Abstract
High-protein diets are commonly utilized for weight loss, yet they have been reported to raise cardiovascular risk. The mechanisms underlying this risk are unknown. Here, we show that dietary protein drives atherosclerosis and lesion complexity. Protein ingestion acutely elevates amino acid levels in blood and atherosclerotic plaques, stimulating macrophage mammalian target of rapamycin (mTOR) signalling. This is causal in plaque progression, because the effects of dietary protein are abrogated in macrophage-specific Raptor-null mice. Mechanistically, we find amino acids exacerbate macrophage apoptosis induced by atherogenic lipids, a process that involves mammalian target of rapamycin complex 1 (mTORC1)-dependent inhibition of mitochondrial autophagy (mitophagy), accumulation of dysfunctional mitochondria and mitochondrial apoptosis. Using macrophage-specific mTORC1- and autophagy-deficient mice, we confirm this amino acid–mTORC1–autophagy signalling axis in vivo. Our data provide insights into the deleterious impact of excessive protein ingestion on macrophages and atherosclerotic progression. Incorporation of these concepts in clinical studies is important to define the vascular effects of protein-based weight loss regimens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Change history
09 September 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Moore, K. J. & Tabas, I. Macrophages in the pathogenesis of atherosclerosis. Cell 145, 341–355 (2011).
Gardner, C. D. et al. Comparison of the Atkins, Zone, Ornish, and LEARN diets for change in weight and related risk factors among overweight premenopausal women: the A to Z Weight Loss Study: a randomized trial. JAMA 297, 969–977 (2007).
Halton, T. L. et al. Low-carbohydrate-diet score and the risk of coronary heart disease in women. N. Engl. J. Med. 355, 1991–2002 (2006).
Hu, F. B. et al. Dietary protein and risk of ischemic heart disease in women. Am. J. Clin. Nutr. 70, 221–227 (1999).
Lagiou, P. et al. Low carbohydrate-high protein diet and incidence of cardiovascular diseases in Swedish women: prospective cohort study. BMJ 344, e4026 (2012).
Debry, G. Dietary Proteins and Atherosclerosis (CRC Press, 2004).
Foo, S. Y. et al. Vascular effects of a low-carbohydrate high-protein diet. Proc. Natl Acad. Sci. USA 106, 15418–15423 (2009).
Wolfson, R. L. & Sabatini, D. M. The dawn of the age of amino acid sensors for the mTORC1 pathway. Cell Metab. 26, 301–309 (2017).
Ma, Y. et al. A dietary quality comparison of popular weight-loss plans. J. Am. Diet. Assoc. 107, 1786–1791 (2007).
Anderson, J. W., Konz, E. C. & Jenkins, D. J. Health advantages and disadvantages of weight-reducing diets: a computer analysis and critical review. J. Am. Coll. Nutr. 19, 578–590 (2000).
Razani, B. et al. Autophagy links inflammasomes to atherosclerotic progression. Cell Metab. 15, 534–544 (2012).
Sergin, I. et al. Inclusion bodies enriched for p62 and polyubiquitinated proteins in macrophages protect against atherosclerosis. Sci. Signal. 9, ra2 (2016).
Sancak, Y. et al. Ragulator-Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell 141, 290–303 (2010).
Zoncu, R. et al. mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H+-ATPase. Science 334, 678–683 (2011).
Sengupta, S., Peterson, T. R., Laplante, M., Oh, S. & Sabatini, D. M. mTORC1 controls fasting-induced ketogenesis and its modulation by ageing. Nature 468, 1100–1104 (2010).
Ai, D. et al. Disruption of mammalian target of rapamycin complex 1 in macrophages decreases chemokine gene expression and atherosclerosis. Circ. Res. 114, 1576–1584 (2014).
Li, N. et al. Mitochondrial complex I inhibitor rotenone induces apoptosis through enhancing mitochondrial reactive oxygen species production. J. Biol. Chem. 278, 8516–8525 (2003).
Emanuel, R. et al. Induction of lysosomal biogenesis in atherosclerotic macrophages can rescue lipid-induced lysosomal dysfunction and downstream sequelae. Arterioscler. Thromb. Vasc. Biol. 34, 1942–1952 (2014).
Liao, X. et al. Macrophage autophagy plays a protective role in advanced atherosclerosis. Cell Metab. 15, 545–553 (2012).
Ouimet, M. et al. Autophagy regulates cholesterol efflux from macrophage foam cells via lysosomal acid lipase. Cell Metab. 13, 655–667 (2011).
Sergin, I. et al. Exploiting macrophage autophagy-lysosomal biogenesis as a therapy for atherosclerosis. Nat. Commun. 8, 15750 (2017).
Evans, T. D., Sergin, I., Zhang, X. & Razani, B. Target acquired: selective autophagy in cardiometabolic disease. Sci. Signal. 10, eaag2298 (2017).
Sun, N. et al. Measuring in vivo mitophagy. Mol. Cell 60, 685–696 (2015).
Laplante, M. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Food and Agriculture Organization, Food Policy and Food Science Service, Nutrition Division. Amino-Acid Content of Foods and Biological Data on Proteins (FAO, 1970).
Castellano, B. M. et al. Lysosomal cholesterol activates mTORC1 via an SLC38A9–Niemann-Pick C1 signaling complex. Science 355, 1306–1311 (2017).
Bernstein, A. M. et al. Major dietary protein sources and risk of coronary heart disease in women. Circulation 122, 876–883 (2010).
Solon-Biet, S. M. et al. The ratio of macronutrients, not caloric intake, dictates cardiometabolic health, aging, and longevity in ad libitum-fed mice. Cell Metab. 19, 418–430 (2014).
Kurdi, A., De Meyer, G. R. & Martinet, W. Potential therapeutic effects of mTOR inhibition in atherosclerosis. Br. J. Clin. Pharmacol. 82, 1267–1279 (2016).
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889 (2006).
Razani, B. et al. Fatty acid synthase modulates homeostatic responses to myocardial stress. J. Biol. Chem. 286, 30949–30961 (2011).
Jewell, J. L. et al. Metabolism. Differential regulation of mTORC1 by leucine and glutamine. Science 347, 194–198 (2015).
Febbraio, M. et al. Targeted disruption of the class B scavenger receptor CD36 protects against atherosclerotic lesion development in mice. J. Clin. Invest. 105, 1049–1056 (2000).
Razani, B. et al. Caveolin-1 null mice are viable but show evidence of hyperproliferative and vascular abnormalities. J. Biol. Chem. 276, 38121–38138 (2001).
Katayama, H., Kogure, T., Mizushima, N., Yoshimori, T. & Miyawaki, A. A sensitive and quantitative technique for detecting autophagic events based on lysosomal delivery. Chem. Biol. 18, 1042–1052 (2011).
Sun, N. et al. A fluorescence-based imaging method to measure in vitro and in vivo mitophagy using mt-Keima. Nat. Protoc. 12, 1576–1587 (2017).
Acknowledgements
This work was supported by National Institutes of Health grant no. R01 HL125838, VA MERIT I01 BX003415, American Diabetes Association grant no. 1-18-IBS-029, Washington University Diabetic Cardiovascular Disease Center and Diabetes Research Center grant no. P30 DK020579, Washington University Mass Spectrometry core grant nos. P41GM103422 and P30DK056341, a grant from the Longer Life Foundation and the Foundation for Barnes-Jewish Hospital.
Author information
Authors and Affiliations
Contributions
X.Z. and B.R. designed the studies and wrote the manuscript. X.Z., I.S., T.D.E., S.J., A.R., D.K., S.C., E.S., K.B.H. and J.R.C. performed and analysed the experiments. X.Z. and I.S. prepared the figures. D.K., S.E., C.C.W., A.D., D.F., B.M., N.O.S., J.D.S., I.J.L. and B.R. provided the reagents, advised on the experimental design and performed critical reading of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Primary Handling Editor: Pooja Jha.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 High Protein diets increase atherosclerotic plaque formation and plaque complexity without altering serum metabolites.
(a) Average daily food intake over 1 week for ApoE-KO mice fed a standard Western diet (n = 5) or high protein Western diet (n = 5) (b–d) Cohorts of ApoE-KO mice were placed on a standard Western diet (Std. WD) or high protein Western diet (HP WD) and after 8 weeks, (b) Body composition (fat and lean weights) (Std. WD: n = 4; HP WD: n = 5), (c) glucose tolerance test (GTT) and glucose AUC (Std. WD: n = 7; HP WD: n = 9), and (d) serum cholesterol, glucose, triglycerides, and free fatty acids (Std. WD: n = 11; HP WD: n = 14) were measured. (e) Quantification of atherosclerotic plaque burden using Oil Red O-stained aortic root sections from mice fed standard or high protein Western diets for 16 weeks; representative roots shown on right (Std. WD: n = 12; HP WD: n = 11). (f) Measurements of serum cholesterol in cohorts of ApoE-KO mice after 16 weeks of standard or high protein Western diets (Std WD: n = 6; HP WD: n = 8). (g-i) Plaque composition quantified by immunofluorescence staining of aortic root sections for (g) macrophage (MOMA-2+) (Std. WD: n = 12; HP WD: n = 13), (h) apoptosis (TUNEL+) (Std. WD: n = 13; HP WD: n = 13), (i) and necrotic core (acellular) (Std. WD: n = 13; HP WD: n = 13). For all graphs, data are presented as mean ±SEM. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed unpaired t-test.
Extended Data Fig. 2 Levels of select L-amino acids in serum, aorta, and splenic macrophages after high protein challenge.
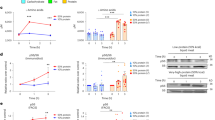
(a) Serum levels of 8 L-amino acids by mass spectrometry from mice fed standard or high protein Western diets for 8 weeks (Std. WD: n = 8; HP WD: n = 8). (b–d) Time course measurement of the levels of 8 L-amino acids in serum (n = 3) (b), splenic macrophages (n = 2) (c), and atherosclerotic aortas (n = 2) (d) by mass spectrometry after high protein gavage. For all graphs, data are presented as mean ±SEM. *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed unpaired t-test for a and one-way ANOVA with Dunnett’s test for b–d.
Extended Data Fig. 3 Leucine is amongst the best amino acid inducers of mTOR signaling in cultured primary macrophages.
(a, b) Immunoblot analysis of thioglycollate-elicited peritoneal macrophages (PMACs) (a) and bone marrow derived macrophages (BMDM) (b) after 30min of incubation with amino acid-free or regular/amino acid-sufficient medium to assess mTORC1 signaling using phospho- and total S6K and S6 levels as readouts. (c) Representative immunofluorescence images of BMDMs after 15min of incubation with amino acid-free or regular/amino acid-sufficient medium to assess co-localization between mTOR and Lamp2. (d, e) Immunoblot analysis of macrophages after 30min of incubation with regular medium or amino acid-free medium with and without 20 different L-amino acids to assess mTORC1 signaling using phospho- and total S6 levels as readouts. Representative blots (d) and quantification of pS6/total S6 ratio for five independent experiments (e). The best three mTOR inducers and three non-inducers are listed at right. (f, g) Immunofluorescence imaging of macrophages after 15min of incubation with regular medium or amino acid-free medium with and without the best three mTOR inducers and non-inducers to assess co-localization between mTOR and Lamp2. Representative images (f) and quantification of the mTOR/Lamp2 co-localization (+aa: n = 42; -aa: n = 74; Leu: n = 26; Arg: n = 38; Glu: n = 12; Gln: n = 34; Phe: n = 37; Thr: n = 43 cells) (g). For all graphs, data are presented as mean ±SEM.
Extended Data Fig. 4 Body weight and common serum metabolites of control and mϕRaptor KO mice fed a standard or LCHP Western diet.
(a) Immunoblot analysis of control and Raptor KO macrophages after 30min of incubation with regular medium or amino acid-free medium with and without leucine to assess mTORC1 activity using phospho- and total S6K and S6 as readouts. (b) Immunofluorescence imaging of macrophages after 15min of incubation with regular medium or amino acid-free medium with and without leucine to assess co-localization between mTOR and Lamp2. Representative images (left) and quantification of the mTOR/Lamp2 co-localization (right) (Control: +aa: n = 42; -aa: n = 52; Leu: n = 44; ATG5 KO: +aa: n = 27; -aa: n = 40; Leu: n = 43 cells). (c) Total body weights of control (n = 22) and mϕRaptor-KO mice (n = 18) (all on ApoE-KO background) fed a standard Western diet for 8 weeks. (d) Measurements of serum cholesterol, glucose, triglycerides, and free fatty acids in control and mϕRaptor-KO mice after 8 weeks of Western diet feeding (n = 15–25 per group). (e) Measurements of serum cholesterol, glucose, triglycerides, and free fatty acids in cohorts of mϕRaptor-KO mice (on ApoE-null background) fed a standard Western diet or high protein Western diet for 8 weeks (n = 15 per group). For all graphs, data are presented as mean ±SEM. *P < 0.05, **P < 0.01, one-way ANOVA with Tukey’s test for b, two-tailed unpaired t-test for c.
Extended Data Fig. 5 Leucine-mediated activation of mTORC1 synergizes with atherogenic lipids to induce mitochondrial-dependent apoptosis in macrophages.
(a, b) Control and Raptor-KO macrophages were treated with vehicle, 500µg/ml cholesterol crystals (CC) with or without leucine and apoptosis assessed by Caspase-3/7 immunofluorescence staining. Shown are (a) representative images and (b) quantification of >103 cells from acquired images (Control: -aa: n = 11; Leu: n = 9; cc: n = 11; cc+Leu: n = 11; Raptor-KO: n = 11 image fields). For all graphs, data are presented as mean ±SEM. *P < 0.05, ***P < 0.001, one-way ANOVA with Tukey’s test.
Extended Data Fig. 6 Leucine-mediated activation of mTORC1 inhibits autophagy in macrophages.
(a) Immunoblot analysis of phospho- and total ULK1 levels in macrophages incubated with amino acid-free medium with and without leucine for the indicated times. (b, c) The autophagy marker LC3 was evaluated in macrophages incubated with regular medium or amino acid-free medium with and without leucine for 30 minutes using (b) immunoblot analysis or (c) quantification of LC3 intensity (+aa: n = 52; -aa: n = 52; Leu: n = 52 cells) and number of puncta (+aa: n = 97; -aa: n = 87; Leu: n = 97 cells) by immunofluorescence microscopy. (d) Phospho- and total ULK1 levels were determined in Control and Raptor KO macrophages incubated with regular medium or amino acid-free medium with and without leucine for 30 minutes. (e) LC3 levels in control and Raptor KO macrophages incubated with vehicle or 100nM Bafilomycin for 2 hours. (f) LC3 intensity (Control: +aa: n = 39; -aa: n = 27; Leu: n = 30; Raptor-KO: +aa: n = 31; -aa: n = 30; Leu: n = 30 cells) and number of puncta (n = 52 cells/group) were analyzed by immunofluorescence microscopy in control and Raptor KO macrophages incubated with regular medium or amino acid-free medium with and without leucine for 30 minutes. (g) Quantification of LC3 intensity by immunofluorescence microscopy in control and ATG5-KO macrophages incubated with regular medium or amino acid-free medium with and without leucine for 30 minutes (Control: +aa: n = 52; -aa: n = 50; Leu: n = 51; ATG5 KO: +aa: n = 36; -aa: n = 35; Leu: n = 47 cells). (h) ATG5-KO macrophages were co-incubated with vehicle or CC ± leucine and percent of caspase 3/7-positive cells were quantified in >103 cells from acquired images (-aa: n = 13; -aa+cc: n = 11; Leu: n = 10; Leu+ cc.: n = 12 image fields). For all graphs, data presented as mean ±SEM. *P < 0.05, one-way ANOVA with Tukey’s test.
Extended Data Fig. 7 Common serum metabolites of mϕRaptor-KO and dual mϕRaptor/mϕATG5-KO (DKO) mice fed a standard Western diet.
Measurements of serum cholesterol, glucose, triglycerides, and free fatty acids in cohorts of mϕRaptor-KO (n = 18) and mϕRaptor/ mϕATG5-KO (DKO) mice (n = 14) (all on ApoE-null background) fed a standard Western diet for 8 weeks.
Supplementary information
Supplementary Information
Supplementary Fig. 1
Source data
Source Data Fig. 4
Unprocessed western blots of Fig. 4
Source Data Fig. 8
Unprocessed western blots of Fig. 8
Source Data Extended Data Fig. 3
Unprocessed western blots of Extended Data Fig. 3
Source Data Extended Data Fig. 4
Unprocessed western blots of Extended Data Fig. 4
Source Data Extended Data Fig. 6
Unprocessed western blots of Extended Data Fig. 6
Rights and permissions
About this article
Cite this article
Zhang, X., Sergin, I., Evans, T.D. et al. High-protein diets increase cardiovascular risk by activating macrophage mTOR to suppress mitophagy. Nat Metab 2, 110–125 (2020). https://doi.org/10.1038/s42255-019-0162-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-019-0162-4
This article is cited by
-
Rab4A-directed endosome traffic shapes pro-inflammatory mitochondrial metabolism in T cells via mitophagy, CD98 expression, and kynurenine-sensitive mTOR activation
Nature Communications (2024)
-
Dysregulated cellular metabolism in atherosclerosis: mediators and therapeutic opportunities
Nature Metabolism (2024)
-
A leucine–macrophage mTORC1 connection drives increased risk of atherosclerosis with high-protein diets
Nature Metabolism (2024)
-
Dietary cysteine and methionine promote peroxisome elevation and fat loss by induction of CG33474 expression in Drosophila adipose tissue
Cellular and Molecular Life Sciences (2024)
-
Tryptophanylation of insulin receptor by WARS attenuates insulin signaling
Cellular and Molecular Life Sciences (2024)