Featured
-
-
Article
| Open Access The TDRD3-USP9X complex and MIB1 regulate TOP3B homeostasis and prevent deleterious TOP3B cleavage complexes
The TDRD3-USP9X complex and MIB1 regulate TOP3B homeostasis and prevent deleterious TOP3B cleavage complexes
TDRD3 is a key interaction partner of TOP3B. Here the authors provide molecular mechanisms by which TDRD3 stabilizes TOP3B protein by recruiting the deubiquitylase USP9X. In addition, they show that TDRD3 protects cells from deleterious TOP3B linked DNA and RNA cleavage complexes.
- Sourav Saha
- , Shar-yin Naomi Huang
- & Yves Pommier
-
Article
| Open Access A CK2 and SUMO-dependent, PML NB-involved regulatory mechanism controlling BLM ubiquitination and G-quadruplex resolution
A CK2 and SUMO-dependent, PML NB-involved regulatory mechanism controlling BLM ubiquitination and G-quadruplex resolution
The Bloom syndrome helicase (BLM) unwinds a variety of complex DNA structures including G quadruplex. Here the authors report RNF111-ARKL1-dependent ubiquitination of BLM in PML NBs, which limits BLM protein levels and maintains G quadruplex abundance in the nucleus.
- Shichang Liu
- , Erin Atkinson
- & Bin Wang
-
Article
| Open Access Conformational dynamics of cohesin/Scc2 loading complex are regulated by Smc3 acetylation and ATP binding
Conformational dynamics of cohesin/Scc2 loading complex are regulated by Smc3 acetylation and ATP binding
Cohesin loading on DNA is a highly dynamic process. Here the authors find that Smc3 acetylation blocks the reconfiguration of the Scc2/cohesin complex by preventing Scc2 from interacting with Smc3, and that ATP binding regulates the DNA clamping step by promoting the Scc2/Smc3 coiled-coil interaction.
- Aditi Kaushik
- , Thane Than
- & Bin Hu
-
Article
| Open Access ATR kinase supports normal proliferation in the early S phase by preventing replication resource exhaustion
ATR kinase supports normal proliferation in the early S phase by preventing replication resource exhaustion
The ATR kinase has essential functions apart from its role in DNA replication stress. Here the authors find that in mouse primary B cells ATR tempers the pace of origin firing during the early S phase to avoid exhaustion of dNTPs and other replication factors.
- Demis Menolfi
- , Brian J. Lee
- & Shan Zha
-
Article
| Open Access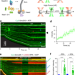 Smc5/6’s multifaceted DNA binding capacities stabilize branched DNA structures
Smc5/6’s multifaceted DNA binding capacities stabilize branched DNA structures
Using single-molecule visualization and manipulation, Chang et al. show that the eukaryotic Smc5/6 complex preferentially binds to and stabilizes ssDNA-dsDNA junctions, which could serve as the molecular basis for its diverse roles in genome maintenance.
- Jeremy T-H. Chang
- , Shibai Li
- & Shixin Liu
-
Article
| Open Access Endocytosis-like DNA uptake by cell wall-deficient bacteria
Endocytosis-like DNA uptake by cell wall-deficient bacteria
Horizontal gene transfer in bacteria can occur through mechanisms such as conjugation, transduction and transformation, which facilitate the passage of DNA across the cell wall. Here, Kapteijn et al. show that cell wall-deficient bacteria can take up DNA and other extracellular materials via an endocytosis-like process.
- Renée Kapteijn
- , Shraddha Shitut
- & Dennis Claessen
-
Article
| Open Access 2-Deoxy-D-glucose couples mitochondrial DNA replication with mitochondrial fitness and promotes the selection of wild-type over mutant mitochondrial DNA
2-Deoxy-D-glucose couples mitochondrial DNA replication with mitochondrial fitness and promotes the selection of wild-type over mutant mitochondrial DNA
It has been a longstanding goal to promote the propagation of functional mitochondrial DNAs at the expense of pathological molecules in cells where the two species coexist. Here, the authors show that restricting the availability of glucose and glutamine can achieve this outcome.
- Boris Pantic
- , Daniel Ives
- & Antonella Spinazzola
-
Article
| Open Access Supercoiling and looping promote DNA base accessibility and coordination among distant sites
Supercoiling and looping promote DNA base accessibility and coordination among distant sites
DNA supercoiling can result in underwinding with negative supercoiling or overwinding with positive supercoiling of the DNA double helix. Here the authors reveal insights into the dynamic relationship between DNA supercoiling-induced sequence-dependent disruptions to base pairing, DNA looping, and the shape of the DNA molecule.
- Jonathan M. Fogg
- , Allison K. Judge
- & Lynn Zechiedrich
-
Article
| Open Access Characterization of a triad of genes in cyanophage S-2L sufficient to replace adenine by 2-aminoadenine in bacterial DNA
Characterization of a triad of genes in cyanophage S-2L sufficient to replace adenine by 2-aminoadenine in bacterial DNA
The cyanophage S-2L incorporates 2-aminoadenine (Z) instead of adenine into its DNA, which still pairs with thymine forming a triple hydrogen bond. Here, the authors identify a third gene mazZ located between purZ and datZ that is required for 2-aminoadenine biosynthesis and determine the crystal structures of MazZ and PurZ. They further show that co-expression of these three genes in E.coli enables 2-aminoadenine incorporation into the bacterial genome.
- Dariusz Czernecki
- , Frédéric Bonhomme
- & Marc Delarue
-
Article
| Open Access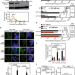 Harmful R-loops are prevented via different cell cycle-specific mechanisms
Harmful R-loops are prevented via different cell cycle-specific mechanisms
Different factors protect cells from harmful R-loops, but the way these are formed is still unclear. Authors show here that R-loops form co-transcriptionally by different manners and cells possess specialized mechanisms to prevent them in each case, a major mechanism being independent of replication and another one being linked to replication.
- Marta San Martin-Alonso
- , María E. Soler-Oliva
- & Andrés Aguilera
-
Article
| Open Access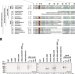 Mechanism of MRX inhibition by Rif2 at telomeres
Mechanism of MRX inhibition by Rif2 at telomeres
Different proteins localised at telomeres ensure chromosome end stability to prevent double strand-end break recognition. Here the authors provide new insight into how in S. cerevisiae the interaction between Rif2 and Rad50 inhibits MRX functions at telomeres.
- Florian Roisné-Hamelin
- , Sabrina Pobiega
- & Stéphane Marcand
-
Article
| Open Access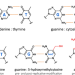 How cyanophage S-2L rejects adenine and incorporates 2-aminoadenine to saturate hydrogen bonding in its DNA
How cyanophage S-2L rejects adenine and incorporates 2-aminoadenine to saturate hydrogen bonding in its DNA
The cyanophage S-2L incorporates 2-aminoadenine (Z) instead of adenine (A) in its genome. Here, the authors provide an explanation for the absence of A in S-2L genome by identifying and characterising functionally and structurally both the HD phosphohydrolase (datZ) that specifically cleaves dATP, and the sole DNA primase-polymerase of S-2L, nonspecific of dATP or dZTP.
- Dariusz Czernecki
- , Pierre Legrand
- & Marc Delarue
-
Article
| Open Access Metabolic pathways inferred from a bacterial marker gene illuminate ecological changes across South Pacific frontal boundaries
Metabolic pathways inferred from a bacterial marker gene illuminate ecological changes across South Pacific frontal boundaries
Extracting functional information from 16S rRNA data surveys would provide a valuable tool for large-scale functional ecology. Here, the authors use PICRUSt2 to infer metabolic functions from bacterial marker gene data across the South Pacific Ocean, and compare them with rate data, biomass estimators and predictions based on shotgun metagenomes.
- Eric J. Raes
- , Kristen Karsh
- & Anya M. Waite
-
Article
| Open Access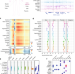 A human tissue map of 5-hydroxymethylcytosines exhibits tissue specificity through gene and enhancer modulation
A human tissue map of 5-hydroxymethylcytosines exhibits tissue specificity through gene and enhancer modulation
DNA 5-hydroxymethylcytosine (5hmC) modification is associated with gene transcription and used as a mark of mammalian development. Here the authors report a comprehensive 5hmC tissue map and analysis of 5hmC genomic distributions in 19 human tissues derived from 10 organ systems, thus providing insights into the role of 5hmC in tissue-specific development.
- Xiao-Long Cui
- , Ji Nie
- & Chuan He
-
Article
| Open Access Telomere damage induces internal loops that generate telomeric circles
Telomere damage induces internal loops that generate telomeric circles
Extrachromosomal circular DNA made of telomeric repeats have been found to have an effect on telomere maintenance. By combining electron microscopy with a telomere purification procedure the authors identify damage-induced i-loops as a key intermediate in telomere circle formation.
- Giulia Mazzucco
- , Armela Huda
- & Ylli Doksani
-
Article
| Open Access The lncRNA lincNMR regulates nucleotide metabolism via a YBX1 - RRM2 axis in cancer
The lncRNA lincNMR regulates nucleotide metabolism via a YBX1 - RRM2 axis in cancer
Despite some well-characterized functions in cancer, the impact of most long non-coding RNAs remains unknown. Here, the authors discover the lncRNA lincNMR which is upregulated in cancer and drives cell proliferation by interacting with YBX1 and controlling nucleotide metabolism.
- Minakshi Gandhi
- , Matthias Groß
- & Sven Diederichs
-
Article
| Open Access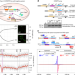 Detection of spacer precursors formed in vivo during primed CRISPR adaptation
Detection of spacer precursors formed in vivo during primed CRISPR adaptation
Primed adaptation in the CRISPR-Cas system helps recognition of previously encountered sequence elements and promotes the formation of new memories. Here the authors characterized spacer precursors of type I-E and type I-F CRISPR-Cas system using in vivo models.
- Anna A. Shiriaeva
- , Ekaterina Savitskaya
- & Ekaterina Semenova
-
Article
| Open Access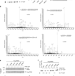 Acetylation regulates ribonucleotide reductase activity and cancer cell growth
Acetylation regulates ribonucleotide reductase activity and cancer cell growth
Ribonucleotide reductase (RNR) catalyzes the de novo synthesis of deoxyribonucleoside diphosphates to provide dNTP precursors for DNA synthesis. Here the authors show that the availability of dNTPs, DNA replication, and cellular proliferation, are modulated by acetylation and deacetylation of RRM2 by KAT7 and Sirt2 respectively.
- Guo Chen
- , Yin Luo
- & Xingming Deng
-
Article
| Open Access Linear mitochondrial DNA is rapidly degraded by components of the replication machinery
Linear mitochondrial DNA is rapidly degraded by components of the replication machinery
Damaged linearized mtDNA needs to be removed from the cell for mitochondrial genome stability. Here the authors shed light into the identity of the machinery responsible for rapidly degrading linearized DNA, implicating the role of mtDNA replication factors.
- Viktoriya Peeva
- , Daniel Blei
- & Wolfram S. Kunz
-
Article
| Open Access ATR inhibition facilitates targeting of leukemia dependence on convergent nucleotide biosynthetic pathways
ATR inhibition facilitates targeting of leukemia dependence on convergent nucleotide biosynthetic pathways
Leukemic cells depend on the nucleotide synthesis pathway to proliferate. Here the authors use metabolomics and proteomics to show that inhibition of ATR reduced the activity of these pathways thus providing a valuable therapeutic target in leukemia.
- Thuc M. Le
- , Soumya Poddar
- & Caius G. Radu
-
Article
| Open Access Structural basis of human PCNA sliding on DNA
Structural basis of human PCNA sliding on DNA
DNA sliding clamps are ring-shaped proteins that encircle DNA and harbour polymerases and other factors that promote processive DNA replication. Here the authors use X-ray crystallography, NMR and MD simulations to propose a model for a PCNA sliding mechanism that relies on short-lived polar interactions.
- Matteo De March
- , Nekane Merino
- & Alfredo De Biasio
-
Article
| Open Access Structural basis of asymmetric DNA methylation and ATP-triggered long-range diffusion by EcoP15I
Structural basis of asymmetric DNA methylation and ATP-triggered long-range diffusion by EcoP15I
Type III restriction–modification enzymes consists of two methylation and one or two restriction subunits. Here the authors report the structure of the full EcoP15I complex bound to DNA, which suggests mechanisms for ATP hydrolysis dependent diffusion along DNA and how a dimeric methyltransferase modifies only one DNA strand.
- Yogesh K. Gupta
- , Siu-Hong Chan
- & Aneel K. Aggarwal
-
Article
| Open Access Targeted DNA degradation using a CRISPR device stably carried in the host genome
Targeted DNA degradation using a CRISPR device stably carried in the host genome
The ability to contain and destroy synthetically engineered microorganisms is an important consideration with environmental, industrial and intellectual property implications. Here Caliando et al. design and demonstrate a stably integrated CRISPR-based system for targeted DNA destruction.
- Brian J. Caliando
- & Christopher A. Voigt

 Mesoscale DNA features impact APOBEC3A and APOBEC3B deaminase activity and shape tumor mutational landscapes
Mesoscale DNA features impact APOBEC3A and APOBEC3B deaminase activity and shape tumor mutational landscapes