Abstract
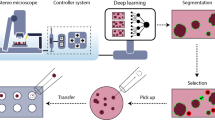
To produce abundant cell culture samples to generate large, standardized image datasets of human induced pluripotent stem (hiPS) cells, we developed an automated workflow on a Hamilton STAR liquid handler system. This was developed specifically for culturing hiPS cell lines expressing fluorescently tagged proteins, which we have used to study the principles by which cells establish and maintain robust dynamic localization of cellular structures. This protocol includes all details for the maintenance, passage and seeding of cells, as well as Matrigel coating of 6-well plastic plates and 96-well optical-grade, glass plates. We also developed an automated image-based hiPS cell colony segmentation and feature extraction pipeline to streamline the process of predicting cell count and selecting wells with consistent morphology for high-resolution three-dimensional (3D) microscopy. The imaging samples produced with this protocol have been used to study the integrated intracellular organization and cell-to-cell variability of hiPS cells to train and develop deep learning-based label-free predictions from transmitted-light microscopy images and to develop deep learning-based generative models of single-cell organization. This protocol requires some experience with robotic equipment. However, we provide details and source code to facilitate implementation by biologists less experienced with robotics. The protocol is completed in less than 10 h with minimal human interaction. Overall, automation of our cell culture procedures increased our imaging samples’ standardization, reproducibility, scalability and consistency. It also reduced the need for stringent culturist training and eliminated culturist-to-culturist variability, both of which were previous pain points of our original manual pipeline workflow.
Key points
-
This protocol describes an automated workflow for the high-throughput culture of human induced pluripotent stem cells expressing fluorescently tagged proteins, and their seeding on 96-well optical-grade, glass-bottom plates for high-quality, live-cell three-dimensional microscopy on a large scale.
-
This produces large, standardized image datasets that we have used to study integrated intracellular organization and cell-to-cell variability, and to generate deep learning-based models of three-dimensional single-cell organization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Source data are provided in this paper. Source data are provided with this paper.
Code availability
Workflow scripts are available from GitHub: https://github.com/AllenInstitute/aics-automated-cell-culture-workflow. Calibration, priming and initialization information can be found in the Supplementary Information file. The entire methods is also available as a python package ‘PyHamilton’ (github.com/dgretton/pyhamilton) available at https://github.com/stefangolas/ipsc-robot.
References
Viana, M. P. et al. Integrated intracellular organization and its variations in human iPS cells. Nature 613, 345–354 (2023).
Ounkomol, C., Seshamani, S., Maleckar, M. M., Collman, F. & Johnson, G. R. Label-free prediction of three-dimensional fluorescence images from transmitted-light microscopy. Nat. Methods 15, 917–920 (2018).
Donovan-Maiye, R. M. et al. A deep generative model of 3D single-cell organization. PLoS Comput. Biol. 18, e1009155 (2022).
Mahla, R. S. Stem cells applications in regenerative medicine and disease therapeutics. Int. J. Cell Biol. 2016, 6940283 (2016).
Roberts, B. et al. Systematic gene tagging using CRISPR/Cas9 in human stem cells to illuminate cell organization. Mol. Biol. Cell 28, 2854–2874 (2017).
Paull, D. et al. Automated, high-throughput derivation, characterization and differentiation of induced pluripotent stem cells. Nat. Methods 12, 885–892 (2015).
Baghbaderani, A. A. et al. cGMP-manufactured human induced pluripotent stem cells are available for pre-clinical and clinical applications. Stem Cell Rep. 5, 647–659 (2015).
Archibald, P. R. T. et al. Comparability of automated human induced pluripotent stem cell culture: a pilot study. Bioprocess Biosyst. Eng. 39, 1847–1858 (2016).
Crombie, D. E. et al. Development of a modular automated system for maintenance and differentiation of adherent human pluripotent stem cells. SLAS Discov. 22, 1016–1025 (2017).
Koike, H. et al. Establishment of automated culture system for murine induced pluripotent stem cells. BMC Biotechnol. 12, 81 (2012).
Kami, D. et al. Large-scale cell production of stem cells for clinical application using the automated cell processing machine. BMC Biotechnol. 13, 102 (2013).
Terstegge, S. et al. Automated maintenance of embryonic stem cell cultures. Biotechnol. Bioeng. 96, 195–201 (2007).
Thomas, R. J. et al. Automated, scalable culture of human embryonic stem cells in feeder-free conditions. Biotechnol. Bioeng. 102, 1636–1644 (2009).
Daniszewski, M. et al. Automated cell culture systems and their applications to human pluripotent stem cell studies. SLAS Technol. 23, 315–325 (2018).
Wang, Y., Cheng, L. & Gerecht, S. Efficient and scalable expansion of human pluripotent stem cells under clinically compliant settings: a view in 2013. Ann. Biomed. Eng. 42, 1357–1372 (2014).
Hussain, W. et al. Reproducible culture and differentiation of mouse embryonic stem cells using an automated microwell platform. Biochem. Eng. J. 77, 246–257 (2013).
Konagaya, S., Ando, T., Yamauchi, T., Suemori, H. & Iwata, H. Long-term maintenance of human induced pluripotent stem cells by automated cell culture system. Sci. Rep. 5, 16647 (2015).
Webb, D. J. & Brown, C. M. Epi-fluorescence microscopy. Methods Mol. Biol. 931, 29–59 (2013).
Chatterjee, I. et al. in Induced Pluripotent Stem (iPS) Cells. Vol. 1357 (eds. Turksen, K. & Nagy, A.) 311–327 (Springer, 2015).
Le, M. N. T. & Hasegawa, K. Expansion culture of human pluripotent stem cells and production of cardiomyocytes. Bioengineering 6, 48 (2019).
Conway, M. K. et al. Scalable 96-well plate based iPSC culture and production using a robotic liquid handling system. J. Vis. Exp. https://doi.org/10.3791/52755 (2015).
Chan, S. W., Rizwan, M. & Yim, E. K. F. Emerging methods for enhancing pluripotent stem cell expansion. Front. Cell Dev. Biol. 8, 70 (2020).
Haenel, F. & Garbow, N. Application note: Cell counting and confluency analysis as quality controls in cell-based assays. PerkinElmer https://resources.perkinelmer.com/lab-solutions/resources/docs/011833_01_app.pdf (2014).
Jaccard, N. et al. Automated method for the rapid and precise estimation of adherent cell culture characteristics from phase contrast microscopy images. Biotechnol. Bioeng. 111, 504–517 (2014).
Kreitzer, F. R. et al. A robust method to derive functional neural crest cells from human pluripotent stem cells. Am. J. Stem Cells 2, 119–131 (2013).
Schulz, T. C. et al. Directed neuronal differentiation of human embryonic stem cells. BMC Neurosci. 4, 27 (2003).
Zemirli, N., Morel, E. & Molino, D. Mitochondrial dynamics in basal and stressful conditions. Int. J. Mol. Sci. 19, 564 (2018).
Chen, J. et al. The Allen Cell Structure Segmenter: a new open source toolkit for segmenting 3D intracellular structures in fluorescence microscopy images. Preprint at bioRxiv https://doi.org/10.1101/491035 (2018).
Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232 (2019).
van der Walt, S. et al. scikit-image: image processing in Python. PeerJ 2, e453 (2014).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Kamentsky, L. et al. Improved structure, function and compatibility for CellProfiler: modular high-throughput image analysis software. Bioinformatics 27, 1179–1180 (2011).
McQuin, C. et al. CellProfiler 3.0: next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Acknowledgements
We thank J. Burn for editing the video tutorial, A. R. Horwitz and the Allen Institute for Cell Science Team for helpful discussions. We also thank S. Golas from the Massachusetts Institute of Technology Media Lab for converting the Hamilton methods to pyHamilton code. The WTC line that we used to create our gene-edited cell lines was provided by the Bruce R. Conklin Laboratory at the Gladstone Institute and University of California, San Francisco. We acknowledge the Allen Institute for Cell Science founder, P. G. Allen, for his vision, encouragement and support.
Author information
Authors and Affiliations
Contributions
Conceptualization: S.M.R., W.W., R.N.G. and N.G. Methodology: M.E.C., B.W.G., J.A., M.A.F., A.H., M.C.H., I.A.M., A.M.N., D.J.T., C.Y., R.N.G. and N.G. Software: M.J.S.-B., A.N. and B.P.W. Validation: M.E.C., B.W.G., J.A., A.B., M.A.F., A.H., M.C.H., W.L., I.A.M., A.M.N., W.J.T., D.J.T., C.Y. and N.G. Formal analysis: M.E.C., A.B., D.J.T. and C.Y. Investigation: M.E.C., B.W.G., M.C.H., W.L. and W.J.T. Resources: W.W. Data curation: M.E.C., A.B., M.A.F., A.H., M.C.H., W.L., I.A.M., A.M.N., W.J.T. and N.G. Writing — original draft: M.E.C., B.W.G., A.B., D.J.T., C.Y. and N.G. Writing — review and editing: M.E.C., B.W.G., A.B., E.M.A., M.A.F., I.A.M., A.M.N., A.N., S.M.R., E.E.S., D.J.T., C.Y., R.N.G. and N.G. Visualization: M.E.C., B.W.G., A.B., T.P.D., D.J.T., C.Y. and N.G. Supervision: S.M.R., W.W., R.N.G. and N.G. Project administration: I.A.M. and N.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Virgile Viasnoff and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Viana, M. P. et al. Nature 613, 345–354 (2023): https://doi.org/10.1038/s41586-022-05563-7
Donovan-Maiye, R. M. et al. PLoS Comput. Biol. 18, e1009155 (2022): https://doi.org/10.1371/journal.pcbi.1009155
Supplementary information
Supplementary Information
Supplementary Procedures and Fig. 1
Source data
Source Data Fig. 5, 6, 7 and 8
Statistical data and a link to a Quilt repository for the source image data.
Rights and permissions
About this article
Cite this article
Gregor, B.W., Coston, M.E., Adams, E.M. et al. Automated human induced pluripotent stem cell culture and sample preparation for 3D live-cell microscopy. Nat Protoc 19, 565–594 (2024). https://doi.org/10.1038/s41596-023-00912-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00912-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.