Abstract
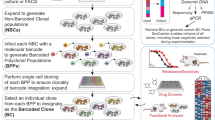
Lineage-tracing methods have enabled characterization of clonal dynamics in complex populations, but generally lack the ability to integrate genomic, epigenomic and transcriptomic measurements with live-cell manipulation of specific clones of interest. We developed a functionalized lineage-tracing system, ClonMapper, which integrates DNA barcoding with single-cell RNA sequencing and clonal isolation to comprehensively characterize thousands of clones within heterogeneous populations. Using ClonMapper, we identified subpopulations of a chronic lymphocytic leukemia cell line with distinct clonal compositions, transcriptional signatures and chemotherapy survivorship trajectories; patterns that were also observed in primary human chronic lymphocytic leukemia. The ability to retrieve specific clones before, during and after treatment enabled direct measurements of clonal diversification and durable subpopulation transcriptional signatures. ClonMapper is a powerful multifunctional approach to dissect the complex clonal dynamics of tumor progression and therapeutic response.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All HG3 epigenomic sequencing data that support the findings of this study have been deposited in the Gene Expression Omnibus under accession code GSE151431. Previously published whole-exome sequencing data for fludarabine and ibrutinib-treated patients were from dbGaP with accession numbers phs000922.v1.p1 and phs001431.v1.p, respectively. Previously published scRNA-seq data for ibrutinib-treated patients are available from dbGaP with accession number phs2335.v1. All other data supporting the findings of this study are available from the corresponding authors on reasonable request. Source data are provided with this paper.
Code availability
The single-cell RNA data were processed using Cell Ranger v.3.1 (https://www.10xgenomics.com/) and analyzed with the Python package scanpy v.1.4.5 (https://scanpy.readthedocs.io/en/stable/). Barcode sequence processing was conducted using our cashier algorithm (v.0.1) (https://github.com/brocklab/cashier). Code for modeling growth dynamics is also available on GitHub (https://github.com/kej1993johnson/Lineage_transitions). Any additional code is available from the corresponding authors upon request.
References
Marusyk, A., Janiszewska, M. & Polyak, K. Intratumor heterogeneity: the rosetta stone of therapy resistance. Cancer Cell 37, 471–484 (2020).
Turajlic, S., Sottoriva, A., Graham, T. & Swanton, C. Resolving genetic heterogeneity in cancer. Nat. Rev. Genet. 20, 404–416 (2019).
Egan, J. B. et al. Whole-genome sequencing of multiple myeloma from diagnosis to plasma cell leukemia reveals genomic initiating events, evolution, and clonal tides. Blood 120, 1060–1066 (2012).
Kasar, S. et al. Whole-genome sequencing reveals activation-induced cytidine deaminase signatures during indolent chronic lymphocytic leukaemia evolution. Nat. Commun. 6, 8866 (2015).
Landau, D. A. et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 152, 714–726 (2013).
Roth, A. et al. PyClone: statistical inference of clonal population structure in cancer. Nat. Methods 11, 396–398 (2014).
Williams, M. J., Sottoriva, A. & Graham, T. A. Measuring clonal evolution in cancer with genomics. Annu. Rev. Genomics Hum. Genet. 20, 309–329 (2019).
Bhang, H. E. et al. Studying clonal dynamics in response to cancer therapy using high-complexity barcoding. Nat. Med. 21, 440–448 (2015).
Hata, A. N. et al. Tumor cells can follow distinct evolutionary paths to become resistant to epidermal growth factor receptor inhibition. Nat. Med. 22, 262–269 (2016).
Kalhor, R., Mali, P. & Church, G. M. Rapidly evolving homing CRISPR barcodes. Nat. Methods 14, 195–200 (2017).
Pei, W. et al. Polylox barcoding reveals haematopoietic stem cell fates realized in vivo. Nature 548, 456–460 (2017).
Kebschull, J. M. & Zador, A. M. Cellular barcoding: lineage tracing, screening and beyond. Nat. Methods 15, 871–879 (2018).
Kong, W. et al. CellTagging: combinatorial indexing to simultaneously map lineage and identity at single-cell resolution. Nat. Protoc. 15, 750–772 (2020).
Weinreb, C., Rodriguez-Fraticelli, A., Camargo, F. D. & Klein, A. M. Lineage tracing on transcriptional landscapes links state to fate during differentiation. Science https://doi.org/10.1126/science.aaw3381 (2020).
Wagner, D. E. et al. Single-cell mapping of gene expression landscapes and lineage in the zebrafish embryo. Science 360, 981–987 (2018).
Wagner, D. E. & Klein, A. M. Lineage tracing meets single-cell omics: opportunities and challenges. Nat. Rev. Genet. https://doi.org/10.1038/s41576-020-0223-2 (2020).
Raj, B., Gagnon, J. A. & Schier, A. F. Large-scale reconstruction of cell lineages using single-cell readout of transcriptomes and CRISPR-Cas9 barcodes by scGESTALT. Nat. Protoc. 13, 2685–2713 (2018).
Alemany, A., Florescu, M., Baron, C. S., Peterson-Maduro, J. & van Oudenaarden, A. Whole-organism clone tracing using single-cell sequencing. Nature 556, 108–112 (2018).
Al’Khafaji, A. M., Deatherage, D. & Brock, A. Control of lineage-specific gene expression by functionalized gRNA barcodes. ACS Synth Biol. 7, 2468–2474 (2018).
Feldman, D. et al. CloneSifter: enrichment of rare clones from heterogeneous cell populations. BMC Biol. 18, 177 (2020).
Umkehrer, C. et al. Isolating live cell clones from barcoded populations using CRISPRa-inducible reporters. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0614-0 (2020).
Datlinger, P. et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods 14, 297–301 (2017).
Neftel, C. et al. An integrative model of cellular states, plasticity, and genetics for glioblastoma. Cell 178, 835–849 (2019).
Landau, D. A. et al. Mutations driving CLL and their evolution in progression and relapse. Nature 526, 525–530 (2015).
Gruber, M. et al. Growth dynamics in naturally progressing chronic lymphocytic leukaemia. Nature 570, 474–479 (2019).
Chen, L. et al. CXCR4 upregulation is an indicator of sensitivity to B-cell receptor/PI3K blockade and a potential resistance mechanism in B-cell receptor-dependent diffuse large B-cell lymphomas. Haematologica 105, 1361–1368 (2020).
Li, L., Chai, Y., Wu, C. & Zhao, L. Chemokine receptor CXCR4: an important player affecting the molecular-targeted drugs commonly used in hematological malignancies. Expert Rev. Hematol. 13, 1387–1396 (2020).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Giovannone, N. et al. Galectin-9 suppresses B cell receptor signaling and is regulated by I-branching of N-glycans. Nat. Commun. 9, 3287 (2018).
Giovannone, N., Smith, L. K., Treanor, B. & Dimitroff, C. J. Galectin–glycan interactions as regulators of B cell immunity. Front. Immunol. 9, 2839 (2018).
Kobayashi, T. et al. Galectin-9 exhibits anti-myeloma activity through JNK and p38 MAP kinase pathways. Leukemia 24, 843–850 (2010).
Kuroda, J. et al. Targeting activating transcription factor 3 by galectin-9 induces apoptosis and overcomes various types of treatment resistance in chronic myelogenous leukemia. Mol. Cancer Res. 8, 994–1001 (2010).
Kageshita, T. et al. Possible role of galectin-9 in cell aggregation and apoptosis of human melanoma cell lines and its clinical significance. Int. J. Cancer 99, 809–816 (2002).
Wiersma, V. R. et al. The glycan-binding protein galectin-9 has direct apoptotic activity toward melanoma cells. J. Invest. Dermatol. 132, 2302–2305 (2012).
Cohen, S. et al. The cytokine midkine and its receptor RPTPzeta regulate B cell survival in a pathway induced by CD74. J. Immunol. 188, 259–CD269 (2012).
Cohen, S. & Shachar, I. Midkine as a regulator of B cell survival in health and disease. Br. J. Pharmacol. 171, 888–895 (2014).
Pietila, M. et al. SORLA regulates endosomal trafficking and oncogenic fitness of HER2. Nat. Commun. 10, 2340 (2019).
Campion, O. et al. Contribution of the low-density lipoprotein receptor family to breast cancer progression. Front. Oncol. 10, 882 (2020).
Knisbacher, B. et al. The CLL-1100 project: towards complete genomic characterization and improved prognostics for CLL. In 62nd ASH Annual Meeting and Exposition, 5–8 December (2020).
Feng, Y. C. et al. c-Myc inactivation of p53 through the pan-cancer lncRNA MILIP drives cancer pathogenesis. Nat. Commun. 11, 4980 (2020).
Hayashi, F. et al. Increased expression of tripartite motif (TRIM) like 2 promotes tumoral growth in human oral cancer. Biochem. Biophys. Res. Commun. 508, 1133–1138 (2019).
Kung, C. P., Khaku, S., Jennis, M., Zhou, Y. & Murphy, M. E. Identification of TRIML2, a novel p53 target, that enhances p53 SUMOylation and regulates the transactivation of proapoptotic genes. Mol. Cancer Res. 13, 250–262 (2015).
Xu, L. et al. Acquired mutations associated with ibrutinib resistance in Waldenstrom macroglobulinemia. Blood 129, 2519–2525 (2017).
Efferth, T. et al. Collateral sensitivity of natural products in drug-resistant cancer cells. Biotechnol. Adv. 38, 107342 (2020).
Goldman, A. et al. Targeting tumor phenotypic plasticity and metabolic remodeling in adaptive cross-drug tolerance. Sci. Signal. https://doi.org/10.1126/scisignal.aas8779 (2019).
Imamovic, L. et al. Drug-driven phenotypic convergence supports rational treatment strategies of chronic infections. Cell 172, 121–134 (2018).
Lin, K. H. et al. Using antagonistic pleiotropy to design a chemotherapy-induced evolutionary trap to target drug resistance in cancer. Nat. Genet. 52, 408–417 (2020).
Tong, M. et al. New insights from the widening homogeneity perspective to target intratumor heterogeneity. Cancer Commun. 38, 17 (2018).
Acar, A. et al. Exploiting evolutionary steering to induce collateral drug sensitivity in cancer. Nat. Commun. 11, 1923 (2020).
Zhang, J., Cunningham, J. J., Brown, J. S. & Gatenby, R. A. Integrating evolutionary dynamics into treatment of metastatic castrate-resistant prostate cancer. Nat. Commun. 8, 1816 (2017).
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
Buttner, M., Miao, Z., Wolf, F. A., Teichmann, S. A. & Theis, F. J. A test metric for assessing single-cell RNA-seq batch correction. Nat. Methods 16, 43–49 (2019).
Vieth, B., Parekh, S., Ziegenhain, C., Enard, W. & Hellmann, I. A systematic evaluation of single cell RNA-seq analysis pipelines. Nat. Commun. 10, 4667 (2019).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Lun, A. T., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
Zhang, J. M., Kamath, G. M. & Tse, D. N. Valid post-clustering differential analysis for single-cell RNA-seq. Cell Syst. 9, 383–392 (2019).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Corces, M. R. et al. Lineage-specific and single-cell chromatin accessibility charts human hematopoiesis and leukemia evolution. Nat. Genet. 48, 1193–1203 (2016).
Liao, Y., Smyth, G. K. & Shi, W. The R package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nucleic Acids Res. 47, e47 (2019).
Ryan, J. A., Brunelle, J. K. & Letai, A. Heightened mitochondrial priming is the basis for apoptotic hypersensitivity of CD4+CD8+ thymocytes. Proc. Natl Acad. Sci. USA 107, 12895–12900 (2010).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Benjamin, D. et al. Calling somatic SNVs and indels with Mutect2. Preprint at bioRxiv https://doi.org/10.1101/861054 (2019).
Kim, S. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods 15, 591–594 (2018).
Chen, X. et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics 32, 1220–1222 (2016).
Wala, J. A. et al. SvABA: genome-wide detection of structural variants and indels by local assembly. Genome Res. 28, 581–591 (2018).
Sun, R. et al. Breakpointer: using local mapping artifacts to support sequence breakpoint discovery from single-end reads. Bioinformatics 28, 1024–1025 (2012).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. JMLR 12, 2825–2830 (2011).
Stegle, O., Parts, L., Durbin, R. & Winn, J. A Bayesian framework to account for complex non-genetic factors in gene expression levels greatly increases power in eQTL studies. PLoS Comput. Biol. 6, e1000770 (2010).
Acknowledgements
We acknowledge S. Pollock, L. Nguyen, H. Lyon and C. Patterson for expert project management. We thank C. Hahn, I. Leschiner and B. Persaud for excellent input on human sequencing analysis. We acknowledge support from the National Institutes of Health (5R21CA212928, 5R01CA226258 to A.B.; and 3P01CA206978-03S1, 1U10CA180861-01, 1P01CA206978-01 to C.J.W.). C.J.W. is a Scholar of the Leukemia and Lymphoma Society and C.G. is a Scholar through the American Society of Hematology MMSAP Program and the F31 Diversity Individual Predoctoral Fellowship program through the NCI. E.B. is a recipient of the University of Texas at Austin Provost’s Graduate Excellence Fellowship and the F.M. Jones & H.L. Bruce Endowed Graduate Fellowship. K.J. is grateful for support through a National Science Foundation Graduate Research Fellowship (1610403). B.A.K. was supported by a long-term EMBO fellowship (ALTF 14-2018). We thank the Dana-Farber Flow Cytometry Core, the Broad Institute Walk-Up Sequencing Core and the Genome Sequencing and Analysis Facility at the University of Texas at Austin for their services. All scRNA-seq workflows were conducted in the Translational Immunogenomics Laboratory at the Dana-Farber Cancer Institute.
Author information
Authors and Affiliations
Contributions
C.G. and A.M.A. conceptualized and designed the research. C.G., A.M.A. and E.B. performed and analyzed the bulk of the experiments, under supervision by A. Brock. and C.J.W. K.E.J. performed all mathematical modeling and growth and death rate analyses. Z.L. performed all WGS analyses, supervised by G.G. R.D. generated the computational pipelines for barcode analysis and processed the RNA-seq and ATAC-seq data. S.L. and K.J.L. prepared the 10X Genomics scRNA-seq libraries. S.P. and A.L. performed and analyzed the BH3 profiling analyses. S.G. performed single-cell experiments and provided and facilitated transfer of data from patient samples. B.A.K. collected, processed and analyzed bulk RNA-seq from patients with CLL and compared to HG3. W.Z. and A. Biran processed patient samples for analysis. L.R. and T.J.K. provided primary CLL samples. E.B., K.E.J. and D.N. contributed to statistical analyses. All authors participated in data analysis. C.G., A.M.A. and E.B. wrote the manuscript. All authors reviewed the manuscript before submission. C.J.W. and A. Brock. equally and jointly supervised this work.
Corresponding authors
Ethics declarations
Competing interests
C.J.W. holds equity in BioNTech Inc. and receives research funding from Pharmacyclics. C.J.W. and D.N. have been consultants for H3 Biomedicine and received research funding from Celgene. G.G. receives research funds from IBM and Pharmacyclics and is an inventor of several bioinformatics-related patents, including related to MuTect and ABSOLUTE. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Generation and analysis of high-diversity barcode populations.
a, UMAP of expression profiles derived from bulk RNA-seq of 6 untreated HG3 samples (TP0) and 610 treatment-naïve primary CLLs annotated by expression cluster (left) and IGHV mutation status (right)39. All 616 samples were pre-processed together by using PEER factors to remove technical effects69. b, Hierarchical clustering of the expression profiles used in (a) further supports the similarity between HG3 and IGHV-unmutated primary CLLs as noted by the dendrogram sub-tree highlights (red, HG3; yellow, IGHV-unmutated). c, Diversity of the ClonMapper library as estimated by deep sequencing. Green dots correspond to total unique read counts at corresponding read depths. Approximately 68 million unique barcodes were identified at a read depth of 138 million reads (red line), and fitting of the sampled data (blue line) yields unique barcode estimates of ~7.6 × 107. d, Barcode frequency distribution of the ClonMapper library as quantified by deep sequencing (depth of 138 million reads with no base below Q30). e, CellTiter-Glo analysis of 72-hour dose curves using each chemotherapeutic alone (n = 2 cell culture replicates from one independent experiment, respectively) or (f) in combination at molar ratios equivalent to each drug’s respective LD50 concentrations (HG3 n = 2, REC1 n = 3 cell culture replicates as in d above). g, Ridgeline plots representing the log2 fold change of barcode proportions from TP0 to TP1, where barcodes are grouped by their presence across 8 barcoded cell culture replicates. Only barcodes at or above 0.005% (HG3) and 0.0005% (REC1) abundance at TP0 are shown. h, Unique barcode read counts of TP0 and TP1 barcoded cell culture replicates 1-8 normalized to log10 counts per million. Barcodes are sorted in descending order by the sum of their counts across TP1 columns. i, Venn diagrams of clones occupying HS versus LS clusters at TP0 and TP1.
Extended Data Fig. 2 Marker gene expression over time across survivorship clusters.
a, Heatmap of marker gene expression upregulated in TP0 vs TP1 HS (top; rows annotated by vertical orange bar), and TP1 vs TP0 HS (bottom; rows annotated by vertical red bar). b, Heatmap of marker gene expression upregulated in TP0 vs TP1 LS (top; rows annotated by vertical blue bar), and TP1 vs TP0 LS (top; rows annotated by vertical green bar). All with log2(FC) > 2, q < 0.05.
Extended Data Fig. 3 Molecular characteristics of high and low survivorship populatios.
a, Correlation plots of unique barcode abundance as determined by barcode sampling of CD18+ and CXCR4+ isolated populations (n = 1 biologically independent sample). b, Venn diagram of overlap between unique barcodes identified by barcode sampling of FACS-sorted populations and unique barcodes identified by scRNA-seq. c, Correlation plot of differentially expressed genes between HS and LS subpopulations as measured by scRNA-seq and bulk RNA-seq (n = 1,921 genes). d, Two top pathways (FDR < 0.25, nominal P < 0.05) identified through gene set enrichment analysis of differentially expressed genes identified through bulk RNA-sequencing. e, Correlation plot of genes differentially expressed by RNA-seq that also have differentially accessible peaks by ATAC-seq. Spearman correlation = 0.63; n = 4010 genes. f, Transcription factor motif enrichment within differentially accessible ATAC-seq peaks (DESeq2 two-sided Wald test q < 0.05, n = 6 paired samples) in HS versus LS subpopulations. Motif scores are E-values produced by AME using Fisher’s exact test. Red; transcription factors involved in Wnt and Notch signaling. g, Log2(FC) of transcription factor motif enrichment within differentially accessible ATAC-seq peaks (DESeq2 2-sided Wald test q < 0.05, n = 6 paired samples) in HS versus LS subpopulations, colored by gene expression. h, Viability assay of the HS subpopulation treated with small molecule inhibitors of the Wnt, Notch, or CXCR4 pathways. ANOVA with Dunnett’s test was used to adjust the p values for comparison of each experimental condition to the single control (n = 3 biologically independent samples). Whiskers, minimum and maximum values, box, first and third quartile, line, median.
Extended Data Fig. 4 High and low survivorship signatures in primary CLL samples.
a, Transcriptomic enrichment analysis of the CXCR4, Wnt, and Notch signaling scores (n = 23 patients; one-sided t-test; adjustments were not made for multiple comparisons). b, UMAP of scRNA-seq from patient samples (n = 4 patients) post-FCR therapy. Samples are annotated by HS and LS signatures. c, UMAP of scRNA-seq from patients 1 and 3 annotated by Wnt signature. d, Signaling enrichment in matched transcriptomes collected from a 13-patient cohort with IGHV-unmutated CLL before and 6-12 months after initiating ibrutinib therapy (one-sided t-test; adjustments were not made for multiple comparisons). e, UMAP of scRNA-seq from patient samples before and during ibrutinib therapy (n = 2 patients). f, Schematic of CXCR4, Wnt and Notch signaling in HS clones. Transmitting ligands and receiving receptors are both upregulated in HS as compared to LS clones. Intensity of color (blue if transmitting, red if receiving) indicates the log2(FC) in expression identified through bulk RNA-sequencing.
Extended Data Fig. 5 Differential growth dynamics of high and low survivorship subpopulations.
a, Fitted growth rate of unique clones at TP0 across 4 parallel barcoded cell culture replicates labeled by HS (red) vs LS (blue) classification. b, Percentage of population belonging to HS or LS subpopulations over 96-hour outgrowth. c, Cell counts of bulk, HS and LS subpopulations at TP0 and TP1. LS TP1 cell counts were statistically significantly higher (P = 0.00029) than TP0 cell counts. There was no statistically significant difference between HS TP0 and TP1 cell counts (P = 0.639). P-values calculated using a two-sided two-sample Student’s t-test (n = 1 of 2 biologically independent experiments with 3 technical replicates each, error bars denote mean±SD; adjustments not made for multiple comparison). d, Stacked bar graphs depict HS and LS subpopulations (marked by GFP or RFP, respectively) co-cultured at different proportions and analyzed by flow cytometry over time. e, UMAP of scRNA-seq data from patient samples post-FCR therapy annotated by LGALS9 and SORL1 expression (red outline, high survivorship expression; blue outline, low survivorship expression - as demonstrated in Extended Data Fig. 4c). f, BH3 profiling of LS versus HS subpopulations. Statistical significance determined using the Holm-Sidak method for multiple comparison two-sided t-tests, alpha=0.05 (n = 1 biologically independent experiment with 3 technical replicates, error bars denote mean ± SD). g, Percent abundance of barcodes at TP0 and TP0.5 across 4 parallel barcoded cell culture replicates labeled by HS (red), LS (blue), and unassigned (grey) classification. h, Ordered distribution of barcode abundance at each timepoint, where each bar represents a lineage. Red, HS. Blue, LS. Gray, unassigned. i-j, UMAP of scRNA-seq data from TP0, TP0.5 and TP1 with top 10 lineages and Leiden clusters annotated (n = 14,505 cells). k, Scaled expression of genes corresponding to pathways from GSEA analysis (input from 2-sided Wilcoxon rank-sum comparing TP0.5 HS versus LS clusters).
Extended Data Fig. 6 Clonal diversification.
a, Cancer cell fractions (CCF) of representative HS mutations enriched upon chemotherapy relapse in cohort of 71 patients with CLL treated with fludarabine, cyclophosphamide and rituximab24,25. b, Change in tumor mutation burden (TMB) in 71 patients with relapsed CLL treated with fludarabine and cyclophosphamide or fludarabine, cyclophosphamide, and rituximab. Raw difference (left; two-tailed paired t-test, t = −2.72, P = 0.008). Right, log2(FC) fold change.
Supplementary information
Supplementary Tables
Supplementary Tables 1–9
Source data
Fig. 1
Statistical source data.
Fig. 2
Statistical source data.
Fig. 2
Reconstructed western blot replicates 1–3.
Fig. 3
Statistical source data.
Fig. 4
Statistical source data.
Fig. 5
Statistical source data.
Extended Data Fig. 1
Statistical source data.
Extended Data Fig. 2
Statistical source data.
Extended Data Fig. 3
Statistical source data.
Extended Data Fig. 4
Statistical source data.
Extended Data Fig. 5
Statistical source data.
Extended Data Fig. 6
Statistical source data.
Rights and permissions
About this article
Cite this article
Gutierrez, C., Al’Khafaji, A.M., Brenner, E. et al. Multifunctional barcoding with ClonMapper enables high-resolution study of clonal dynamics during tumor evolution and treatment. Nat Cancer 2, 758–772 (2021). https://doi.org/10.1038/s43018-021-00222-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-021-00222-8
This article is cited by
-
Decoding leukemia at the single-cell level: clonal architecture, classification, microenvironment, and drug resistance
Experimental Hematology & Oncology (2024)
-
Tracing back primed resistance in cancer via sister cells
Nature Communications (2024)
-
Unraveling non-genetic heterogeneity in cancer with dynamical models and computational tools
Nature Computational Science (2023)
-
A global sensitivity analysis of a mechanistic model of neoadjuvant chemotherapy for triple negative breast cancer constrained by in vitro and in vivo imaging data
Engineering with Computers (2023)
-
Best Practices in Designing, Sequencing, and Identifying Random DNA Barcodes
Journal of Molecular Evolution (2023)