Article
|
Open Access
Featured
-
-
Article
| Open Access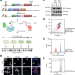 A mutational atlas for Parkin proteostasis
A mutational atlas for Parkin proteostasis
Gene variants can affect folding and stability of the encoded protein. Here, the authors apply deep mutational scanning to provide genotype-phenotype information for 99% of the possible PRKN variants and reveal mechanistic details on how some variants cause loss-of-function and Parkinsons disease.
- Lene Clausen
- , Vasileios Voutsinos
- & Rasmus Hartmann-Petersen
-
Article
| Open Access SEL1L-HRD1 interaction is required to form a functional HRD1 ERAD complex
SEL1L-HRD1 interaction is required to form a functional HRD1 ERAD complex
The importance of the SEL1L-HRD1 interaction in vivo was unclear. Here, authors reported that SEL1L-HRD1 interaction is required to form a functional HRD1 ERAD complex by recruiting the E2 enzyme UBE2J1 and DERLIN to HRD1.
- Liangguang Leo Lin
- , Huilun Helen Wang
- & Ling Qi
-
Article
| Open Access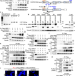 The phosphatase DUSP22 inhibits UBR2-mediated K63-ubiquitination and activation of Lck downstream of TCR signalling
The phosphatase DUSP22 inhibits UBR2-mediated K63-ubiquitination and activation of Lck downstream of TCR signalling
The T cell receptor signalosome integrates multiple positive and negative regulatory elements to finetune the response and limit harmful inflammation. Here authors show a regulatory cascade of T cell activation, in which DUSP22 negatively regulates UBR2, which is an activator of the kinase Lck via K63 ubiquitination.
- Ying-Chun Shih
- , Hsueh-Fen Chen
- & Tse-Hua Tan
-
Article
| Open Access Doa10/MARCH6 architecture interconnects E3 ligase activity with lipid-binding transmembrane channel to regulate SQLE
Doa10/MARCH6 architecture interconnects E3 ligase activity with lipid-binding transmembrane channel to regulate SQLE
Transmembrane E3 ligases are crucial in cellular homeostasis and metabolic regulation. Here, the authors provide the structural details of the ER-resident E3 ligase MARCH6/Doa10, uncovering its unique circular membrane structure and its role in ubiquitylation processes, essential for protein quality control.
- J. Josephine Botsch
- , Roswitha Junker
- & Bastian Bräuning
-
Article
| Open Access Orphan quality control by an SCF ubiquitin ligase directed to pervasive C-degrons
Orphan quality control by an SCF ubiquitin ligase directed to pervasive C-degrons
The prevalence and conservation of C-degrons across eukaryotes is unclear. Here, the authors perform an unbiased survey of C-degrons in budding yeast and identify a C-degron pathway of broad specificity operated by the SCFDas1 ubiquitin ligase.
- Ka-Yiu Edwin Kong
- , Susmitha Shankar
- & Anton Khmelinskii
-
Article
| Open Access The TDRD3-USP9X complex and MIB1 regulate TOP3B homeostasis and prevent deleterious TOP3B cleavage complexes
The TDRD3-USP9X complex and MIB1 regulate TOP3B homeostasis and prevent deleterious TOP3B cleavage complexes
TDRD3 is a key interaction partner of TOP3B. Here the authors provide molecular mechanisms by which TDRD3 stabilizes TOP3B protein by recruiting the deubiquitylase USP9X. In addition, they show that TDRD3 protects cells from deleterious TOP3B linked DNA and RNA cleavage complexes.
- Sourav Saha
- , Shar-yin Naomi Huang
- & Yves Pommier
-
Article
| Open Access Expanding PROTACtable genome universe of E3 ligases
Expanding PROTACtable genome universe of E3 ligases
Proteolysis-targeting chimeras (PROTACs) offer new avenues for drug development. Here the authors investigate E3 ligases—key to PROTAC function—and identify candidate targets for cancer drivers such as KRAS and EGFR.
- Yuan Liu
- , Jingwen Yang
- & Leng Han
-
Article
| Open Access Palmitoylation-driven PHF2 ubiquitination remodels lipid metabolism through the SREBP1c axis in hepatocellular carcinoma
Palmitoylation-driven PHF2 ubiquitination remodels lipid metabolism through the SREBP1c axis in hepatocellular carcinoma
Palmitoylation of proteins can have pathophysiological implications. Here, the authors show that palmitoylation enhances the proteasomal degradation of the histone demethylase PHF2, leading to increased lipogenesis and cell proliferation in an SREBP1c dependent manner and further show that PHF2 acts as an E3 ligase of SREBP1c, suppressing the growth of liver cancer cells.
- Do-Won Jeong
- , Jong-Wan Park
- & Yang-Sook Chun
-
Article
| Open Access A CK2 and SUMO-dependent, PML NB-involved regulatory mechanism controlling BLM ubiquitination and G-quadruplex resolution
A CK2 and SUMO-dependent, PML NB-involved regulatory mechanism controlling BLM ubiquitination and G-quadruplex resolution
The Bloom syndrome helicase (BLM) unwinds a variety of complex DNA structures including G quadruplex. Here the authors report RNF111-ARKL1-dependent ubiquitination of BLM in PML NBs, which limits BLM protein levels and maintains G quadruplex abundance in the nucleus.
- Shichang Liu
- , Erin Atkinson
- & Bin Wang
-
Article
| Open Access Recognition of an Ala-rich C-degron by the E3 ligase Pirh2
Recognition of an Ala-rich C-degron by the E3 ligase Pirh2
Incompletely synthesized nascent polypeptides resulting from ribosome stalling during translation are under surveillance by ribosome-associated quality control. Here, the authors report the molecular mechanism by which the E3 ligase Pirh2 targets the polyalanine tail of aberrant nascent chains for degradation via the C-degron pathway.
- Xiaolu Wang
- , Yao Li
- & Cheng Dong
-
Article
| Open Access Trim-Away ubiquitinates and degrades lysine-less and N-terminally acetylated substrates
Trim-Away ubiquitinates and degrades lysine-less and N-terminally acetylated substrates
TRIM21 mediates intracellular antibody immunity and is exploited for targeted protein degradation using Trim-Away technology. Here, the authors dissect the ubiquitination requirements for Trim-Away, providing an explanation for how TRIM21 can target diverse substrates for degradation.
- Leo Kiss
- , Tyler Rhinesmith
- & Leo C. James
-
Article
| Open Access CRL2ZER1/ZYG11B recognizes small N-terminal residues for degradation
CRL2ZER1/ZYG11B recognizes small N-terminal residues for degradation
N-degron pathways play an important role in maintaining protein homeostasis. Here, Li et al. demonstrates an additional non-Ac/N-degron pathway, in which N-terminal non-acetylated small residue degrons (Ser, Ala, or Cys) are recognized by CRL2ZER1/ZYG11B and targeted for protein degradation.
- Yao Li
- , Yueling Zhao
- & Wenyi Mi
-
Article
| Open Access The assembly of mammalian SWI/SNF chromatin remodeling complexes is regulated by lysine-methylation dependent proteolysis
The assembly of mammalian SWI/SNF chromatin remodeling complexes is regulated by lysine-methylation dependent proteolysis
Here the authors show the assembly/disassembly of mammalian SWI/SNF complexes is dynamically regulated by a lysine methylation mechanism involving the demethylase LSD1 and the methyl-lysine reader L3MBTL3, which recognizes SET7-methylated lysines in SMARCC1 and SMARCC2 to target them for CRL4 ubiquitin ligase-mediated proteolysis.
- Pengfei Guo
- , Nam Hoang
- & Hui Zhang
-
Article
| Open Access Ubiquitin proteolysis of a CDK-related kinase regulates titan cell formation and virulence in the fungal pathogen Cryptococcus neoformans
Ubiquitin proteolysis of a CDK-related kinase regulates titan cell formation and virulence in the fungal pathogen Cryptococcus neoformans
The pathogenic yeast Cryptococcus neoformans forms large, so-called ‘titan cells’ during infection. Here, Cao et al. show that a ubiquitin ligase inhibits this process by targeting for degradation a CDK-related kinase that stimulates titan cell formation.
- Chengjun Cao
- , Keyi Wang
- & Chaoyang Xue
-
Article
| Open Access Predicting the structural basis of targeted protein degradation by integrating molecular dynamics simulations with structural mass spectrometry
Predicting the structural basis of targeted protein degradation by integrating molecular dynamics simulations with structural mass spectrometry
The formation of ternary degrader-protein complexes is a key step in the targeted degradation of proteins of interest. Here, the authors explore the structure and dynamics of such complexes applying high-performance computer simulations augmented with experimental data.
- Tom Dixon
- , Derek MacPherson
- & Jesus A. Izaguirre
-
Article
| Open Access Cryo-EM structures of Gid12-bound GID E3 reveal steric blockade as a mechanism inhibiting substrate ubiquitylation
Cryo-EM structures of Gid12-bound GID E3 reveal steric blockade as a mechanism inhibiting substrate ubiquitylation
The GID E3 ligase regulates glucose-induced degradation in yeast, and key physiology. This study unveils E3 ligase regulation by reshaping the substrate binding site, blocking substrate access to ubiquitination active sites, and a Cage-like assembly.
- Shuai Qiao
- , Chia-Wei Lee
- & Brenda A. Schulman
-
Article
| Open Access Proteolysis of adaptor protein Mmr1 during budding is necessary for mitochondrial homeostasis in Saccharomyces cerevisiae
Proteolysis of adaptor protein Mmr1 during budding is necessary for mitochondrial homeostasis in Saccharomyces cerevisiae
Mitochondria are transported to daughter cells by myosin via an adaptor protein Mmr1 during budding in yeast. Here they show that the regulated proteolysis of Mmr1 after mitochondria inheritance is required for mitochondrial homeostasis.
- Keisuke Obara
- , Taku Yoshikawa
- & Takumi Kamura
-
Article
| Open Access UBR4/POE facilitates secretory trafficking to maintain circadian clock synchrony
UBR4/POE facilitates secretory trafficking to maintain circadian clock synchrony
Although ubiquitin ligases are known to control clock protein degradation, their other roles in clock neurons are unclear. Here the authors report that UBR4 promotes export of neuropeptides from the Golgi for axonal trafficking, which is important for circadian clock synchrony in mice and flies.
- Sara Hegazi
- , Arthur H. Cheng
- & Hai-Ying Mary Cheng
-
Article
| Open Access Ubiquitin and a charged loop regulate the ubiquitin E3 ligase activity of Ark2C
Ubiquitin and a charged loop regulate the ubiquitin E3 ligase activity of Ark2C
Attachment of ubiquitin to proteins is tightly regulated and controls many signalling pathways. Here, the authors show that addition of ubiquitin by the RING E3 ligases Arkadia and Ark2C is enhanced by ubiquitin and a charged loop that precedes the RING domain.
- Andrej Paluda
- , Adam J. Middleton
- & Catherine L. Day
-
Article
| Open Access Structural basis for the E3 ligase activity enhancement of yeast Nse2 by SUMO-interacting motifs
Structural basis for the E3 ligase activity enhancement of yeast Nse2 by SUMO-interacting motifs
Nse2 is a SUMO E3 ligase component of the Smc5/6 multisubunit complex involved in the DNA repair and chromosome integrity. Here, the structure of the Nse2 in complex with an E2-SUMO thioester mimetic reveals the combined action of two SIM motifs during the E3- dependent conjugation reaction.
- Nathalia Varejão
- , Jara Lascorz
- & David Reverter
-
Article
| Open Access The NUCKS1-SKP2-p21/p27 axis controls S phase entry
The NUCKS1-SKP2-p21/p27 axis controls S phase entry
Entry into S phase of the cell cycle is regulated positively by mitogens and negatively by DNA damage; however, how balance of these signals is achieved is not well known. Here the authors show that the NUCKS1-SKP2- p21/p27 axis integrates this information, where the NUCKS1 transcription factor affects levels of p21/p27 to readout the mitogen:DNA damage balance and regulate S phase entry decision.
- Samuel Hume
- , Claudia P. Grou
- & Grigory L. Dianov
-
Article
| Open Access G3BP1 inhibits Cul3SPOP to amplify AR signaling and promote prostate cancer
G3BP1 inhibits Cul3SPOP to amplify AR signaling and promote prostate cancer
SPOP functions as a tumour suppressor in prostate cancer but how the protein is regulated is unclear. Here, the authors identify G3BP1 as a competitive inhibitor of SPOP and show that G3BP1-SPOP axis activates androgen signalling to drive tumorigenesis.
- Chandrani Mukhopadhyay
- , Chenyi Yang
- & Pengbo Zhou
-
Article
| Open Access Genetic fusions favor tumorigenesis through degron loss in oncogenes
Genetic fusions favor tumorigenesis through degron loss in oncogenes
The impact of genetic fusions on degrons, which are motifs for ubiquitin-mediated protein degradation, has not been fully explored. Here, the authors analyse fusion genes affecting degrons in pan-cancer genomics data, validate their functional impact and find enrichment for both internal and C-terminal degron losses.
- Jing Liu
- , Collin Tokheim
- & Wenyi Wei
-
Article
| Open Access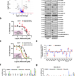 Inhibition of CK1ε potentiates the therapeutic efficacy of CDK4/6 inhibitor in breast cancer
Inhibition of CK1ε potentiates the therapeutic efficacy of CDK4/6 inhibitor in breast cancer
Acquisition of CDK4/6i resistance represents a major clinical challenge. Here, the authors report that inhibition of CK1ε can prevent acquisition of CDK4/6i resistance, potentiating the therapeutic efficacy of CDK4/6i in human breast cancer.
- Fabin Dang
- , Li Nie
- & Wenyi Wei
-
Article
| Open Access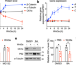 The E3 ubiquitin ligase component, Cereblon, is an evolutionarily conserved regulator of Wnt signaling
The E3 ubiquitin ligase component, Cereblon, is an evolutionarily conserved regulator of Wnt signaling
Cereblon (CRBN) is an E3 ubiquitin ligase substrate receptor that is involved in cancer cell death, although its regulation is poorly understood. Here, the authors show that Wnt ligand increases CRBN-dependent protein degradation and demonstrate CRBN’s importance in physiological Wnt signaling.
- Chen Shen
- , Anmada Nayak
- & David J. Robbins
-
Article
| Open Access MARCH8 inhibits influenza A virus infection by targeting viral M2 protein for ubiquitination-dependent degradation in lysosomes
MARCH8 inhibits influenza A virus infection by targeting viral M2 protein for ubiquitination-dependent degradation in lysosomes
The membrane-associated RING-CH (MARCH) proteins are E3 ligases regulating stability of plasma membrane (PM) proteins. Here, Liu et al. show that MARCH8 suppresses Influenza A virus infection in vitro and in vivo through redirecting M2 protein from the PM to lysosomes for degradation.
- Xiaoman Liu
- , Fengwen Xu
- & Fei Guo
-
Article
| Open Access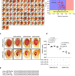 UBE4B, a microRNA-9 target gene, promotes autophagy-mediated Tau degradation
UBE4B, a microRNA-9 target gene, promotes autophagy-mediated Tau degradation
Hyperphosphorylated Tau accumulation promotes neurodegeneration in Alzheimer’s disease. Here, the authors screen a miRNA library in Drosophila and identify a conserved ubiquitin ligase that directs Tau for autophagic degradation, uncovering a potential target to treat Tau-mediated neurodegeneration.
- Manivannan Subramanian
- , Seung Jae Hyeon
- & Kweon Yu
-
Article
| Open Access MAEA is an E3 ubiquitin ligase promoting autophagy and maintenance of haematopoietic stem cells
MAEA is an E3 ubiquitin ligase promoting autophagy and maintenance of haematopoietic stem cells
Haematopoietic stem cells (HSCs) are metabolically quiescent, with balanced myeloid and lymphoid potential. Here the authors show that MAEA is required in HSCs for ubiquitination and downregulation of surface cytokine receptors via autophagy; MAEA loss leads to impaired HSC quiescence and a myeloproliferative disorder.
- Qiaozhi Wei
- , Sandra Pinho
- & Paul S. Frenette
-
Article
| Open Access IP6-assisted CSN-COP1 competition regulates a CRL4-ETV5 proteolytic checkpoint to safeguard glucose-induced insulin secretion
IP6-assisted CSN-COP1 competition regulates a CRL4-ETV5 proteolytic checkpoint to safeguard glucose-induced insulin secretion
Mediators of insulin signalling are targets of cullin-RING ubiquitin ligases (CRL) that mediate protein degradation, but the role of protein degradation in insulin signalling is incompletely understood. Here, the authors identified a glucose-responsive CRL4-COP1-ETV5 proteolytic axis that promotes insulin secretion, and is inhibited under hypoglycemia.
- Hong Lin
- , Yuan Yan
- & Feng Rao
-
Article
| Open Access Proteasomal degradation of the tumour suppressor FBW7 requires branched ubiquitylation by TRIP12
Proteasomal degradation of the tumour suppressor FBW7 requires branched ubiquitylation by TRIP12
The tumor suppressor FBW7 is a substrate adaptor for the E3 ubiquitin ligase complex SKP1-CUL1-F-box (SCF) and itself a target for ubiquitylation. Here, the authors show that TRIP12 mediates branched K11-linked ubiquitylation of FBW7, to regulate its stability and thus abundance of a subset of SCFFBW7 substrates.
- Omar M. Khan
- , Jorge Almagro
- & Axel Behrens
-
Article
| Open Access Antagonistic control of myofiber size and muscle protein quality control by the ubiquitin ligase UBR4 during aging
Antagonistic control of myofiber size and muscle protein quality control by the ubiquitin ligase UBR4 during aging
Sarcopenia is the age-associated functional decline and atrophy of muscle fibers, and it has been proposed that it might be counteracted by inducing myofiber hypertrophy. Here, the authors show that expression levels of the ubiquitin ligase UBR4 are increased with ageing, and that whilst its genetic ablation rescues muscle atrophy, it is also associated with reduced protein quality and impaired force production in Drosophila and mouse models.
- Liam C. Hunt
- , Bronwen Schadeberg
- & Fabio Demontis
-
Article
| Open Access RETRACTED ARTICLE:The Arabidopsis NOT4A E3 ligase promotes PGR3 expression and regulates chloroplast translation
RETRACTED ARTICLE:The Arabidopsis NOT4A E3 ligase promotes PGR3 expression and regulates chloroplast translation
NOT4 proteins associate with ribosomes and are required for co-translational quality control in yeast and animals. Here, Bailey et al. show that Arabidopsis NOT4A positively regulates the expression of the nuclear encoded pentatricopeptide repeat (PPR) protein PGR3 and is required for ribosome biogenesis and mRNA translation in the chloroplast.
- Mark Bailey
- , Aiste Ivanauskaite
- & Daniel J. Gibbs
-
Article
| Open Access Structural basis for substrate recognition and chemical inhibition of oncogenic MAGE ubiquitin ligases
Structural basis for substrate recognition and chemical inhibition of oncogenic MAGE ubiquitin ligases
Testis-restricted melanoma antigen (MAGE) proteins function as substrate adapters for E3 ubiquitin ligases. Biochemical and structural analyses of MAGE-A11 provide insight into the substrate binding mode of MAGE proteins and enable discovery of potent, cytotoxic inhibitors of MAGE-A11:substrate interaction.
- Seung Wook Yang
- , Xin Huang
- & Patrick Ryan Potts
-
Article
| Open Access A phospho-switch controls RNF43-mediated degradation of Wnt receptors to suppress tumorigenesis
A phospho-switch controls RNF43-mediated degradation of Wnt receptors to suppress tumorigenesis
RNF43 is frequently mutated in cancers and negatively regulates Wnt signalling. Here, the authors report that RNF43 phosphorylation at a serine triplet is required for the negative regulation of Wnt signalling and that the phosphorylation of RNF43 suppresses cancer-associated oncogenic RNF43 mutants.
- Tadasuke Tsukiyama
- , Juqi Zou
- & Shigetsugu Hatakeyama
-
Article
| Open Access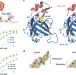 Structural bases of IMiD selectivity that emerges by 5-hydroxythalidomide
Structural bases of IMiD selectivity that emerges by 5-hydroxythalidomide
5-hydroxythalidomide is a primary thalidomide metabolite generated by the cytochrome P450 isozymes. The reported data, including crystal structure of the 5-hydroxythalidomide-mediated complex of CRBN with SALL4, elucidate how additional hydroxy group of the metabolite enhances the interaction of CRBN with the neosubstrate SALL4.
- Hirotake Furihata
- , Satoshi Yamanaka
- & Takuya Miyakawa
-
Article
| Open Access Structural basis for DNA damage-induced phosphoregulation of MDM2 RING domain
Structural basis for DNA damage-induced phosphoregulation of MDM2 RING domain
p53 is an important tumor suppressor protein which is regulated by the E3 ubiquitin ligase MDM2. Here the authors reveal that DNA damage-induced Ser429 phosphorylation of MDM2 serve to boost the activity of MDM2 homodimer by stabilizing the active E2–ubiquitin complex and promote its self-destruction to enable rapid p53 stabilization.
- Helge M. Magnussen
- , Syed F. Ahmed
- & Danny T. Huang
-
Article
| Open Access SKP2 attenuates autophagy through Beclin1-ubiquitination and its inhibition reduces MERS-Coronavirus infection
SKP2 attenuates autophagy through Beclin1-ubiquitination and its inhibition reduces MERS-Coronavirus infection
Here, Gassen et al. show that S-phase kinase-associated protein 2 (SKP2) is responsible for lysine-48-linked poly-ubiquitination of beclin 1, resulting in its proteasomal degradation, and that inhibition of SKP2 enhances autophagy and reduces replication of MERS coronavirus.
- Nils C. Gassen
- , Daniela Niemeyer
- & Theo Rein
-
Article
| Open Access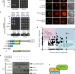 GIGANTEA recruits the UBP12 and UBP13 deubiquitylases to regulate accumulation of the ZTL photoreceptor complex
GIGANTEA recruits the UBP12 and UBP13 deubiquitylases to regulate accumulation of the ZTL photoreceptor complex
The daily accumulation of the ZEITLUPE (ZTL) photoreceptor/E3 ubiquitin ligase relies on a light-dependent interaction with GIGANTEA (GI). Here the authors show that GI recruits two deubiquitylases to help stabilize the ZTL-GI complex during the day and likely counterbalance the activity of ZTL.
- Chin-Mei Lee
- , Man-Wah Li
- & Joshua M. Gendron
-
Article
| Open Access WWP2 regulates pathological cardiac fibrosis by modulating SMAD2 signaling
WWP2 regulates pathological cardiac fibrosis by modulating SMAD2 signaling
Pathological cardiac fibrosis is a hallmark of diseases leading to heart failure. Here, the authors used systems genetics to identify a pro-fibrotic gene network regulated by WWP2, a E3 ubiquitin ligase, which orchestrates the nucleocytoplasmic shuttling and transcriptional activity of SMAD2 in the diseased heart.
- Huimei Chen
- , Aida Moreno-Moral
- & Enrico Petretto
-
Article
| Open Access Degron-tagged reporters probe membrane topology and enable the specific labelling of membrane-wrapped structures
Degron-tagged reporters probe membrane topology and enable the specific labelling of membrane-wrapped structures
Visualising certain organelles and their dynamics is challenging in living cells. Here the authors co-opt selective degradation to label membrane-bound compartments in worm embryos and mammalian cells, revealing membrane topology during cell division.
- Katharina B. Beer
- , Gholamreza Fazeli
- & Ann M. Wehman
-
Article
| Open Access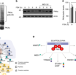 Feedback inhibition of cAMP effector signaling by a chaperone-assisted ubiquitin system
Feedback inhibition of cAMP effector signaling by a chaperone-assisted ubiquitin system
How intracellular cAMP activate PKA is well-characterized, but PKA inactivation remains poorly understood. Here, Rinaldi et al. show that CHIP/HSP70 ubiquitinates the catalytic subunit of PKA, with implications for the human disease spinocerebellar ataxia 16, as patients often have CHIP mutations.
- Laura Rinaldi
- , Rossella Delle Donne
- & Antonio Feliciello
-
Article
| Open Access Vps11 and Vps18 of Vps-C membrane traffic complexes are E3 ubiquitin ligases and fine-tune signalling
Vps11 and Vps18 of Vps-C membrane traffic complexes are E3 ubiquitin ligases and fine-tune signalling
Endosomal fusion depends on the HOPS and CORVET complexes but whether and how their subunits modulate signal transduction is not fully understood. Here, the authors show that the HOPS/CORVET subunits Vps11 and Vps18 are E3 ubiquitin ligases that are involved in the regulation of ERα signalling.
- Gregory Segala
- , Marcela A. Bennesch
- & Didier Picard
-
Article
| Open Access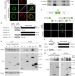 Gigaxonin E3 ligase governs ATG16L1 turnover to control autophagosome production
Gigaxonin E3 ligase governs ATG16L1 turnover to control autophagosome production
Membrane elongation is fundamental to autophagy and is controlled by an ubiquitin-conjugating cascade orchestrated by ATG16L1. Here, the authors identify that the E3 ligase Gigaxonin regulates autophagosome formation by controlling ATG16L1 turnover.
- Aurora Scrivo
- , Patrice Codogno
- & Pascale Bomont
-
Article
| Open Access ER-associated ubiquitin ligase HRD1 programs liver metabolism by targeting multiple metabolic enzymes
ER-associated ubiquitin ligase HRD1 programs liver metabolism by targeting multiple metabolic enzymes
HRD1 is an E3 ligase known to play a role in targeting degradation of misfolded proteins in the ER. Here the authors show that HRD1 interacts with metabolic enzymes and its liver specific deficiency results in lower body weight, blood glucose and plasma lipids during high fat diet in mice.
- Juncheng Wei
- , Yanzhi Yuan
- & Deyu Fang
-
Article
| Open Access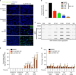 The ubiquitin ligase UBR5 suppresses proteostasis collapse in pluripotent stem cells from Huntington’s disease patients
The ubiquitin ligase UBR5 suppresses proteostasis collapse in pluripotent stem cells from Huntington’s disease patients
Induced pluripotent stem cells (iPSCs) suppress the aggregation of Huntington’s disease (HD) polyQ-expanded huntingtin (HTT). Here the authors show that proteasome activity determines the levels of mutant HTT in HD-iPSCs and find that UBR5 is a modulator of super-vigilant proteostasis of iPSCs.
- Seda Koyuncu
- , Isabel Saez
- & David Vilchez
-
Article
| Open Access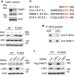 Methylated DNMT1 and E2F1 are targeted for proteolysis by L3MBTL3 and CRL4DCAF5 ubiquitin ligase
Methylated DNMT1 and E2F1 are targeted for proteolysis by L3MBTL3 and CRL4DCAF5 ubiquitin ligase
Lysine methylation is increasingly being implicated in the modification of non-histone proteins. Here the authors find that the methylation of DNMT1 and E2F1 are recognized by the protein L3MBTL3 and the ubiquitin E3 ligase CRL4DCAF5, which cooperatively target these methylated proteins for ubiquitin-dependent proteolysis.
- Feng Leng
- , Jiekai Yu
- & Hui Zhang
-
Article
| Open Access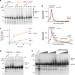 UFD-2 is an adaptor-assisted E3 ligase targeting unfolded proteins
UFD-2 is an adaptor-assisted E3 ligase targeting unfolded proteins
The U-box ubiquitin ligase UFD-2 is one of the most abundant components of the ubiquitin proteasome system in muscle cells. Here the authors perform in vitro and in vivo experiments and show that UFD-2 has E3 ligase activity and that it ubiquitinates unfolded myosin using the C. elegans myosin chaperone UNC-45 as an adaptor protein.
- Doris Hellerschmied
- , Max Roessler
- & Tim Clausen
-
Article
| Open Access RNF8/UBC13 ubiquitin signaling suppresses synapse formation in the mammalian brain
RNF8/UBC13 ubiquitin signaling suppresses synapse formation in the mammalian brain
Ubiquitin ligases play critical roles in neuronal connectivity in the brain. Here, Valnegri and colleagues show that ubiquitin ligase RNF8 and ubiquitin-conjugating enzyme UBC13 regulate synapse number in cerebellar granule neurons and rodent cerebellar learning.
- Pamela Valnegri
- , Ju Huang
- & Azad Bonni
-
Article
| Open Access An integrated bioinformatics platform for investigating the human E3 ubiquitin ligase-substrate interaction network
An integrated bioinformatics platform for investigating the human E3 ubiquitin ligase-substrate interaction network
Protein stability modulation by E3 ubiquitin ligases is an important layer of functional regulation, but screening for E3 ligase-substrate interactions is time-consuming and costly. Here, the authors take an in silico naïve Bayesian classifier approach to integrate multiple lines of evidence for E3-substrate prediction, enabling prediction of the proteome-wide human E3 ligase interaction network.
- Yang Li
- , Ping Xie
- & Fuchu He

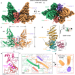 Structural insights into the functional mechanism of the ubiquitin ligase E6AP
Structural insights into the functional mechanism of the ubiquitin ligase E6AP