Featured
-
-
Article
| Open Access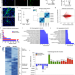 HIRA stabilizes skeletal muscle lineage identity
HIRA stabilizes skeletal muscle lineage identity
The epigenetic mechanisms coordinating the maintenance of adult cellular lineages remain poorly understood. Here the authors demonstrate that HIRA, a H3.3 histone chaperone, establishes the chromatin landscape required for skeletal muscle cell identity.
- Joana Esteves de Lima
- , Reem Bou Akar
- & Frédéric Relaix
-
Article
| Open Access Transcription shapes genome-wide histone acetylation patterns
Transcription shapes genome-wide histone acetylation patterns
Histone acetylation is a ubiquitous hallmark of transcription. Here the authors provide evidence that the majority of histone acetylation is dependent on transcription, specifically due to the requirement of RNAPII for the recruitment and activity of histone acetyltransferases.
- Benjamin J. E. Martin
- , Julie Brind’Amour
- & LeAnn J. Howe
-
Article
| Open Access Hydroxamic acid-modified peptide microarrays for profiling isozyme-selective interactions and inhibition of histone deacetylases
Hydroxamic acid-modified peptide microarrays for profiling isozyme-selective interactions and inhibition of histone deacetylases
Current histone microarrays cannot be used to directly study the transient interactions of histone deacetylases (HDACs). Here, the authors show that hydroxamic acid-modified microarrays can capture HDACs, provide insights into their substrate specificity, and serve to develop peptide inhibitors.
- Carlos Moreno-Yruela
- , Michael Bæk
- & Christian A. Olsen
-
Article
| Open Access An epigenetic gene silencing pathway selectively acting on transgenic DNA in the green alga Chlamydomonas
An epigenetic gene silencing pathway selectively acting on transgenic DNA in the green alga Chlamydomonas
Strong transgene suppression has been observed in Chlamydomonas reinhardtii, but the underlying mechanism is unknown. Here, the authors identify a sirtuin-type histone deacetylase that selectively acts on transgenic DNA to repress gene expression by assembling a repressive chromatin structure composed of deacetylated histones.
- Juliane Neupert
- , Sean D. Gallaher
- & Ralph Bock
-
Article
| Open Access An embryonic stem cell-specific heterochromatin state promotes core histone exchange in the absence of DNA accessibility
An embryonic stem cell-specific heterochromatin state promotes core histone exchange in the absence of DNA accessibility
Nucleosome turnover concomitant with incorporation of the histone variant H3.3 is a hallmark of regulatory regions in the animal genome. Here, the authors demonstrate that fast histone turnover and H3.3 incorporation defines a dynamic heterochromatin state in pluripotent stem cells.
- Carmen Navarro
- , Jing Lyu
- & Simon J. Elsässer
-
Article
| Open Access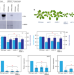 Arabidopsis SWR1-associated protein methyl-CpG-binding domain 9 is required for histone H2A.Z deposition
Arabidopsis SWR1-associated protein methyl-CpG-binding domain 9 is required for histone H2A.Z deposition
The SWI2/SNF2-Related 1 chromatin remodeling complex (SWR1-C) is important for gene regulation, but its composition remains largely uncharacterized in plants. Here, the authors report that methyl-CpG-binding domain 9 (MBD9) is a SWR1-C interacting protein required for histone H2A.Z deposition in Arabidopsis.
- Magdalena E. Potok
- , Yafei Wang
- & Steven E. Jacobsen
-
Article
| Open Access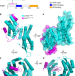 Paf1 and Ctr9 subcomplex formation is essential for Paf1 complex assembly and functional regulation
Paf1 and Ctr9 subcomplex formation is essential for Paf1 complex assembly and functional regulation
The evolutionarily conserved multifunctional polymerase-associated factor 1 (Paf1) complex plays an essential role in gene regulation. Here, the authors report two structures of the human and yeast Ctr9/Paf1 subcomplexes, and find that this is essential for Paf1 complex assembly and function.
- Ying Xie
- , Minying Zheng
- & Jiafu Long
-
Article
| Open Access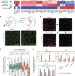 Histone H3.3 sub-variant H3mm7 is required for normal skeletal muscle regeneration
Histone H3.3 sub-variant H3mm7 is required for normal skeletal muscle regeneration
Incorporation of histone H3 variant H3.3 into chromatin regulates transcription. Here the authors find that H3.3 sub-variant H3mm7 is required for skeletal muscle regeneration and that H3mm7 nucleosomes are unstable and exhibit higher mobility, with H3mm7 promoting open chromatin around promoters.
- Akihito Harada
- , Kazumitsu Maehara
- & Yasuyuki Ohkawa
-
Article
| Open Access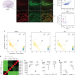 A comprehensive map coupling histone modifications with gene regulation in adult dopaminergic and serotonergic neurons
A comprehensive map coupling histone modifications with gene regulation in adult dopaminergic and serotonergic neurons
The limited size of some neuronal types and their entangled environment renders it difficult to study their transcription regulation. Here the authors present a comparative analysis of histone modifications and transcription in dopaminergic and serotonergic neurons and embryonic neural progenitors.
- Erik Södersten
- , Konstantinos Toskas
- & Johan Holmberg
-
Article
| Open Access H3K14ac is linked to methylation of H3K9 by the triple Tudor domain of SETDB1
H3K14ac is linked to methylation of H3K9 by the triple Tudor domain of SETDB1
SETDB1 is a histone methyltransferase that generates H3K9me3 marks in euchromatic regions. Here the authors show that the triple Tudor domain (3TD) of SETDB1 binds histone H3 tails containing K14 acetylation combined with K9 methylation, and that the K9me–K14ac modification defines a novel chromatin state enriched at SETDB1 binding sites.
- Renata Z. Jurkowska
- , Su Qin
- & Albert Jeltsch

 Learning the histone codes with large genomic windows and three-dimensional chromatin interactions using transformer
Learning the histone codes with large genomic windows and three-dimensional chromatin interactions using transformer