotherapy. Nat. Mater. 20, 421–430 (2021).
Zhao, S. et al. A DNA origami-based aptamer nanoarray for potent and reversible anticoagulation in hemodialysis. Nat. Commun. 12, 358 (2021).
Sun, W. et al. Casting inorganic structures with DNA molds. Science 346, 1258361 (2014).
Jing, X. et al. Solidifying framework nucleic acids with silica. Nat. Protoc. 14, 2416–2436 (2019).
Kuzyk, A. et al. DNA-based self-assembly of chiral plasmonic nanostructures with tailored optical response. Nature 483, 311–314 (2012).
Shen, J., Sun, W., Liu, D., Schaus, T. & Yin, P. Three-dimensional nanolithography guided by DNA modular epitaxy. Nat. Mater. 20, 683–690 (2021).
Kong, G. et al. DNA origami-based protein networks: from basic construction to emerging applications. Chem. Soc. Rev. 50, 1846–1873 (2021).
Knudsen, J. B. et al. Routing of individual polymers in designed patterns. Nat. Nanotechnol. 10, 892–898 (2015).
Zhang, P. et al. Capturing transient antibody conformations with DNA origami epitopes. Nat. Commun. 11, 3114 (2020).
Shaw, A. et al. Binding to nanopatterned antigens is dominated by the spatial tolerance of antibodies. Nat. Nanotechnol. 14, 184–190 (2019).
Lee, H. et al. Molecularly self-assembled nucleic acid nanoparticles for targeted in vivo siRNA delivery. Nat. Nanotechnol. 7, 389–393 (2012).
M. van den Brink. Continued scaling in semiconductor manufacturing enabled by advances in lithography. In 2019 IEEE International Electron Devices Meeting (IEDM). 1.2.1–1.2.5 (IEEE, 2019).
Wagner, C. & Harned, N. Lithography gets extreme. Nat. Photonics 4, 24–26 (2010).
The International Technology Roadmap for Semiconductors 2.0: 2015 Edition. (Semiconductor Industry Association, 2015).
Pei, H. et al. Organizing end-site-specific SWCNTs in specific loci using DNA. J. Am. Chem. Soc. 141, 11923–11928 (2019).
Maune, H. T. et al. Self-assembly of carbon nanotubes into two-dimensional geometries using DNA origami templates. Nat. Nanotechnol. 5, 61–66 (2010).
Zhang, Y. et al. Nanoparticle-assisted alignment of carbon nanotubes on DNA origami. Angew. Chem. Int. Ed. 59, 4892–4896 (2020).
Mangalum, A., Rahman, M. & Norton, M. L. Site-specific immobilization of single-walled carbon nanotubes onto single and one-dimensional DNA origami. J. Am. Chem. Soc. 135, 2451–2454 (2013).
Zhao, M. et al. DNA-directed nanofabrication of high-performance carbon nanotube field-effect transistors. Science 368, 878–881 (2020).
Sun, W. et al. Precise pitch-scaling of carbon nanotube arrays within three-dimensional DNA nanotrenches. Science 368, 874–877 (2020).
Chen, Y., Yang, C., Zhu, Z. & Sun, W. Suppressing high-dimensional crystallographic defects for ultra-scaled DNA arrays. Nat. Commun. 13, 2707 (2022).
Liu, L. et al. Aligned, high-density semiconducting carbon nanotube arrays for high-performance electronics. Science 368, 850–856 (2020).
Qiu, C. et al. Scaling carbon nanotube complementary transistors to 5-nm gate lengths. Science 355, 271–276 (2017).
Tulevski, G. S. et al. Toward high-performance digital logic technology with carbon nanotubes. ACS Nano 8, 8730–8745 (2014).
Chau, R. et al. Benchmarking nanotechnology for high-performance and low-power logic transistor applications. IEEE Trans. Nanotechnol. 4, 153–158 (2005).
Peng, L.-M., Zhang, Z. & Qiu, C. Carbon nanotube digital electronics. Nat. Electron. 2, 499–505 (2019).
Cao, Q. et al. Arrays of single-walled carbon nanotubes with full surface coverage for high-performance electronics. Nat. Nanotechnol. 8, 180–186 (2013).
He, X. et al. Wafer-scale monodomain films of spontaneously aligned single-walled carbon nanotubes. Nat. Nanotechnol. 11, 633–638 (2016).
Yang, Y., Ding, L., Han, J., Zhang, Z. & Peng, L.-M. High-performance complementary transistors and medium-scale integrated circuits based on carbon nanotube thin films. ACS Nano 11, 4124–4132 (2017).
Brady, G. J. et al. Quasi-ballistic carbon nanotube array transistors with current density exceeding Si and GaAs. Sci. Adv. 2, e1601240 (2016).
Cao, Q., Han, S. & Tulevski, G. S. Fringing-field dielectrophoretic assembly of ultrahigh-density semiconducting nanotube arrays with a self-limited pitch. Nat. Commun. 5, 5071 (2014).
Park, H. et al. High-density integration of carbon nanotubes via chemical self-assembly. Nat. Nanotechnol. 7, 787–791 (2012).
Han, S.-J. et al. High-speed logic integrated circuits with solution-processed self-assembled carbon nanotubes. Nat. Nanotechnol. 12, 861–865 (2017).
Shulaker, M. M. et al. Three-dimensional integration of nanotechnologies for computing and data storage on a single chip. Nature 547, 74–78 (2017).
Jin, S. H. et al. Using nanoscale thermocapillary flows to create arrays of purely semiconducting single-walled carbon nanotubes. Nat. Nanotechnol. 8, 347–355 (2013).
Guo, Y. et al. Soft-lock drawing of super-aligned carbon nanotube bundles for nanometre electrical contacts. Nat. Nanotechnol. 17, 278–284 (2022).
Benson, E. et al. DNA rendering of polyhedral meshes at the nanoscale. Nature 523, 441–444 (2015).
Qiu, S. et al. Solution-processing of high-purity semiconducting single-walled carbon nanotubes for electronics devices. Adv. Mater. 31, 1800750 (2019).
Tu, X., Manohar, S., Jagota, A. & Zheng, M. DNA sequence motifs for structure-specific recognition and separation of carbon nanotubes. Nature 460, 250–253 (2009).
Zheng, M. et al. Structure-based carbon nanotube sorting by sequence-dependent DNA assembly. Science 302, 1545–1548 (2003).
Liu, H., Nishide, D., Tanaka, T. & Kataura, H. Large-scale single-chirality separation of single-wall carbon nanotubes by simple gel chromatography. Nat. Commun. 2, 309 (2011).
Ma, Z., Han, J., Yao, S., Wang, S. & Peng, L.-M. Improving the performance and uniformity of carbon-nanotube-network-based photodiodes via yttrium oxide coating and decoating. ACS Appl. Mater. Interfaces 11, 11736–11742 (2019).
Cao, Q., Tersoff, J., Farmer, D. B., Zhu, Y. & Han, S.-J. Carbon nanotube transistors scaled to a 40-nanometer footprint. Science 356, 1369–1372 (2017).
Shulaker, M. M. et al. High-performance carbon nanotube field-effect transistors. In 2014 IEEE International Electron Devices Meeting (IEDM) 33.6.1–33.6.4 (IEEE, 2014).
2018 Semiconductor synthetic biology roadmap. Semicond. Res. Corp. 36 (2018).
Bui, H. et al. Programmable periodicity of quantum dot arrays with DNA origami nanotubes. Nano Lett. 10, 3367–3372 (2010).
Tikhomirov, G., Petersen, P. & Qian, L. Fractal assembly of micrometre-scale DNA origami arrays with arbitrary patterns. Nature 552, 67–71 (2017).
Woods, D. et al. Diverse and robust molecular algorithms using reprogrammable DNA self-assembly. Nature 567, 366–372 (2019).
Woo, S. & Rothemund, P. W. K. Self-assembly of two-dimensional DNA origami lattices using cation-controlled surface diffusion. Nat. Commun. 5, 4889 (2014).
Han, S.-P., Maune, H. T., Barish, R. D., Bockrath, M. & Goddard, W. A. DNA-linker-induced surface assembly of ultra dense parallel single walled carbon nanotube arrays. Nano Lett. 12, 1129–1135 (2012).
Zhong, D. et al. Gigahertz integrated circuits based on carbon nanotube films. Nat. Electron. 1, 40–45 (2018).
Lei, T. et al. Low-voltage high-performance flexible digital and analog circuits based on ultrahigh-purity semiconducting carbon nanotubes. Nat. Commun. 10, 2161 (2019).
Fu, Y. et al. Single-step rapid assembly of DNA origami nanostructures for addressable nanoscale bioreactors. J. Am. Chem. Soc. 135, 696–702 (2013).
Sobczak, J.-P. J., Martin, T. G., Gerling, T. & Dietz, H. Rapid folding of DNA into nanoscale shapes at constant temperature. Science 338, 1458–1461 (2012).
Praetorius, F. et al. Biotechnological mass production of DNA origami. Nature 552, 84–87 (2017).
Wang, Y. et al. What molecular assembly can learn from catalytic chemistry. Chem. Soc. Rev. 43, 399–411 (2014).
Zhang, H. et al. Programming chain-growth copolymerization of DNA hairpin tiles for in-vitro hierarchical supramolecular organization. Nat. Commun. 10, 1006 (2019).
Yao, G. et al. Meta-DNA structures. Nat. Chem. 12, 1067–1075 (2020).
Zhang, Z. et al. Doping-free fabrication of carbon nanotube based ballistic CMOS devices and circuits. Nano Lett. 7, 3603–3607 (2007).
Lin, Z., Yang, Y., Jagota, A. & Zheng, M. Machine learning-guided systematic search of DNA sequences for sorting carbon nanotubes. ACS Nano 16, 4705–4713 (2022).
Lin, Z. et al. DNA-guided lattice remodeling of carbon nanotubes. Science 377, 535–539 (2022).
Tang, J. et al. Flexible CMOS integrated circuits based on carbon nanotubes with sub-10 ns stage delays. Nat. Electron. 1, 191–196 (2018).
Arnold, M. S., Green, A. A., Hulvat, J. F., Stupp, S. I. & Hersam, M. C. Sorting carbon nanotubes by electronic structure using density differentiation. Nat. Nanotechnol. 1, 60–65 (2006).
Arnold, M. S., Stupp, S. I. & Hersam, M. C. Enrichment of single-walled carbon nanotubes by diameter in density gradients. Nano Lett. 5, 713–718 (2005).
Tulevski, G. S., Franklin, A. D. & Afzali, A. High purity isolation and quantification of semiconducting carbon nanotubes via column chromatography. ACS Nano 7, 2971–2976 (2013).
M. M. Shulaker et al. Efficient metallic carbon nanotube removal for highly-scaled technologies. In 2015 IEEE International Electron Devices Meeting (IEDM). 32.4.1–32.4.4 (IEEE, 2015)
Kang, S. J. et al. High-performance electronics using dense, perfectly aligned arrays of single-walled carbon nanotubes. Nat. Nanotechnol. 2, 230–2326 (2007).
Acknowledgements
Y.C., M.Z., Y.O., Z.L., K.W., Z. Zhang and W.S. acknowledge the National Key R&D Program of China (2021YFF1200300, 2020YFA0714703), the National Science Foundation of China (grant nos. 21875003, 21991134 and T2125001), Beijing Municipal Science and Technology Commission (grant no. JQ20007), Peking University, and Zhangjiang Laboratory for financial support. Y.C., S.Z., C.Y. and Z. Zhu acknowledge the National Natural Science Foundation of China (grant nos. 21974113, 21735004 and 21521004), National Key R&D Program of China (2021YFA0909400) and the Fundamental Research Funds for the Central Universities (20720210001). Y.L. acknowledges City University of Hong Kong for financial support. J.S. acknowledges the National Key R&D Program of China (2022YFB3503700) and the National Science Foundation of China (grant no. 22271003). We acknowledge the Peking Nanofab for the support of device fabrication.
Author information
Authors and Affiliations
Contributions
Y.C. and M.Z. developed the protocol, conducted the whole experiments, interpreted the data and wrote the manuscript. Y.O., S.Z., Z.L., K.W. and Z. Zhang contributed to the protocol development, conducted the experiments and analyzed the data. Y.L. and C.Y. analyzed the data and supervised the study. W.S. conceived, designed and supervised the study, interpreted the data and wrote the manuscript. J.S. designed and supervised the study, and interpreted the data. Z. Zhu designed, supervised the study, interpreted the data and wrote the manuscript. All the authors edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Mihai Irimia and Haitao Liu for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Ke, Y. et al. Nat. Chem. 6, 994–1002 (2014): https://doi.org/10.1038/nchem.2083
Sun, W. et al. Science 368, 874–877 (2020): https://doi.org/10.1126/science.aaz7440
Zhao, M. et al. Science 368, 878–881 (2020): https://doi.org/10.1126/science.aaz7435
Shen, J. et al. Nat. Mater. 20, 683–690 (2021): https://doi.org/10.1038/s41563-021-00930-7
Chen, Y. et al. Nat. Commun. 13, 2707 (2022): https://doi.org/10.1038/s41467-022-30441-1
Supplementary information
Supplementary Information
Supplementary Figs. 1–11.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, Y., Zhao, M., Ouyang, Y. et al. Biotemplated precise assembly approach toward ultra-scaled high-performance electronics. Nat Protoc 18, 2975–2997 (2023). https://doi.org/10.1038/s41596-023-00870-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00870-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.