Featured
-
-
Article
| Open Access Structured information extraction from scientific text with large language models
Structured information extraction from scientific text with large language models
Extracting scientific data from published research is a complex task required specialised tools. Here the authors present a scheme based on large language models to automatise the retrieval of information from text in a flexible and accessible manner.
- John Dagdelen
- , Alexander Dunn
- & Anubhav Jain
-
Article
| Open Access Overlay databank unlocks data-driven analyses of biomolecules for all
Overlay databank unlocks data-driven analyses of biomolecules for all
In this work, the authors report NMR lipids Databank to promote decentralised sharing of biomolecular molecular dynamics (MD) simulation data with an overlay design. Programmatic access enables analyses of rare phenomena and advances the training of machine learning models.
- Anne M. Kiirikki
- , Hanne S. Antila
- & O. H. Samuli Ollila
-
Article
| Open Access vcfdist: accurately benchmarking phased small variant calls in human genomes
vcfdist: accurately benchmarking phased small variant calls in human genomes
Accurately benchmarking small variant calling accuracy is critical for the continued improvement of human genome sequencing. Here, the authors show that current approaches are biased towards certain variant representations and develop a new approach to ensure consistent and accurate benchmarking, regardless of the original variant representations.
- Tim Dunn
- & Satish Narayanasamy
-
Article
| Open Access Poor sleep and shift work associate with increased blood pressure and inflammation in UK Biobank participants
Poor sleep and shift work associate with increased blood pressure and inflammation in UK Biobank participants
Circadian disruption is linked to increased blood pressure and heart disease risk. Here, the authors show a positive association between circadian disruption and blood pressure (SBP/DBP) regulation in males and females irrespective of age, weight and inflammatory status.
- Monica Kanki
- , Artika P. Nath
- & Morag J. Young
-
Article
| Open Access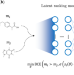 Extracting medicinal chemistry intuition via preference machine learning
Extracting medicinal chemistry intuition via preference machine learning
Over their careers, medicinal chemists develop a gut feeling for what is a promising molecule. Here, the authors use machine learning models to learn this intuition and show that it can be successfully applied in several drug discovery scenarios.
- Oh-Hyeon Choung
- , Riccardo Vianello
- & José Jiménez-Luna
-
Article
| Open Access lesSDRF is more: maximizing the value of proteomics data through streamlined metadata annotation
lesSDRF is more: maximizing the value of proteomics data through streamlined metadata annotation
Public proteomics data often lack essential metadata, limiting their potential. To address this, the authors developed lesSDRF, a tool to simplify the process of metadata annotation, thereby ensuring that data leave a lasting, impactful legacy well beyond their initial publication.
- Tine Claeys
- , Tim Van Den Bossche
- & Lennart Martens
-
Article
| Open Access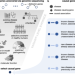 Simulation of undiagnosed patients with novel genetic conditions
Simulation of undiagnosed patients with novel genetic conditions
Rare Mendelian disorders pose a major diagnostic challenge, but evaluation of automated tools that aim to uncover causal genes tools is limited. Here, the authors present a computational pipeline that simulates realistic clinical datasets to address this deficit.
- Emily Alsentzer
- , Samuel G. Finlayson
- & Isaac S. Kohane
-
Article
| Open Access Systematic transcriptional analysis of human cell lines for gene expression landscape and tumor representation
Systematic transcriptional analysis of human cell lines for gene expression landscape and tumor representation
During preclinical drug development, the ability of cancer cell lines to faithfully model human disease is important for identifying potential therapeutic strategies. Here, using transcriptomic datasets of over 1000 cell lines, the authors evaluate how representative each line is of its cancer type and present their cell line selection tool.
- Han Jin
- , Cheng Zhang
- & Adil Mardinoglu
-
Article
| Open Access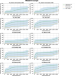 Uncertainty in non-CO2 greenhouse gas mitigation contributes to ambiguity in global climate policy feasibility
Uncertainty in non-CO2 greenhouse gas mitigation contributes to ambiguity in global climate policy feasibility
The potential for the mitigation of global non-CO2 greenhouse gases is highly uncertain. Harmsen et al. estimate this uncertainty and show that it has large implications for the feasibility of reaching the Paris Climate Agreement targets.
- Mathijs Harmsen
- , Charlotte Tabak
- & Detlef van Vuuren
-
Article
| Open Access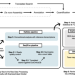 ExpressAnalyst: A unified platform for RNA-sequencing analysis in non-model species
ExpressAnalyst: A unified platform for RNA-sequencing analysis in non-model species
RNA-sequencing data analysis is difficult for non-model species that have no reference genome. ExpressAnalyst enables RNA-sequencing analysis for any eukaryotic species in less than 24 h, on a laptop, and without any programming.
- Peng Liu
- , Jessica Ewald
- & Jianguo Xia
-
Article
| Open Access Fast, accurate antibody structure prediction from deep learning on massive set of natural antibodies
Fast, accurate antibody structure prediction from deep learning on massive set of natural antibodies
Prediction of antibody structures is critical for understanding and designing novel therapeutic and diagnostic molecules. Here, the authors present IgFold: a fast, accurate method for antibody structure prediction using an end-to-end deep learning model.
- Jeffrey A. Ruffolo
- , Lee-Shin Chu
- & Jeffrey J. Gray
-
Article
| Open Access Bladder cancer organoids as a functional system to model different disease stages and therapy response
Bladder cancer organoids as a functional system to model different disease stages and therapy response
Bladder cancer heterogeneity can limit treatment efficacy in individual patients. Here, the authors use patient derived organoids to develop a drug screening pipeline and identify markers of treatment response.
- Martina Minoli
- , Thomas Cantore
- & Marianna Kruithof-de Julio
-
Article
| Open Access Sex differences in allometry for phenotypic traits in mice indicate that females are not scaled males
Sex differences in allometry for phenotypic traits in mice indicate that females are not scaled males
Research aimed at improving healthcare has largely focused on male animals and cells. Here, the authors use data from the International Mouse Phenotyping Consortium to show that body weight does not account for all phenotypic differences between male and female mice, supporting more female-focused research.
- Laura A. B. Wilson
- , Susanne R. K. Zajitschek
- & Shinichi Nakagawa
-
Article
| Open Access Transcriptomic architecture of nuclei in the marmoset CNS
Transcriptomic architecture of nuclei in the marmoset CNS
Studies of cell heterogeneity in white matter in primates have been limited to date. Here the authors describe a marmoset brain cell atlas that bridges rodent and human data, revealing strong gray-white matter glial segregation.
- Jing-Ping Lin
- , Hannah M. Kelly
- & Daniel S. Reich
-
Article
| Open Access Systematic evidence and gap map of research linking food security and nutrition to mental health
Systematic evidence and gap map of research linking food security and nutrition to mental health
There is a broad range of research available on the relationship between food security and mental health. Here the authors carry out a systematic mapping of evidence on food security and nutrition related to mental health and identifies trends in themes, setting, and study design over the 20 year period studied.
- Thalia M. Sparling
- , Megan Deeney
- & Suneetha Kadiyala
-
Article
| Open Access Endothelial cell heterogeneity and microglia regulons revealed by a pig cell landscape at single-cell level
Endothelial cell heterogeneity and microglia regulons revealed by a pig cell landscape at single-cell level
Pigs are important large animal models for biomedical research. Here, the authors construct a single-cell landscape of pig tissues, unravelling the phenotypic heterogeneity of blood endothelial cells in adipose tissues and the evolutionally conserved regulons of microglia in brains.
- Fei Wang
- , Peiwen Ding
- & Yonglun Luo
-
Article
| Open Access ChIP-Hub provides an integrative platform for exploring plant regulome
ChIP-Hub provides an integrative platform for exploring plant regulome
A comprehensive data portal to explore plant regulomes is still unavailable. Here, the authors develop a web-based platform ChIP-Hub in the ENCODE standards and demonstrate its applications in the identification of hierarchical regulatory network, tissue-specific chromatin dynamics, putative enhancers and chromatin states.
- Liang-Yu Fu
- , Tao Zhu
- & Dijun Chen
-
Article
| Open Access The 4D Nucleome Data Portal as a resource for searching and visualizing curated nucleomics data
The 4D Nucleome Data Portal as a resource for searching and visualizing curated nucleomics data
This paper describes the ‘4DN Data Portal’ that hosts data generated by the 4D Nucleome network, including Hi-C and other chromatin conformation capture assays, as well as various sequencing-based and imaging-based assays. Raw data have been uniformly processed to increase comparability and the portal is implemented with visualization tools to browse the data without download.
- Sarah B. Reiff
- , Andrew J. Schroeder
- & Peter J. Park
-
Article
| Open Access Knowledge integration and decision support for accelerated discovery of antibiotic resistance genes
Knowledge integration and decision support for accelerated discovery of antibiotic resistance genes
Here the authors present KIDS, a knowledge graph integration and phenotypic prediction framework. When applied on antibiotic data, it identifies 6 novel antibiotic resistant E. coli genes that the authors subsequently validate.
- Jason Youn
- , Navneet Rai
- & Ilias Tagkopoulos
-
Comment
| Open Access Multilateral benefit-sharing from digital sequence information will support both science and biodiversity conservation
Multilateral benefit-sharing from digital sequence information will support both science and biodiversity conservation
Ensuring international benefit-sharing from sequence data without jeopardising open sharing is a major obstacle for the Convention on Biological Diversity and other UN negotiations. Here, the authors propose a solution to address the concerns of both developing countries and life scientists.
- Amber Hartman Scholz
- , Jens Freitag
- & Jörg Overmann
-
Article
| Open Access Helical structure motifs made searchable for functional peptide design
Helical structure motifs made searchable for functional peptide design
Here, we present TP-DB; a pattern-based search engine based on 1.67 million helices from the Protein Database (PDB). We demonstrate the utility of TP-DB in identifying microbe-specific antigens, as well as the design of antimicrobial peptides and Protein-protein interaction blockers.
- Cheng-Yu Tsai
- , Emmanuel Oluwatobi Salawu
- & Lee-Wei Yang
-
Article
| Open Access Network medicine for disease module identification and drug repurposing with the NeDRex platform
Network medicine for disease module identification and drug repurposing with the NeDRex platform
There is an unmet need for adaptable tools allowing biomedical researchers to employ network-based drug repurposing approaches for their individual use cases. Here, the authors close this gap with NeDRex, an integrative and interactive platform.
- Sepideh Sadegh
- , James Skelton
- & Tim Kacprowski
-
Article
| Open Access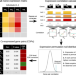 The molecular basis, genetic control and pleiotropic effects of local gene co-expression
The molecular basis, genetic control and pleiotropic effects of local gene co-expression
Local gene co-expression is found throughout the genome, but systematic analysis of these co-expressed genes is needed. Here, the authors identify local co-expressed genes in 49 tissues and characterize the genetic variants which may affect their expression and contribute to disease.
- Diogo M. Ribeiro
- , Simone Rubinacci
- & Olivier Delaneau
-
Article
| Open Access Enhancing CRISPR-Cas9 gRNA efficiency prediction by data integration and deep learning
Enhancing CRISPR-Cas9 gRNA efficiency prediction by data integration and deep learning
High-quality gRNA activity data is needed for accurate on-target efficiency predictions. Here the authors generate activity data for over 10,000 gRNA and build a deep learning model CRISPRon for improved performance predictions.
- Xi Xiang
- , Giulia I. Corsi
- & Yonglun Luo
-
Article
| Open Access Landscape of allele-specific transcription factor binding in the human genome
Landscape of allele-specific transcription factor binding in the human genome
Single-nucleotide variants in enhancers or promoters may affect gene transcription by altering transcription factor binding sites. Here the authors present a meta-analysis empowered by a new statistical method covering thousands of ChIP-Seq experiments resulting in the identification of more than 500 thousand allele-specific binding (ASB) events in the human genome.
- Sergey Abramov
- , Alexandr Boytsov
- & Ivan V. Kulakovskiy
-
Article
| Open Access Sarcoma classification by DNA methylation profiling
Sarcoma classification by DNA methylation profiling
Sarcomas are morphologically heterogeneous tumours rendering their classification challenging. Here the authors developed a classifier using DNA methylation data from several soft tissue and bone sarcoma subtypes, which has the potential to improve classification for research and clinical purposes.
- Christian Koelsche
- , Daniel Schrimpf
- & Andreas von Deimling
-
Perspective
| Open Access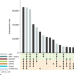 Towards a unified open access dataset of molecular interactions
Towards a unified open access dataset of molecular interactions
The IMEx consortium provides one of the largest resources of curated, experimentally verified molecular interaction data. Here, the authors review how IMEx evolved into a fundamental resource for life scientists and describe how IMEx data can support biomedical research.
- Pablo Porras
- , Elisabet Barrera
- & Sandra Orchard
-
Article
| Open Access Retrospective evaluation of whole exome and genome mutation calls in 746 cancer samples
Retrospective evaluation of whole exome and genome mutation calls in 746 cancer samples
With the generation of large pan-cancer whole-exome and whole-genome sequencing projects, a question remains about how comparable these datasets are. Here, using The Cancer Genome Atlas samples analysed as part of the Pan-Cancer Analysis of Whole Genomes project, the authors explore the concordance of mutations called by whole exome sequencing and whole genome sequencing techniques.
- Matthew H. Bailey
- , William U. Meyerson
- & Christian von Mering
-
Article
| Open Access Ion mobility collision cross-section atlas for known and unknown metabolite annotation in untargeted metabolomics
Ion mobility collision cross-section atlas for known and unknown metabolite annotation in untargeted metabolomics
Collision cross section (CCS) information can aid the annotation of unknown metabolites. Here, the authors optimize the machine-learning based prediction of metabolite CCS values and curate a 1.6 million compound CCS atlas, improving annotation accuracy and coverage for known and unknown metabolites.
- Zhiwei Zhou
- , Mingdu Luo
- & Zheng-Jiang Zhu
-
Article
| Open Access Different scaling of linear models and deep learning in UKBiobank brain images versus machine-learning datasets
Different scaling of linear models and deep learning in UKBiobank brain images versus machine-learning datasets
Schulz et al. systematically benchmark performance scaling with increasingly sophisticated prediction algorithms and with increasing sample size in reference machine-learning and biomedical datasets. Complicated nonlinear intervariable relationships remain largely inaccessible for predicting key phenotypes from typical brain scans.
- Marc-Andre Schulz
- , B. T. Thomas Yeo
- & Danilo Bzdok
-
Article
| Open Access Searching large-scale scRNA-seq databases via unbiased cell embedding with Cell BLAST
Searching large-scale scRNA-seq databases via unbiased cell embedding with Cell BLAST
Single-cell RNA-seq (scRNA-seq) is being widely used to resolve cellular heterogeneity. Here, the authors present a cell-querying method built on a neural network-based generative model and a customized cell-to-cell similarity metric.
- Zhi-Jie Cao
- , Lin Wei
- & Ge Gao
-
Article
| Open Access Construction of a web-based nanomaterial database by big data curation and modeling friendly nanostructure annotations
Construction of a web-based nanomaterial database by big data curation and modeling friendly nanostructure annotations
The low curation of existing nanomaterials’s databases is limiting their application in modeling studies. Here the authors report a publicly available nanomaterial database that contains annotated nanostructures of diverse nanomaterials immediately available for modeling research studies.
- Xiliang Yan
- , Alexander Sedykh
- & Hao Zhu
-
Article
| Open Access A comprehensive non-redundant gene catalog reveals extensive within-community intraspecies diversity in the human vagina
A comprehensive non-redundant gene catalog reveals extensive within-community intraspecies diversity in the human vagina
Reference databases are essential for studies on host-microbiota interactions. Here, the authors present the construction of VIRGO, a human vaginal non-redundant gene catalog, which represents a comprehensive resource for taxonomic and functional profiling of vaginal microbiomes from metagenomic and metatranscriptomic datasets.
- Bing Ma
- , Michael T. France
- & Jacques Ravel
-
Article
| Open Access ProtCID: a data resource for structural information on protein interactions
ProtCID: a data resource for structural information on protein interactions
The authors previously developed the Protein Common Interface Database (ProtCID), which compares and clusters the interfaces of pairs of full-length protein chains with defined Pfam domain architectures in different PDB entries to identify biological assemblies. Here the authors extend ProtCID to the clustering of domain-domain interactions that also allows analyzing domain interactions with peptides, nucleic acids, and ligands.
- Qifang Xu
- & Roland L. Dunbrack Jr.
-
Article
| Open Access A machine-compiled database of genome-wide association studies
A machine-compiled database of genome-wide association studies
Most databases of genotype-phenotype associations are manually curated. Here, Kuleshov et al. describe a machine curation system that extracts such relationships from the GWAS literature and synthesizes them into a structured knowledge base called GWASkb that can complement manually curated databases.
- Volodymyr Kuleshov
- , Jialin Ding
- & Michael Snyder
-
Article
| Open Access FDA-ARGOS is a database with public quality-controlled reference genomes for diagnostic use and regulatory science
FDA-ARGOS is a database with public quality-controlled reference genomes for diagnostic use and regulatory science
To be able to use infectious disease next generation sequencing as a diagnostic tool, appropriate reference datasets are required. Here, Sichtig et al. describe FDA-ARGOS, a reference database for high-quality microbial reference genomes, and demonstrate its utility on the example of two use cases.
- Heike Sichtig
- , Timothy Minogue
- & Uwe Scherf
-
Review Article
| Open Access Towards a standardized bioinformatics infrastructure for N- and O-glycomics
Towards a standardized bioinformatics infrastructure for N- and O-glycomics
Glycomics is gaining momentum in basic, translational and clinical research. Here, the authors review current reporting standards and analysis tools for mass-spectrometry-based glycomics, and propose an e-infrastructure for standardized reporting and online deposition of glycomics data.
- Miguel A. Rojas-Macias
- , Julien Mariethoz
- & Niclas G. Karlsson
-
Perspective
| Open Access Inferring causation from time series in Earth system sciences
Inferring causation from time series in Earth system sciences
Questions of causality are ubiquitous in Earth system sciences and beyond, yet correlation techniques still prevail. This Perspective provides an overview of causal inference methods, identifies promising applications and methodological challenges, and initiates a causality benchmark platform.
- Jakob Runge
- , Sebastian Bathiany
- & Jakob Zscheischler
-
Article
| Open Access Capturing variation impact on molecular interactions in the IMEx Consortium mutations data set
Capturing variation impact on molecular interactions in the IMEx Consortium mutations data set
Genetic variants might exert their functional effects via influencing molecular interaction. Here, the authors present a resource featuring almost 28,000 annotations describing the effect of small sequence changes on physical protein interactions, curated by IMEx Consortium curators.
- J. Khadake
- , B. Meldal
- & P. Porras
-
Article
| Open Access A reference haplotype panel for genome-wide imputation of short tandem repeats
A reference haplotype panel for genome-wide imputation of short tandem repeats
Short-tandem repeats (STR), similar to single nucleotide polymorphisms (SNP), contribute to complex traits, but their ascertainment by next-generation sequencing is costly. Here, Saini et al. provide a SNP+STR haplotype reference panel that allows imputation of STRs from SNP array data.
- Shubham Saini
- , Ileena Mitra
- & Melissa Gymrek
-
Article
| Open Access Haplosaurus computes protein haplotypes for use in precision drug design
Haplosaurus computes protein haplotypes for use in precision drug design
Proteoforms arise as protein isoforms or as protein haplotypes, which are the result of genetic variation. Here, the authors develop Haplosaurus, a database that computes protein haplotypes genome-wide from existing genotype data and analyse protein haplotype variability in the 1000 Genomes dataset.
- William Spooner
- , William McLaren
- & Catherine Chaillan Huntington
-
Article
| Open Access Assessment of the impact of shared brain imaging data on the scientific literature
Assessment of the impact of shared brain imaging data on the scientific literature
Data sharing is recognized as a way to promote scientific collaboration and reproducibility, but some are concerned over whether research based on shared data can achieve high impact. Here, the authors show that neuroimaging papers using shared data are no less likely to appear in top-ranked journals.
- Michael P. Milham
- , R. Cameron Craddock
- & Arno Klein
-
Article
| Open Access Information recovery from low coverage whole-genome bisulfite sequencing
Information recovery from low coverage whole-genome bisulfite sequencing
Here, Libertini and colleagues devise a computation tool that can analyze whole-genome bisulfite sequencing (WGBS) data to recover of ∼30% of the lost differential methylation position information. They use COMETgazer and COMETvintage to analyze 13 diffferent methylome data to demonstrate their performance.
- Emanuele Libertini
- , Simon C. Heath
- & Stephan Beck

 Extracting accurate materials data from research papers with conversational language models and prompt engineering
Extracting accurate materials data from research papers with conversational language models and prompt engineering