Abstract
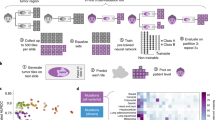
We use deep transfer learning to quantify histopathological patterns across 17,355 hematoxylin and eosin-stained histopathology slide images from 28 cancer types and correlate these with matched genomic, transcriptomic and survival data. This approach accurately classifies cancer types and provides spatially resolved tumor and normal tissue distinction. Automatically learned computational histopathological features correlate with a large range of recurrent genetic aberrations across cancer types. This includes whole-genome duplications, which display universal features across cancer types, individual chromosomal aneuploidies, focal amplifications and deletions, as well as driver gene mutations. There are widespread associations between bulk gene expression levels and histopathology, which reflect tumor composition and enable the localization of transcriptomically defined tumor-infiltrating lymphocytes. Computational histopathology augments prognosis based on histopathological subtyping and grading, and highlights prognostically relevant areas such as necrosis or lymphocytic aggregates. These findings show the remarkable potential of computer vision in characterizing the molecular basis of tumor histopathology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
TCGA data (images, as well as genomic, transcriptomic and clinical data) are publically available from http://gdc.cancer.gov. For METABRIC, images and genomic and transcriptomic data are available under controlled access at the European Genome-phenome Archive (https://ega-archive.org/) under study accession EGAS00000000098, and clinical data are available at https://www.cbioportal.org/. For BASIS, genomic data are freely available from ftp://ftp.sanger.ac.uk/pub/cancer/Nik-ZainalEtAl-560BreastGenomes, clinical data are published42, and histopathology images are available under controlled access at the European Genome-phenome Archive via accession EGAS00001001178. All other data supporting the findings of this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
The computational histopathology algorithm and analysis code are available at https://github.com/gerstung-lab/PC-CHiP. The retrained checkpoints for Inception-V4 and amended Inception-V4 architecture are available from the BioStudies database (https://www.ebi.ac.uk/biostudies/) under accession number S-BSST292. Source data are provided with this paper.
References
Lindeman, N. I. et al. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: guideline from the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology. J. Thorac. Oncol. 8, 823–859 (2013).
Woodman, S. E., Lazar, A. J., Aldape, K. D. & Davies, M. A. New strategies in melanoma: molecular testing in advanced disease. Clin. Cancer Res. 18, 1195–1200 (2012).
Russnes, H. G., Lingjærde, O. C., Børresen-Dale, A.-L. & Caldas, C. Breast cancer molecular stratification: from intrinsic subtypes to integrative clusters. Am. J. Pathol. 187, 2152–2162 (2017).
Dienstmann, R. et al. Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat. Rev. Cancer 17, 79–92 (2017).
Cancer Genome Atlas Research Network. The molecular taxonomy of primary prostate cancer. Cell 163, 1011–1025 (2015).
Bailey, P. et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531, 47–52 (2016).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115–118 (2017).
Coudray, N. et al. Classification and mutation prediction from non-small cell lung cancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25, 1301–1309 (2019).
Bera, K., Schalper, K. A., Rimm, D. L., Velcheti, V. & Madabhushi, A. Artificial intelligence in digital pathology—new tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 16, 703–715 (2019).
Hegde, N. et al. Similar image search for histopathology: SMILY. NPJ Digit. Med. 2, 56 (2019).
Saltz, J. et al. Spatial organization and molecular correlation of tumor-infiltrating lymphocytes using deep learning on pathology images. Cell Rep. 23, 181–193.e7 (2018).
Shia, J. et al. Morphological characterization of colorectal cancers in The Cancer Genome Atlas reveals distinct morphology–molecular associations: clinical and biological implications. Mod. Pathol. 30, 599–609 (2017).
Schaumberg, A. J., Rubin, M. A. & Fuchs, T. J. H&E-stained whole slide image deep learning predicts SPOP mutation state in prostate cancer. Preprint at bioRxiv https://doi.org/10.1101/064279 (2018).
Tsou, P. & Wu, C.-J. Mapping driver mutations to histopathological subtypes in papillary thyroid carcinoma: applying a deep convolutional neural network. J. Clin. Med. Res. 8, 1675 (2019).
Chang, P. et al. Deep-learning convolutional neural networks accurately classify genetic mutations in gliomas. AJNR Am. J. Neuroradiol. 39, 1201–1207 (2018).
Kather, J. N. et al. Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat. Med. 25, 1054–1056 (2019).
Kather, J. N., Heij, L. R., Grabsch, H. I. & Kooreman, L. F. S. Pan-cancer image-based detection of clinically actionable genetic alterations. Nat. Cancer https://doi.org/10.1038/s43018-020-0087-6 (2020).
Kather, J. N., Schulte, J., Grabsch, H. I., Loeffler, C. & Muti, H. S. Deep learning detects virus presence in cancer histology. Preprint at bioRxiv https://doi.org/10.1101/690206 (2019).
Yu, K.-H. et al. Association of omics features with histopathology patterns in lung adenocarcinoma. Cell Syst. 5, 620–627.e3 (2017).
Schmauch, B., Romagnoni, A., Pronier, E. & Saillard, C. Transcriptomic learning for digital pathology. Preprint at bioRxiv https://doi.org/10.1101/760173 (2019).
Mobadersany, P. et al. Predicting cancer outcomes from histology and genomics using convolutional networks. Proc. Natl Acad. Sci. USA 115, E2970–E2979 (2018).
Cheng, J. et al. Integrative analysis of histopathological images and genomic data predicts clear cell renal cell carcinoma prognosis. Cancer Res. 77, e91–e100 (2017).
Yu, K.-H. et al. Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features. Nat. Commun. 7, 12474 (2016).
Szegedy, C., Ioffe, S., Vanhoucke, V. & Alemi, A. A. Inception-v4, Inception-ResNet and the impact of residual connections on learning. in Proceedings of the Thirty-First AAAI Conference on Artificial Intelligence. AAAI Press 4, 4278–4284 (2017).
Cooper, L. A. et al. PanCancer insights from The Cancer Genome Atlas: the pathologist’s perspective. J. Pathol. 244, 512–524 (2018).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature 578, 122–128 (2020).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Cancer Genome Atlas Research Network. Comprehensive and integrated genomic characterization of adult soft tissue sarcomas. Cell 171, 950–965.e28 (2017).
Zack, T. I. et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, 1134–1140 (2013).
Burger, P. C. et al. Small cell architecture—a histological equivalent of EGFR amplification in glioblastoma multiforme? J. Neuropathol. Exp. Neurol. 60, 1099–1104 (2001).
Verhaak, R. G. W. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2010).
Kebebew, E. et al. The prevalence and prognostic value of BRAF mutation in thyroid cancer. Ann. Surg. 246, 466–470 (2007).
O’Hara, A. J. & Bell, D. W. The genomics and genetics of endometrial cancer. Adv. Genomics Genet. 2012, 33–47 (2012).
Thorsson, V. et al. The immune landscape of cancer. Immunity 48, 812–830.e14 (2018).
Nawaz, S., Heindl, A., Koelble, K. & Yuan, Y. Beyond immune density: critical role of spatial heterogeneity in estrogen receptor-negative breast cancer. Mod. Pathol. 28, 766–777 (2015).
Cabrita, R. et al. Tertiary lymphoid structures improve immunotherapy and survival in melanoma. Nature 577, 561–565 (2020).
Pollheimer, M. J. et al. Tumor necrosis is a new promising prognostic factor in colorectal cancer. Hum. Pathol. 41, 1749–1757 (2010).
Jögi, A., Vaapil, M., Johansson, M. & Påhlman, S. Cancer cell differentiation heterogeneity and aggressive behavior in solid tumors. Ups. J. Med. Sci. 117, 217–224 (2012).
Gooden, M. J. M., de Bock, G. H., Leffers, N., Daemen, T. & Nijman, H. W. The prognostic influence of tumour-infiltrating lymphocytes in cancer: a systematic review with meta-analysis. Br. J. Cancer 105, 93–103 (2011).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Nik-Zainal, S. et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534, 47–54 (2016).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Bayraktar, O. A., Bartels, T., Polioudakis, D. & Holmqvist, S. Single-cell in situ transcriptomic map of astrocyte cortical layer diversity. Preprint at bioRxiv https://doi.org/10.1101/432104 (2018).
Ke, R. et al. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10, 857–860 (2013).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2018).
Elston, R. C. On Fisher’s method of combining P-values. Biom. J. 33, 339–345 (1991).
Wilcoxon, F. Individual comparisons by ranking methods. Biom. Bull. 1, 80–83 (1945).
Mann, H. B. & Whitney, D. R. On a test of whether one of two random variables is stochastically larger than the other. Ann. Math. Stat. 18, 50–60 (1947).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B Stat. Methodol. 57, 289–300 (1995).
Best, D. J. & Roberts, D. E. Algorithm AS 89: the upper tail probabilities of Spearman’s rho. J. R. Stat. Soc. C Appl. Stat. 24, 377–379 (1975).
Harrell, F. E. Jr, Califf, R. M., Pryor, D. B., Lee, K. L. & Rosati, R. A. Evaluating the yield of medical tests. J. Am. Med. Assoc. 247, 2543–2546 (1982).
Grossman, R. L. et al. Toward a shared vision for cancer genomic data. N. Engl. J. Med. 375, 1109–1112 (2016).
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J. & Wojna, Z. Rethinking the inception architecture for computer vision. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 2818–2826 (IEEE, 2016).
Tellez, D. et al. Quantifying the effects of data augmentation and stain color normalization in convolutional neural networks for computational pathology. Preprint at https://arxiv.org/abs/1902.06543 (2019).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
LeDell, E., Petersen, M. & van der Laan, M. Computationally efficient confidence intervals for cross-validated area under the ROC curve estimates. Electron. J. Stat. 9, 1583–1607 (2015).
Fieller, E. C., Hartley, H. O. & Pearson, E. S. Tests for rank correlation coefficients. I. Biometrika 44, 470–481 (1957).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041.e21 (2017).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl Acad. Sci. USA 107, 16910–16915 (2010).
Taylor, A. M. et al. Genomic and functional approaches to understanding cancer aneuploidy. Cancer Cell 33, 676–689.e3 (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Croft, D. et al. The Reactome Pathway Knowledgebase. Nucleic Acids Res. 42, D472–D477 (2014).
Cox, D. R. in Breakthroughs in Statistics (eds Kotz, S. & Johnson, N. L.) 527–541 (Springer, 1992).
Simon, N., Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for Cox’s proportional hazards model via coordinate descent. J. Stat. Softw. 39, 1–13 (2011).
Singer, Y. & Duchi, J. C. Efficient learning using forward-backward splitting. Adv. Neural Inf. Proc. Syst. 22, 495–503 (2009).
Snoek, J., Larochelle, H. & Adams, R. P. Practical Bayesian optimization of machine learning algorithms. Adv. Neural Inf. Proc. Syst. 25, 2951–2959 (2012).
Dentro, S. C. et al. Portraits of genetic intra-tumour heterogeneity and subclonal selection across cancer types. Preprint at bioRxiv https://doi.org/10.1101/312041 (2018).
Caicedo, J. C. et al. Nucleus segmentation across imaging experiments: the 2018 Data Science Bowl. Nat. Methods 16, 1247–1253 (2019).
Acknowledgements
A.W.J. and M.G. are supported by grant NNF17OC0027594 from the Novo Nordisk Foundation. L.M. is a recipient of a Cancer Research UK Clinical PhD Fellowship (C20/A20917). L.R.Y. is funded by a Wellcome Trust Clinical Research Career Development Fellowship (214584/Z/18/Z). The results shown here are in part based on data generated by the TCGA Research Network (https://www.cancer.gov/tcga). We thank C. Caldas, S.-F. Chin, Y. Yuan and the METABRIC consortium, as well as M. Stratton, M. Van de Vijver and the BASIS consortium for assistance and sharing data. We also thank all members of the Gerstung laboratory, I, Martincorena and A. Lawson for critical comments on the manuscript.
Author information
Authors and Affiliations
Contributions
Y.F. retrieved and quality controlled all images, developed and trained the deep learning algorithms, performed statistical tests for genomic and molecular association and created all of the figures. A.W.J. performed the survival analysis, reviewed the statistical procedures and applied multiple testing adjustments. R.V.T. and M.G. extended the Inception-V4 algorithm. S.G. provided copy number and annotated mutation data. H.V. extracted mutational signature data. A.S. performed nuclei segmentation. L.R.Y. curated validation data. L.M. oversaw the histopathology review, including blinded assessment of TILs, with help from M.J.-L. M.G. conceived of and supervised the study. Y.F., A.W.J. and M.G. wrote the manuscript with input from L.M. and all other authors, who also approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Computational histopathological features discriminate between different tissue types.
a, UMAP dimensionality reduction representation of the 1,536 histopathological features from randomly selected tiles colored by groups of cancer types (n=200 tiles per tissue type and JPEG quality). b, Example tiles from H&E-stained tissue sections of normal and tumor samples from different cancer types (arranged by row, manually selected from best predicted tiles). All tiles are manually selected from best predicted tiles.
Extended Data Fig. 2 The distribution of predicted tumor purity by histopathological features for samples with different histopathologists evaluated tumor purity.
Each boxplot corresponds to one cancer type, each box corresponds to the predicted tumor purity from histopathological features for samples with the histopathologist evaluated tumor purity indicated on x-axis (total number of slides n=14,862). Boxplots depict the quartiles and median, whiskers extend to 1.5× the inter quartile range.
Extended Data Fig. 3 Pan-cancer morphological features of whole genome duplications.
a, Distribution of cell nucleus size and intensity of samples with and without WGD. Each dot in the scatter plot corresponds to one of 12,000 tiles that were randomly selected across cancer types. The cell nucleus size and intensity were calculated using Cell Profiler with a pipeline provided by the software provider. Boxplots depict the quartiles and median, whiskers extend to 1.5× the inter quartile range. b, AUC from PC-CHiP (y-axis) compared to hard coded features (x-axis) for a set of n=500 randomly selected tiles for each cancer type. Each dot represents a cancer type. Error bars correspond to 95% confidence intervals. c, Histopathological prediction of WGD using 5-fold cross validation (red) and models trained leaving out one cancer type (blue). Error bars correspond to 95% confidence intervals estimated by bootstrap resampling.
Extended Data Fig. 4 Example tiles for associations between computational histopathological and genomic alterations.
a, Four example tiles for chromosome 8q gain (left column) and wild type (right column) breast invasive carcinoma (top row) and esophageal carcinoma (bottom row). b, Four example tiles for chromosome 17p loss (left column) and wild type (right column) for colon adenocarcinoma (top row) and lung squamous cell carcinoma (bottom row). c, Four example tiles for TP53 mutated (left column) and wild type (right column) liver cancer (hepatocellular carcinomas). d, Four example tiles for PTEN mutation (left column) and wild type (right column) for uterine cancer. Representative tiles are selected from 100 best predicted tiles.
Extended Data Fig. 5 Histopathological associations with transcriptomic cell proliferation scores.
a, Example tiles for low proliferation (top row) and high (bottom row) for breast invasive carcinoma, liver hepatocellular carcinoma, thymoma and lung adenocarcinoma. Four example tiles manually selected from best predicted tiles are shown for each tumour type. b, Boxplots show the different transcriptomic proliferation score for tumors with different histological grades for 10 cancer types with available data (n=11,080). G1-G4 corresponds to different grades with G1 being the lowest and G4 the highest, GX stands for “Grade cannot be assessed”, GB stands for “Borderline grade”. p-values were calculated by ANOVA. Boxplots depict the quartiles and median, whiskers extend to 1.5× the inter quartile range. c, Figure shows the increases of predictive accuracy of proliferation score from PC-CHiP compared to conventional histological grades. Each line represents one cancer type with the same colors as in Fig. 1e–g. d, Barplots showing the correlation of transcriptomic proliferation score and the tumor purity estimated by ASCAT (at patient level), histopathology (at patient level) and predicted tumor probability from PC-CHiP (at tile level) in each cancer (n=10,762 tumor samples for ASCAT, n=11,080 tumor samples for histopathology and n=6,188 tumor samples for PC-CHiP).
Extended Data Fig. 6 Accuracy of TIL scores predicted by PC-CHiP.
a, Systematic blinded assessment of TIL raw counts by two expert pathologists for three different cancer types (n=150 for each cancer). Each box plot shows the predicted TIL scores from PC-CHiP for tiles with different TIL raw counts, as independently evaluated by pathologists. b, Publically available slide-level TIL data displays lower concordance compared to with systematic blinded assessment of TIL (n=372 tiles). Each box plot shows the slide level TILs evaluation from TCGA for tiles with different TIL raw counts. Boxplots depict the quartiles and median, whiskers extend to 1.5× the inter quartile range.
Extended Data Fig. 7 Patient risk stratification using histopathological features.
Kaplan-Meier curves for high and low risk groups in different tumor types and stages. a, breast invasive carcinoma. b, stomach adenocarcinoma. c, head and neck squamous cell carcinoma. Only tumor stages with at least 20 patients are shown. Hazard ratios (HR) and the corresponding 95% confidence interval were computed using a Cox proportional hazards model.
Extended Data Fig. 8 Overall performance of PC-CHiP in validation datasets.
a, The validation accuracy in METABRIC (blue) and BASIS (green) datasets compared to TCGA dataset (red) for each significant association discovered in TCGA indicated at the bottom (total number of genomic alterations tested n=82). Each point corresponds to the predicted AUC for the genomic alteration indicated at the bottom. Error bars correspond to 95% confidence intervals. p-value estimated from Wilcox’s rank sum test and adjusted using FDR. b, The distribution of correlation between predicted and true transcript level in METABRIC (x-axis) compared to those in TCGA (y-axis). Each dot represents a gene (n=14,756 genes); blue dots are the genes that can be validated in METABRIC (Spearman’s rank correlation ρ > 0, p-value estimated using two sided t-test, adjusted FDR<0.1).
Extended Data Fig. 9 Histopathological feature representations before and after retraining of Inception-V4.
UMAP representation of the histopathological features from the original Inception model (n=200 tiles randomly selected for each tissue type/JPEG quality) (a, b) and the modified, retrained architecture (c, d). a, lung adenocarcinoma, squamous cell carcinoma and normal lung tissue highlighted. b, breast tumor and normal from TCGA and breast tumor from METABRIC highlighted. c, as in a, but for the modified architecture. d, as c based on the modified architecture. In each figure, the plot on the right side is colored by tissue type and the plot on the left side is colored by jpeg quality.
Extended Data Fig. 10 Molecular associations before and after retraining of Inception-V4.
a, AUC for selected genetic alterations and survival for the original and modified Inception architecture. Error bars denote 95% confidence intervals. Sample sizes are n=149 tumor samples for BASIS; for METABRIC, n=454 tumor samples were used for WGD status and copy number alterations; n=434 tumor samples were used for driver gene mutations. Additional details can be found in Supplementary Table 4. b, Whole-slide average histopathology predictions for TILs from the modified network (x-axis) relative to expert pathologist categories (y-axis). Boxplots depict the quartiles and median, whiskers extend to 1.5× the inter quartile range. Shown are n=36 tumor samples for METABRIC and n=129 tumor samples for BASIS with available TIL annotation c, Distribution of validated (deep green), indeterminate (light green) and invalid (gray) associations in METABRIC and BASIS across different alteration types. Distribution of validated (deep green), indeterminate (light green) and invalid (gray) transcriptomic associations in METABRIC. Sample sizes for genomic associations as in a; n=456 tumor samples were used for transcriptomics. Details can be found in Supplementary Table 4. d, Scatterplots of genomic and transcriptomic association strengths based on the original (x-axis) and modified (y-axis) Inception model for the TCGA cohort. Predictions from the original model are five-fold cross-validated, while those of the modified architecture are evaluated on a single 70% training / 30% testing split. Sample sizes and the number of alterations tested can be found in Supplementary Table 5.
Supplementary information
Supplementary Tables
Supplementary Tables 1–5.
Supplementary Data
High-resolution image tiles at 20× magnification and 512 pixels × 512 pixels (0.5 µm px−1), shown in Figs. 1, 3, 4 and 6–8 and Extended Data Figs. 4 and 5.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Fu, Y., Jung, A.W., Torne, R.V. et al. Pan-cancer computational histopathology reveals mutations, tumor composition and prognosis. Nat Cancer 1, 800–810 (2020). https://doi.org/10.1038/s43018-020-0085-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-0085-8
This article is cited by
-
Slideflow: deep learning for digital histopathology with real-time whole-slide visualization
BMC Bioinformatics (2024)
-
Deep learning in cancer genomics and histopathology
Genome Medicine (2024)
-
Pathogenomics for accurate diagnosis, treatment, prognosis of oncology: a cutting edge overview
Journal of Translational Medicine (2024)
-
Regression-based Deep-Learning predicts molecular biomarkers from pathology slides
Nature Communications (2024)
-
A systematic pan-cancer study on deep learning-based prediction of multi-omic biomarkers from routine pathology images
Communications Medicine (2024)