Abstract
WD-repeat proteins are regulatory beta-propeller platforms that enable the assembly of multiprotein complexes. Here, we report the functional and bioinformatic analysis of human WD-repeat protein Interacting with PhosphoInosides (WIPI)-1α (WIPI49/Atg18), a member of a novel WD-repeat protein family with autophagic capacity in Saccharomyces cerevisiae and Caenorhabditis elegans, recently identified as phospholipid-binding effectors. Our phylogenetic analysis divides the WIPI protein family into two paralogous groups that fold into 7-bladed beta-propellers. Structural modeling identified two evolutionary conserved interaction sites in WIPI propellers, one of which may bind phospholipids. Human WIPI-1α has LXXLL signature motifs for nuclear receptor interactions and binds androgen and estrogen receptors in vitro. Strikingly, human WIPI genes were found aberrantly expressed in a variety of matched tumor tissues including kidney, pancreatic and skin cancer. We found that endogenous hWIPI-1 protein colocalizes in part with the autophagosomal marker LC3 at punctate cytoplasmic structures in human melanoma cells. In addition, hWIPI-1 accumulated in large vesicular and cup-shaped structures in the cytoplasm when autophagy was induced by amino-acid deprivation. These cytoplasmic formations were blocked by wortmannin, a classic inhibitor of PI-3 kinase-mediated autophagy. Our data suggest that WIPI proteins share an evolutionary conserved function in autophagy and that autophagic capacity may be compromised in human cancers.
Similar content being viewed by others
Introduction
Beta-propellers are a diverse group of proteins built of between four and eight ‘blades’ packed in a circular array (Neer et al., 1994; Neer and Smith, 1996; Yu et al., 2000). Each blade consists of a four-stranded antiparallel β-sheet and the ring structure is often stabilized by the circular permutation of one or more β-strands from the C- to the N-terminus, such that the termini are now within one repeat rather than between them (Paoli, 2001a). WD-repeat proteins comprise a superfamily of regulatory proteins within the beta-propeller fold. The WD-repeat was originally defined as a core unit of approximately 40 amino acids, ending with the residues, tryptophan and aspartic acid, hence ‘WD’ (Fong et al., 1986; Garcia-Higuera et al., 1996; Garcia-Higuera et al., 1998; Smith et al., 1999). Despite the strong conservation of structural features (Paoli, 2001b), WD-repeats display a high degree of sequence diversity.
The common function of all WD-repeat proteins is to regulate the assembly of multiprotein complexes by presenting a stable attachment platform for simultaneous and reversible protein–protein interactions (van Nocker and Ludwig, 2003). Based on this property, WD-repeat proteins are key components of many essential biological functions including cell cycle control, apoptosis, signal transduction pathways, RNA metabolism, chromatin assembly and vesicular trafficking (Li and Roberts, 2001). The importance of WD-repeat proteins is further highlighted by their association with inheritable human diseases (Li and Roberts, 2001) and tumor phenotypes (Chernova et al., 2001; Honore et al., 2002). WD-repeat proteins have also been linked to the nonapoptotic cell death pathway because mouse Apg16L, a novel factor harboring a WD-repeat domain at its C-terminus, participates as a scaffold in an 800 kDa complex that functions in mammalian autophagy (Mizushima et al., 2003).
Autophagy (macroautophagy) is the bulk degradation of proteins and organelles (Kim and Klionsky, 2000; Klionsky and Emr, 2000) and is significantly associated with cancer, infection, neurodegenerative diseases and cardiomyopathies (Mizushima et al., 2002; Klionsky, 2004; Tanida et al., 2004). Autophagy is mediated by unique organelles termed autophagosomes that are formed from isolation membranes (also called phagophores) enclosing undigested cytoplasmic material. In yeast, autophagy-related factors accumulate at the autophagosome formation organizing center, the preautophagosomal structures (PAS). The PAS are sensitive to PI-3 kinase inhibitors. To date, mammalian PAS-like structures have not been described (Gozuacik and Kimchi, 2004).
From an unrelated yeast-based screen (Waddell et al., 2001), we have cloned a novel human WD-repeat gene, termed WIPI-1α, and we here present its comprehensive bioinformatic analysis and expression profile in matched human tumor samples. We provide the first evidence of an involvement of human WIPI-1 in mammalian autophagy. Endogenous hWIPI-1 protein colocalizes in part with the recognized autophagosomal marker LC3. Starvation-induced autophagy resulted in a striking accumulation of human WIPI-1 at large cup-shaped, vesicular and lasso-like structures in the cytoplasm. Inhibition of PI-3 kinase activity with wortmannin abolished the formation of these hWIPI-1-containing structures.
Based on our observations in the mammalian system, and reported studies analysing yeast and nematode homologs (Melendez et al., 2003; Stromhaug et al., 2004), we propose that WIPI proteins share an ancestral function in autophagy. Further, we predict the autophagic capacity of WIPI proteins to be defective in human cancers because we found WIPI genes to be aberrantly expressed in a variety of tumor tissues.
Results
Our phylogenetic analysis revealed that our screen isolate represents a human homolog of the yeast Atg18 gene (Barth et al., 2001; Georgakopoulos et al., 2001; Guan et al., 2001), shown to function in autophagy in Saccharomyces cerevisiae (Stromhaug et al., 2004) and Caenorhabditis elegans (Melendez et al., 2003). Further, members of this novel WD-repeat family in yeast (Atg18, Atg21) and humans (WIPI49) have been described as phospholipid-binding effectors (Dove et al., 2004; Jeffries et al., 2004; Stromhaug et al., 2004). To reflect this characteristic, we propose WIPI as the mammalian nomenclature according to human WIPI49 (Jeffries et al., 2004).
Isolation of human WIPI gene family members
We originally isolated human WIPI-1 as a 5′-truncated cDNA fragment from a human liver cDNA library screen (Waddell et al., 2001). We subsequently isolated the missing 5′-fragment by RT–PCR from normal human testis mRNA and generated a fused full-length human WIPI-1α cDNA (AY691424). We identified several additional human WIPI cDNAs by BLAST searching the human genome sequence database and we isolated these species by RT–PCR cloning from normal human testis mRNA (data not shown). Analysis of our isolated cDNAs and NCBI database entries defines four human WIPI genes, two of which give rise to alternative splice variants (Table 1). Human WIPI-1α was our initial isolate and WIPI-1β is an N-terminal variant, which was recently described as WIPI49 (Jeffries et al., 2004). The closest human homologs of WIPI-1 are represented by splice variants of the WIPI-2 gene, whose expression (hWIPI-2β, -δ) we identified by RT–PCR (data not shown). Human WIPI-1 is encoded by the antisense strand of the sulfatase.3 gene, and hWIPI-3 is encoded by the antisense strand of the interleukin enhancer-binding factor 1-like family member (ILF1-like) gene.
To analyse the expression of hWIPI genes in normal human tissues, we generated PCR probes that specifically corresponded to unique 3′-sequences of hWIPI ORFs. Hence, hybridization results (Figures 1 and 5) for hWIPI-1 represent the expression of both the N-terminal α and β variants, and results for hWIPI-2 represent the expression of all the splice variants α–δ (see Table 1). All the human WIPI genes are ubiquitously expressed (Figure 1). Levels are particularly high in skeletal and heart muscle, and in testis. In addition, hWIPI-1 is highly expressed in normal pancreas and placenta (Figure 1a). Brain expression of hWIPI-1 is highest in the adult cerebellum (Figure 1b). Northern analysis also indicates the presence of an as yet uncharacterized but ubiquitous hWIPI-1 splice variant of approximately 7.0 kb, indicated by an arrowhead in Figure 1a.
Human WIPI genes are highly expressed in the skeletal muscle, the heart and the testis. Human WIPI-1 expression in normal human tissues analysed in Northern blots (a) or RNA dot blots (b; hybridization results on top, RNA origin at the bottom). The putative hWIPI-1 splice variant of approximately 7 kb is indicated by an arrowhead. The mRNA expression of hWIPI-2, -3, -4 is shown by Northern blot analysis (c), the β-actin control is also provided (the human heart and skeletal muscle contain two forms of β-actin, 2.0 and 1.8 kb)
Human WIPI genes are aberrantly expressed in a variety of matched tumor samples. The origin of the matched tumor samples is shown on the bottom right-hand side where the filter was hybridized with a 32P-labeled ubiquitin probe to control for template loading. The normal tissue sample from each patient is on the left-hand side and the corresponding tumor tissue is on the right-hand side. Filters were hybridized with 32P-labelled probes specific for each of the four human WIPI members
Human WIPI-1α is a member of a highly conserved WD-repeat protein family that falls into two paralogous groups
A bioinformatic analysis of hWIPI-1α (see Materials and methods) showed that WIPI proteins form a large family with conserved residues that set them aside from other WD-repeat proteins. In a cluster analysis (Figure 2, left panel), WIPI sequences fell into two large paralogs groups, one containing human WIPI-1 and -2, the other human WIPI-3 and -4. Both groups have representatives throughout the crown eukaryotes (plants, animals and fungi); in addition, the WIPI-3/4 group is also present in some protozoans. Phylogenetic analysis showed that, in addition to the deep paralogy at the root of the family, vertebrates have undergone an additional duplication in each of the two groups (Figure 2, right panel). Phylogenetic analysis of individual blades further showed that the prototypical WIPI protein contained fully differentiated blades, suggesting that the phospholipid binding function is likely to be an ancestral property (data not shown).
The WIPI protein family. The left panel shows a clustering analysis of the WIPI family in relation to WD-repeat proteins; the location of the seed sequence (hWIPI-1α) is marked by an open circle. Neighbor-joining phylogeny of the WIPI family is shown in right panel; the two subfamilies identified in the clustering analysis are marked with parentheses
WIPI proteins as 7-bladed beta-propellers
In searches against databases of domain profiles, such as Pfam and SMART (Schultz et al., 1998), at least two regions of human WIPI proteins matched the WD-repeat consensus (data not shown). Further bioinformatic analyses (see Materials and methods) revealed the presence of seven WD-like repeats, starting with strand 1a and ending with strand 7d, and thus not displaying any permutation (Figure 3, upper panel). The 12 residues preceding and 85 residues following the predicted propeller domain in hWIPI-1α appear to be natively unfolded. In addition, the 37-residue insertion between strands 6c and 6d also appears to be essentially unstructured. A homology model of the propeller domain of hWIPI-1α, based on the prototypical WD-repeat proteins TUP1 and G-protein beta subunit, illustrates the general fold of the WIPI family (Figure 3, lower panel) and provided a scaffold for analysing the location of conserved and invariant residues (colored red and magenta, respectively, in Figure 3). The analysis showed that conserved residues cluster largely on one face of the propeller, between blades 1 and 3, and invariant residues on the other face, between blades 4 and 6. The two clusters define two potential binding sites of WIPI proteins, which seem connected through the central pore by a string of conserved polar residues (Figure 3, bottom right). The cluster of invariant residues contains two consecutive arginines, whose mutation in the yeast WIPI protein Svp1p has been shown to severely affect the interaction with phospholipids (Dove et al., 2004). In contrast, mutation of two consecutive arginines outside this cluster had little effect on phospholipid binding. We therefore propose that this is the phospholipid binding site and deduce from its conservation that it represents the common feature of all WIPI proteins.
WIPI proteins as 7-bladed beta-propellers. The assignment of the seven propeller blades in hWIPI-1α, as well as of the four β-strands (a, b c, and d) forming one blade, is shown at the top. Predicted β-strands are underlined. The N- and C-terminal extensions and the large insertion between strands c and d of blade 6 are predicted to be essentially unstructured. The terminal extensions were omitted from the model. The structure of the insert in the figure is arbitrary and shown purely to illustrate its location and size. Residues in conserved hydrophobic positions of the WIPI family are colored blue; conserved polar residues are colored red, and invariant polar residues are colored magenta. Below are shown top and side views of an hWIPI-1α model, colored on the left from blue to yellow for the N- to C-terminal succession of blades and highlighting on the right the location of conserved polar residues. Polar residues are colored as in the alignment above, and two invariant arginines, which have been shown by mutagenesis to be involved in phospholipid binding, are labeled
GST-hWIPI-1α interacts with nuclear steroid hormone receptors
The primary amino-acid sequence of WIPI-1 contains multiple motifs for putative protein–protein interactions (ELM database, Puntervoll et al., 2003), such as the LXXLL signature motif for nuclear receptor interaction. To test for an interaction with nuclear receptors, we conducted GST pull-down experiments with bacterially expressed GST-hWIPI-1α and GST alone as a control. As input samples, we used a set of in vitro-translated nuclear hormone receptors. GST-hWIPI-1α can physically interact with the androgen (AR) and the estrogen receptor (ER) in vitro (Figure 4). Interestingly, the interaction of GST-hWIPI-1α with nuclear receptors was hormone independent because binding to the AR occurred in the presence and absence of 5α-androstan, and binding to ERs (ERα and ERβ) was independent of β-estradiol (Figure 4). Similar results were obtained with the retinoic acid receptors RAR and RXR (data not shown).
GST-hWIPI-1α interacts in vitro with nuclear hormone receptors. GST pull-down experiments were performed with GST alone or GST-hWIPI-1α using in vitro-translated AR, ERα and ERβ as input samples in the presence or absence of 5α-androstan or β-estradiol. The Coomassie blue-stained gel is shown on the bottom and demonstrates that an excess amount of GST control protein was used in all GST pull-down experiments when compared to GST-hWIPI-1α protein. The asterisk indicates an unspecific protein that copurified with GST and GST-hWIPI1α
Human WIPI genes are aberrantly expressed in human cancers
Human WIPI-1 and WIPI-3 localize to regions 17q24.3 and 17q25.3, a segment on chromosome 17q that is allelically imbalanced in breast, ovarian and prostate cancer (Gozuacik and Kimchi, 2004). We therefore examined the mRNA expression profile of the human WIPI genes in several matched tumor tissues (Figure 5). Strikingly, expression of hWIPI-2 and hWIPI-4 was found downregulated in 100% of the matched pancreatic cancer samples (seven patients) and in 50% of kidney tumors (10 patients). This phenotype was also observed for hWIPI-1 (40%, 10 patients) and hWIPI-2 (30%, 10 patients), although at a lesser penetrance. Interestingly, hWIPI-1 expression was upregulated in five samples of 10 tested skin cancer patients and in four out of 10 samples from cervical carcinoma patients. Human WIPI-2 mRNA expression was upregulated in seven out of 10 uterine cancer patients. In addition, hWIPI-3 expression appears slightly elevated in six out of 10 ovarian cancer patients. We used a 32P-labeled ubiquitin cDNA probe to control for equal loading (Figure 5). The mRNA expression profiles in Figure 5 demonstrated that hWIPI-1 expression was upregulated in 50% of skin cancer patients and consistent with the fact that hWIPI-1 was highly expressed in the human melanoma cell line G361 and nondetectable in HeLa cells (Figure 5). These results lead us to test whether hWIPI-1 was misexpressed in a variety of human melanoma cell lines. We confirmed that hWIPI-1 is highly expressed in G361 cells and demonstrated that hWIPI-1 levels are also high in Sk-mel-28, Sk-mel-13, WM852 and WM451 cells (Figure 6a).
Analysis of WIPI-1 mRNA expression in human melanoma cell lines and WIPI-1 protein expression in G361 and HeLa cells. Northern blot analysis of total RNA extracted from human tumor cell lines, hybridized with a 32P-labelled probe specific for hWIPI-1 (a). Western blotting of total cell extracts from G361 and HeLa cells with a polyclonal antiserum recognizing the C-terminus of hWIPI-1 (b). As indicated, G361 or HeLa cells were transiently transfected with a GF-P or GFP-hWIPI-1α-expressing plasmid to further demonstrate specificity of the raised antiserum used in the present study (b)
We generated a rabbit polyclonal antiserum that detects the C-terminus of human WIPI-1. This antiserum detected a high level of hWIPI-1 in G361, but hWIPI-1 was not detectable in HeLa cells (Figure 6b). Our C-terminal antiserum is hWIPI specific because we only detected a single band at the expected size of 49 kDa in G361 total cell extracts (Figure 6b). We also transiently transfected G361 and HeLa cells with a GFP-expressing vector or with a construct that encodes an N-terminal GFP-hWIPI-1α fusion protein. Again, we detected a 49 kDa protein band representing endogenous hWIPI-1 in GFP-transfected G361 cells but not in HeLa cells (Figure 6b). As expected, an additional 76 kDa protein band was detected at the predicted size of GFP-hWIPI-1α and a 49 kDa band was detected that results from an internal transcription start within the GFP-hWIPI-1α open reading frame (marked with an asterisk in HeLa cells).
Human WIPI-1α accumulates at discrete subcellular structures upon amino-acid deprivation
The WIPI homologs in S. cerevisiae and C. elegans were shown to be functionally involved in autophagy. We therefore asked whether human WIPI-1 represents a mammalian autophagy-linked protein and studied the intracellular localization of endogenous hWIPI-1 protein in G361 cells by confocal microscopy. In dividing G361 cells, we observed that hWIPI-1 localizes to vesicle-like, punctate structures in the cytoplasm (Figure 7a). This localization dramatically changed when we starved G361 cells in amino-acid-deficient medium for 3 h (Figure 7b, e–l), a procedure widely used to induce autophagy (Mitchener et al., 1976; Munafo and Colombo, 2001). Upon starvation-induced autophagy, hWIPI-1 accumulates at subcellular structures in the cytoplasm: enlarged vesicular (Figure 7f) and lasso-like (Figure 7e and l) structures, and large cup-shaped structures predominantly around the nucleus (Figure 7f, h–k). The PI-3 kinase inhibitor wortmannin inhibits the formation of autophagosomes (Blommaart et al., 1997; Petiot et al., 2000), so we tested whether the starvation-induced hWIPI-1 subcellular structures were sensitive to wortmannin administration. Strikingly, wortmannin administration to G361 cells that were incubated for 3 h in serum- and amino-acid-free medium inhibited the formation of the large WIPI-positive subcellular structures (Figure 7c). Treatment of cells kept in complete culture medium with wortmannin alone resulted in a more diffuse staining of endogenous hWIPI-1; the punctate WIPI-1 pattern was reduced in size and complexity (Figure 7d).
Starvation-induced autophagy triggers a cytoplasmic formation of hWIPI-1 subcellular structures. Fluorescent images of endogenous hWIPI-1 protein in G361 cells detected with a C-terminal WIPI-1 antiserum and Alexa 488 anti-rabbit IgG (green) in complete medium (a), amino-acid-deficient medium (b), amino-acid-deficient medium in the presence of 200 nM wortmannin (c) or complete medium in the presence of wortmannin (d). The variety of WIPI-1 positive subcellular structures that accumulated upon the induction of autophagy by amino-acid deprivation for 3 h are shown in (e–l). Cell nuclei were stained with TO-PRO-3 (blue). Bars 10 μm
Human WIPI-1 colocalizes with the autophagic marker LC3
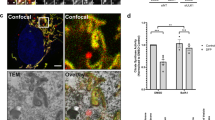
We asked whether hWIPI-1 colocalized with the recognized autophagosomal marker LC3 (Kabeya et al., 2000; Mizushima et al., 2003, 2004). LC3 is present in two forms that show distinct subcellular localization, the diffuse staining of GFP-LC3 represents cytosolic GFP-LC3I and the punctate structures membrane-bound GFP-LC3II, localized at isolation membranes, autophagosomes and some autolysosomes (Kabeya et al., 2000). We transiently expressed GFP-LC3 in G361 cells and observed that GFP-LC3 was distributed throughout the cell as well as present at punctate and large vesicular structures in the cytoplasm under nonstarving conditions (data not shown). Human WIPI-1 colocalized in part with GFP-LC3 at these vesicle-like, punctate structures in dividing G361 cells that were not starved (Figure 8, upper panel). Upon the induction of autophagy by amino-acid depletion, we found LC3- and hWIPI-1-positive subcellular structures in close proximity and at discrete loci (Figure 8, middle panel). Again, we observed a partial colocalization at punctate structures upon amino-acid deprivation (Figure 8, lower panel).
Partial colocalization of endogenous hWIPI-1 and transiently expressed GFP-LC3 in G361 cells. Fluorescent images of GFP-LC3-expressing G361 cells (green); endogenous hWIPI-1 protein detected with a C-terminal WIPI-1 antiserum and Alexa 546 anti-rabbit IgG (red) and cell nuclei stained with TO-PRO-3 (blue). Merged images demonstrate partial colocalization (yellow) of hWIPI-1 and GFP-LC3 prior to (DMEM, upper panel) and after starvation-induced autophagy (EBSS, lower panel). Under starvation conditions (EBSS), hWIPI-1 and GFP-LC3 subcellular structures were also found in close proximity and at discrete loci (middle panel). Bars 10 μm
Discussion
Our bioinformatic analyses demonstrate an ancient origin of the WIPI protein family containing two deeply paralogous groups; these formed during the early evolution of crown eukaryotes and each harbors members from plants, fungi and animals. By homology modeling, we predict that WIPI proteins fold into 7-bladed beta-propellers and that the blades are fully differentiated at the time of evolutionary divergence. Conserved and invariant residues of the WIPI protein family cluster on two opposite sites of the propeller, outlining the location of two conserved binding sites. Recently, it was shown that Atg18/Svp1, WIPI49, Atg21 members of the WIPI protein family are WD-repeat effector modules that bind phospholipids (Dove et al., 2004; Jeffries et al., 2004; Stromhaug et al., 2004). Two critical residues for phospholipid binding (Dove et al., 2004), R227 and R226, are located in one of the clusters (Figure 3), which most likely represents the phospholipid binding site. The other binding site should associate with an as yet unidentified factor, probably by simultaneous binding to phospholipids. In addition, insertions within individual blade units, such as that identified here in blade 6 of hWIPI-1α as well as N- and C-terminal regions may provide the specificity required to bind species-specific factors because these regions show high evolutionary variability and appear ideal for the display of linear motifs. However, we also identified a subset of linear motifs within regions of hWIPI-1α that we predict to participate in folding propeller blade units. Human WIPI-1α has two LXXLL signature motifs for nuclear receptor association and we confirmed the potential of hWIPI-1α to associate with members of class I nuclear steroid hormone receptors (AR, ER, RAR, RXR) by GST pull-down analyses. Although we have yet to determine the physiological significance of this binding potential, it is worth noting that steroid-triggered induction of programmed autophagy has been described in the fruit fly (Lee and Baehrecke, 2001).
The WIPI protein family includes Atg18, the WIPI-1 homolog in S. cerevisiae that was genetically identified as a gene contributing to autophagy (Barth et al., 2001; Georgakopoulos et al., 2001; Guan et al., 2001). A C. elegans ortholog was also recently identified as an autophagic gene essential for dauer development and lifespan extension (Melendez et al., 2003). Furthermore, phospholipid binding is required for vacuole association of Atg18 in S. cerevisiae (Stromhaug et al., 2004), suggesting that phospholipid binding might be important for the autophagic capacity of Atg18 in vivo. We provide evidence here that hWIPI-1 is linked to starvation-induced autophagy in the mammalian system. First, the morphology of endogenous hWIPI-1-labeled structures in dividing G361 cells resembles the classic punctate pattern of autophagy-linked proteins (Kabeya et al., 2000; Simonsen et al., 2004). Second, hWIPI-1 partially colocalizes with the autophagosomal marker GFP-LC3 at those punctate structures. Third, amino-acid deprivation triggered an accumulation of endogenous hWIPI-1 protein to large vesicular and lasso-like or cup-shaped structures that are characteristic for autophagy-linked proteins (Mizushima et al., 2002). Fourth, the starvation-induced hWIPI-1 formation is blocked by wortmannin, a principal inhibitor of PI-3 kinase-induced autophagosome formation (Blommaart et al., 1997; Petiot et al., 2000). We do not know why hWIPI-1 co-localizes with GFP-LC3 at punctated structures prior to the induction of autophagy. It has been shown in detail that the modified form LC3II only binds to autophagic membranes, while LC3I remains diffusely localized (Kabeya et al., 2000). Hence, the punctate GFP-LC3 structures we observe in dividing G361 cells already represent preautophagosomal or autophagosomal membranes. Partial colocalization of endogenous hWIPI-1 with GFP-LC3 suggests that hWIPI-1 is found close to autophagic membranes prior to starvation. Perhaps these structures are the mammalian equivalent of the yeast PAS that might be already present in dividing G361 cells. Consistently, we found hWIPI-1 punctate structures to be sensitive to wortmannin treatment (Figure 7d). Another autophagy-linked protein, Alfy, was recently identified and localized to novel, Alfy-positive structures upon the induction of autophagy (Simonsen et al., 2004). Since Alfy-positive structures were not sensitive to wortmannin, it was postulated that Alfy-positive structures do not represent mammalian PAS-like equivalents (Simonsen et al., 2004).
Strikingly, upon the induction of autophagy by amino-acid depletion, WIPI-1 specific aggregates were identified and shown to be also sensitive to wortmannin administration. However, we were unable to demonstrate a colocalization of GFP-LC3 with the starvation-induced, large hWIPI structures, while we observed again a partial colocalization at punctate structures. It is plausible that the WIPI-1 aggregates that form following the induction of autophagy might represent late autophagosomal structures because autolysosomes have less LC3 in their membrane; hence, the fluorescent signal of GFP-LC3 is weaker (Mizushima et al., 2004). In this context, it is worth noting that Jeffries et al. (2004) reported localization of WIPI49 to endosomal membranes in COS cells. Prior to fusion with lysosomes, autophagosomes can fuse with endosome or endosome-derived vesicles; hence, the WIPI-positive structures in our experiments might reflect this stage of autophagosome formation in mammalian cells. We therefore suggest that (a) co-localization of GFP-LC3 and hWIPI-1 at punctate structures prior to and after the induction of autophagy might represent mammalian PAS-like equivalents, and that (b) the starvation-induced hWIPI-1 formation to large aggregates represents late steps in the autophagic process.
There is increasing evidence that autophagy is a cell death and tumor suppressive mechanism (Gozuacik and Kimchi, 2004), and that defective autophagy aids tumor development (Edinger and Thompson, 2003). It is plausible that WIPI proteins are linked pathologically to cellular transformation because we found that all human WIPI genes were aberrantly expressed in a variety of matched human cancer samples. Strikingly, hWIPI-2 and hWIPI-4 mRNA expression is substantially decreased in 70% of matched kidney (10 patients) and 100% of pancreatic (seven patients) tumor samples. The majority of these samples were derived from advanced stage tumors, such as pancreatic adenocarcinomas stages I–IV. Interestingly, in pancreatic carcinogenesis, autophagic capacity first increases during premalignant stages, but decreases during the pancreatic adenoma to adenocarcinoma transition (Gozuacik and Kimchi, 2004). Hence, pancreatic cancer-associated downregulation of hWIPI-2 and hWIPI-4 supports the possibility that decreased autophagic activity is necessary for the malignant stages of pancreatic cancer (as discussed by Gozuacik and Kimchi, 2004). In addition to decreased mRNA levels in some human cancers, hWIPI-1 is upregulated in 50% of skin cancers tested (10 patients) and in 40% of cervical carcinomas (10 patients); hWIPI-3 mRNA expression is strongly elevated in 70% of uterine cancer patient material and slightly elevated in 60% of the selected ovarian cancers (10 patients). Unraveling the integrity of WIPI genes in the presented matched tumor tissues will be necessary to further understand the consequences of aberrant WIPI gene expression in these cancer tissues. Recently, the tumor suppressor beclin1 has been shown to regulate autophagic cell death (Yu et al., 2004). Our findings of altered hWIPI expression in human cancers and association with autophagic-like structures suggest that dysfunctional WIPI-dependent autophagy may also contribute to tumor progression.
Materials and methods
Isolation of WIPI genes
A partial cDNA fragment of hWIPI-1 (lacking nucleotides 1–152) was isolated (clone 78) from a human liver cDNA library in a screen for p53 inhibitory capacity (Waddell et al., 2001). To generate a full-length hWIPI-1α cDNA clone, the missing 5′-DNA sequence was cloned by RT–PCR (Advantage One-step RT–PCR, BD Biosciences) from human testis mRNA (BD Biosciences). The 3′-oligonucleotide (5′-gag gag aga tct ctg tgc ctt tct tga agt gat-3′) was designed to match a unique BglII site (nucleotides 277/281) within the ORF of hWIPI-1. The 5′-oligonucleotide (5′-aga aga gaa ttc ccg atg gag gcc gag gcc gcg-3′) was designed following alignment of EST sequences retrieved from NCBI (AA482531, Z24843, AA043660, AK079986). This RT–PCR fragment was fused to the initial cDNA clone isolate (clone 78) to generate pAR31CD-hWIPI-1α. To generate GFP-hWIPI-1α, the full length hWIPI-1α cDNA was amplified by PCR (15 cycles; 96°C 1 min, 53°C 1.5 min, 72°C 2.5 min; Pfu Turbo Polymerase/Stratagene) using pAR31CD-hWIPI-1α as a template with the following oligonucleotides: 5′-gag aga ctc gag cta tgg agg ccg agg ccg cgg ac-3′ and 5′-gag aga gaa ttc tca tga ctg ctt cgt ttt gcc ct-3′. The resulting PCR fragment was subcloned into pEGFP.C1 (BD Biosciences; cut XhoI/EcoRI). Cloning of human WIPI family members was conducted by RT–PCR (TITANUM One-Step RT–PCR, BD Biosciences) using human testis mRNA (BD Biosciences) and the following oligonucleotides that were designed according to BLAST search results at NCBI: hWIPI-2α (5′-aga gag gaa ttc tat gaa cct ggc gag cca gag c-3′, 5′-aga gag gga tcc tca gtc agt ccg aag aat cat-3′), hWIPI-2β (5′-aga gag gaa ttc tat gct cct gag gct cca gcg a-3′, 5′-aga gag gga tcc tca gtc agt ccg aag aat cat-3′), hWIPI-3 (5′-aga gag ctc gag cta tgt tat ttc gct gca act at-3′, 5′-aga gag gga tcc tca cag ctt gtc atc ggt cag-3′) and hWIPI-4 (5′-aga gag ctc gag cta tga ctc aac agc cac ttc ga-3′, 5′-aga gag gaa ttc tta aaa gtc atc atc atc aca-3′). The resulting PCR fragments were cloned into pEGFP.C1. All constructs were verified by PCR and overlapping automated DNA sequencing (GENterprise Genomics, Germany). The following GeneBank Accession numbers were assigned to our WIPI isolates: AY691424 (hWIPI-1α), AY691425 (hWIPI-2β), AY691426 (hWIPI-2δ), AY691427 (hWIPI-3) and AY691428 (hWIPI-4).
Bioinformatic analyses
Sequences homologs to hWIPI-1α (GenBank Accession number 31542638) were identified in the nonredundant database at NCBI using the program BLAST (http://www.ncbi.nlm.nih.gov/BLAST/). All sequences with an E-value of 10 or better (295 in all) were extracted and clustered in the program CLANS (Frickey and Lupas, 2004), using P-values ⩽1e−35 as a cutoff for the assignment of an attractive force between sequences (Figure 2, left). CLANS is an implementation of the Fruchterman–Reingold algorithm (Fruchterman and Reingold, 1991). A set of 80 sequences corresponding to the two WIPI subfamilies identified in the clustering analysis and eight sequences representing the WD-repeat protein outgroup were further analysed phylogenetically using the Asatura software (Van de Peer et al., 2002). The neighbor-joining phylogeny, computed with a Poisson distance correction, is shown in Figure 2 (right). In all, 32 sequences of the WIPI family representing all phylogenetic clades and selected for less than 70% sequence identity were aligned interactively using the program MACAW (Schuler et al., 1991). Blocks with significance better than e−20 were retained; these essentially corresponded to the β-strands of the propeller blades, with the exception of strand 7a, which was aligned by the Gibbs sampling procedure. The terminal extensions and the inserted region between strands 6c and 6d were readily apparent as unalignable and allowed the unambiguous assignment of the location for strand 6d. Positions of this alignment that were occupied in at least 75% of the sequences (24) by hydrophobic residues were assigned as conserved hydrophobic and were used together with consensus secondary structure predictions by Psipred (Jones, 1999) and JNet (Cuff and Barton, 1999) to align the propeller blades to the WD-repeat consensus. The quality of the alignments was evaluated in MACAW; all ungapped blocks had a significance better than e−20, except for the alignment for strand 7a. Positions occupied by the same polar residue in at least 24 sequences were marked as conserved (red in Figure 3) and positions with the same polar residue in at least 31 sequences were marked as invariant (magenta in Figure 3). From the alignment to the WD-repeat consensus, we built a molecular model of hWIPI-1α in Modeller 6v2 (Sali et al., 1995), using the structures of G-protein beta subunit and TUP1 as templates (Figure 3).
In vitro translation
The TNT Coupled Reticulocyte Lysate System (Promega) was used according to the manufacturer's protocol with either of the following constructs: pSG5-mERα(MOR); pSG5-ERβ; pSG5-AR (respectively, gifts from Frank Gannon, Heidelberg, Germany; Malcolm Parker, London, UK; Roland Schuele, Freiburg, Germany). Construct integrities were verified by in vitro transcription/translation.
GST pull-down experiments
To generate a GST-hWIPI-1α fusion construct, the hWIPI-1α cDNA lacking the start codon was cloned into pGEX6P3 (Amersham). Escherichia coli BL21 cells (Stratagene) were used as host to induce the expression of GST (4 h at 30°C) or GST-hWIPI-1α (4 h at 25°C) with 1 mM IPTG. Cells were harvested in STE buffer (0.1 M NaCl, 10 mM Tris pH 8.0, 1 mM EDTA), lysed (15 min, 4°C) using lysozyme (0.1 mg/ml) and supplemented with 5 mM DTT and 1.5% sarcosyl. Cleared lysates were coupled to glutathione sepharose 4B (Amersham) by constant rotation (4 h at 4°C) and washed with PBS and incubation buffer (20 mM HEPES/KOH pH 7.9, 20% glycerol, 100 mM KCl, 5 mM MgCl2, 0.2 mM EDTA, 0.01% NP-40, 1 mM DTT, 0.2 mM PMSF, Roche Complete/Protease inhibitor cocktail). Subsequent GST pull-down experiments were conducted with 5–10 μl aliquots of in vitro-translated products (in the presence or absence of 500 nM 5α-androstan or 10 nM β-estradiol) in the presence of 1.5% BSA (rotations routinely for 60 min at 4°C). Thereafter, sediments were washed 3 × (15 min rotations at 4°C) in incubation buffer and heated in Laemmli buffer (15 min, 80°C) to separate proteins from glutathione sepharose. Aliquots of supernatants (6500 r.p.m., 5 min) were resolved by SDS–PAGE (10%). Protein gels were stained with Coomassie blue, fixed, dried and exposed to KODAK BioMAX Film.
WIPI mRNA profiles
To generate specific 32P-labeled cDNA probes for hWIPI family members, the following oligonucleotides were used in standard PCR amplifications (20 cycles: 94°C 30 s, 55°C 30 s, 72°C 1 min, one cycle: 72°C 2 min; REDTaqTM Polymerase, SIGMA): hWIPI-1 (5′-aat aaa gaa aat gac ctc aga-3′, 5′-tca tga ctg ctt cgt ttt gcc-3′), hWIPI-2 (5′-aat gag atc ttg gac tct gcc-3′, 5′-tca gtc agt ccg aag aat cat-3′), hWIPI-3 (5′-tca gcc agt ttc ctt cca aaa-3′, 5′-tca cag ctt gtc atc ggt cat-3′) and hWIPI-4 (5′-cgc gtg ggc aag gtg ggg cct-3′, 5′-tta aaa gtc atc atc atc aca-3′). These PCR fragments were labeled using the Random Primers DNA Labeling System (Invitrogen), and purified with Chroma SpinTM Colums (BD Biosciences) according to the manufacturer's protocols. For Northern hybridizations, the instructions supplied with the following arrays/blots were adhered to: Cancer Profiling Array II (see Supplementary Table), RNA Master Blot, Human MTN Blot I/II (BD Biosciences). The RNeasy Kit (Qiagen) was used for isolation of total RNA from either of the human melanoma cell lines (a gift from Birgit Schittek, Tuebingen, Germany) listed in Figure 6, and standard Northern blotting and hybridization procedures performed by using ExpressHyb Solution (BD Biosciences). To control for equal loading, RNA agarose gels were stained with ethidium bromide (data not shown) and arrays/blots were hybridized with 32P-labelled actin or ubiquitin (BD Biosciences).
WIPI antiserum
The polyclonal hWIPI-1 antiserum was generated by immunizing rabbits (Charles River) with two synthetic peptides representing C-terminal regions in hWIPI-1 (363nkendlrpslp373, 407lrgevipehefatgpv422) and used at a dilution of 1 : 5000 in ECL Western blotting detections (Amersham), and at 1 : 250 for indirect immunofluorescent stainings (see below).
Cell culture, transfections and confocal microscopy
G361 (a gift from Birgit Schittek, Tuebingen, Germany) and HeLa cells (ATCC) were routinely cultured in Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum, 100 μg/ml penicillin and 100 μg/ml streptomycin. For transient transfections, cells were seeded on coverslips and grown to 80% confluence in the absence of antibiotics at 37°C and 5% CO2. Transfections were conducted according to the manufacturer's protocol using LipofectAMINE 2000 (Invitrogen) and a GFP-LC3 construct (a gift from Tamotsu Yoshimori) at a ratio of 2.5 : 1. At 48 h posttransfection, cells were fixed in 3.7% paraformaldehyde (15 min, RT) and analysed by confocal microscopy (see below). To visualize endogenous hWIPI-1 in G361 cells consecutive incubations were conducted using our C-terminal hWIPI-1 antiserum at 1 : 250 (30 min, RT) and Alexa Fluor 546 or Alexa Fluor 488 goat anti-rabbit IgG (Molecular Probes) at 1 : 500 (30 min, RT) in PBS/0.1% Tween (blocking/permeabilization: PBS/0.1% Tween/1% BSA, 30 min RT; washing: PBS/0.1% Tween). For nuclear staining, TO-PRO-3 (Molecular Probes) was used at 1 : 1000. Finally, cells were mounted in Vectashield Mounting Medium (Vector Laboratories) and kept in the dark (4°C). Routinely, control GFP vector (pEGFP.C1, BD Biosciences) transfections were carried out as well as control incubations with secondary fluorescent antibodies alone (data not shown). Confocal laser scanning microscopy was conducted using a LSM510 microscope (Zeiss) and a 63 × 1.4 DIC Plan-Apochromat oil-immersion objective. GFP and Alexa 488 were excited at 488 nm with the internal argon-ion laser. Alexa 546 and TO-PRO-3, respectively, were excited at 543 and 633 nm with helium–neon lasers. A series of 10–20 sections with 0.5 μm spacing along the z-axis was taken and projections were created from confocal stacks by merging the individual confocal slizes.
Autophagy induction
Autophagy was induced by amino-acid starvation in EBSS medium (Sigma) according to Munafo and Colombo (2001) in the presence or absence of wortmannin (200 nM). For wortmannin administration, cells were preincubated with the drug in complete medium for 60 min. Incubation of cells in EBSS occurred for 3 h, an optimum for the accumulation of autophagosomes (Mitchener et al., 1976).
References
Barth H, Meiling-Wesse K, Epple UD and Thumm M . (2001). FEBS Lett., 508, 23–28.
Blommaart EF, Krause U, Schellens JP, Vreeling-Sindelarova H and Meijer AJ . (1997). Eur. J. Biochem., 243, 240–246.
Chernova OB, Hunyadi A, Malaj E, Pan H, Crooks C, Roe B and Cowell JK . (2001). Oncogene, 20, 5378–5392.
Cuff JA and Barton GJ . (1999). Proteins, 40, 502–511.
Dove SK, Piper RC, McEwen RK, Yu JW, King MC, Hughes DC, Thuring J, Holmes AB, Cooke FT, Michell RH, Parker PJ and Lemmon MA . (2004). EMBO J., 23, 1922–1933.
Edinger AL and Thompson CB . (2003). Cancer Cell, 4, 422–424.
Fong HK, Hurley JB, Hopkins RS, Miake-Lye R, Johnson MS, Doolittle RF and Simon MI . (1986). Proc. Natl. Acad. Sci. USA, 83, 2162–2166.
Frickey T and Lupas A . (2004). Bioinformatics [Epub ahead of print].
Fruchterman TM and Reingold EM . (1991). Software Pract. Exp., 21, 1129–1164.
Garcia-Higuera I, Fenoglio J, Li Y, Lewis C, Panchenko MP, Reiner O, Smith TF and Neer EJ . (1996). Biochemistry, 35, 13985–13994.
Garcia-Higuera I, Gaitatzes C, Smith TF and Neer EJ . (1998). J. Biol. Chem., 273, 9041–9049.
Georgakopoulos T, Koutroubas G, Vakonakis I, Tzermia M, Prokova V, Voutsina A and Alexandraki D . (2001). Yeast, 18, 1155–1171.
Gozuacik D and Kimchi A . (2004). Oncogene, 23, 2891–2906.
Guan J, Stromhaug PE, George MD, Habibzadegah-Tari P, Bevan A, Dunn Jr WA and Klionsky DJ . (2001). Mol. Cell. Biol., 12, 3821–3838.
Honore B, Baandrup U, Nielsen S and Vorum H . (2002). Oncogene, 21, 1123–1129.
Jeffries TR, Dove SK, Michell RH and Parker PJ . (2004). Mol. Cell. Biol., 15, 2652–2663.
Jones DT . (1999). J. Mol. Biol., 292, 195–202.
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T, Kominami E, Ohsumi Y and Yoshimori T . (2000). EMBO J., 19, 5720–5728.
Kim J and Klionsky DJ . (2000). Annu. Rev. Biochem., 69, 303–342.
Klionsky DJ and Emr SD . (2000). Science, 290, 1717–1721.
Klionsky DJ . (2004). Nature, 431, 31–32.
Lee C-Y and Baehrecke EH . (2001). Development, 128, 1443–1455.
Li D and Roberts R . (2001). Cell Mol. Life Sci., 58, 2085–2097.
Melendez A, Talloczy Z, Seaman M, Eskelinen EL, Hall DH and Levine B . (2003). Science, 301, 1387–1391.
Mitchener JS, Shelburne JD, Bradford WD and Hawkins HK . (1976). Am. J. Pathol., 83, 485–491.
Mizushima N, Kuma A, Kobayashi Y, Yamamoto A, Matsubae M, Takao T, Natsume T, Ohsumi Y and Yoshimori T . (2003). J. Cell Sci., 116, 1679–1688.
Mizushima N, Ohsumi Y and Yoshimori T . (2002). Cell Struct. Funct., 27, 421–429.
Mizushima N, Yamamoto A, Matsui M, Yoshimori T and Ohsumi Y . (2004). Mol. Cell. Biol., 15, 1101–1111.
Munafo DB and Colombo MI . (2001). J. Cell Sci., 114, 3619–3629.
Neer EJ, Schmidt CJ, Nambudripad R and Smith TF . (1994). Nature, 371, 297–300.
Neer EJ and Smith TF . (1996). Cell, 84, 175–178.
Paoli M . (2001a). Nat. Struct. Biol., 8, 744–745.
Paoli M . (2001b). Prog. Biophys. Mol. Biol., 76, 103–130.
Petiot A, Ogier-Denis E, Blommaart EF, Meijer AJ and Codogno P . (2000). J. Biol. Chem., 275, 992–998.
Puntervoll P, Linding R, Gemund C, Chabanis-Davidson S, Mattingsdal M, Cameron S, Martin DM, Ausiello G, Brannetti B, Costantini A, Ferre F, Maselli V, Via A, Cesareni G, Diella F, Superti-Furga G, Wyrwicz L, Ramu C, McGuigan C, Gudavalli R, Letunic I, Bork P, Rychlewski L, Kuster B, Helmer-Citterich M, Hunter WN, Aasland R and Gibson TJ . (2003). Nucleic Acids Res., 31, 3625–3630.
Sali A, Potterton L, Yuan F, van Vlijmen H and Karplus M . (1995). Proteins, 23, 318–326.
Schuler GD, Altschul SF and Lipman DJ . (1991). Proteins, 9, 180–190.
Schultz J, Milpetz F, Bork P and Ponting CP . (1998). Proc. Natl. Acad. Sci. USA, 95, 5857–5864.
Simonsen A, Birkeland HC, Gillooly DJ, Mizushima N, Kuma A, Yoshimori T, Slagsvold T, Brech A and Stenmark H . (2004). J. Cell Sci., 117, 4239–4251.
Smith TF, Gaitatzes C, Saxena K and Neer EJ . (1999). Trends Biochem. Sci., 24, 181–185.
Stromhaug PE, Reggiori F, Guan J, Wang CW and Klionsky DJ . (2004). Mol. Cell. Biol., 15, 3553–3566.
Tanida I, Ueno T and Kominami E . (2004). Int. J. Biochem. Cell Biol., 36, 2503–2518.
Van de Peer Y, Frickey T, Taylor J and Meyer A . (2002). Gene, 295, 205–211.
van Nocker S and Ludwig P . (2003). BMC Genom., 4, 50.
Waddell S, Jenkins JR and Proikas-Cezanne T . (2001). Oncogene, 20, 6001–6008.
Yu L, Alva A, Su H, Dutt P, Freundt E, Welsh S, Baehrecke EH and Lenardo MJ . (2004). Science, 304, 1500–1502.
Yu L, Gaitatzes C, Neer E and Smith TF . (2000). Protein Sci., 9, 2470–2476.
Acknowledgements
We are most grateful to Tamotsu Yoshimori for kindly providing the GFP-LC3 construct, and thank Johannes Soeding and Yakov Sergeev for assistance with the hWIPI-1α model. We also thank Malcolm Parker, Frank Gannon, Roland Schuele, Hubert Kalbacher and Birgit Schittek for reagents used in this study. We appreciate initial bioinformatic analysis by Stefan Steigele and Kay Nieselt-Struwe, and expert advice on confocal microscopy by Roland Brock. Initial isolation of hWIPI-1α was part of a postdoctoral fellowship by the Deutsche Forschungsgemeinschaft, Germany to TP-C, conducted at the Marie Curie Research Institute, UK. This work is supported by the Fritz Thyssen Stiftung, Germany, and the Fonds der Chemischen Industrie.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Oncogene website (http://www.nature.com/onc)
Supplementary information
Rights and permissions
About this article
Cite this article
Proikas-Cezanne, T., Waddell, S., Gaugel, A. et al. WIPI-1α (WIPI49), a member of the novel 7-bladed WIPI protein family, is aberrantly expressed in human cancer and is linked to starvation-induced autophagy. Oncogene 23, 9314–9325 (2004). https://doi.org/10.1038/sj.onc.1208331
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1208331
Keywords
This article is cited by
-
The ABL-MYC axis controls WIPI1-enhanced autophagy in lifespan extension
Communications Biology (2023)
-
Exploring and exploiting the host cell autophagy during Mycobacterium tuberculosis infection
European Journal of Clinical Microbiology & Infectious Diseases (2023)
-
A comprehensive phenotypic characterization of a whole-body Wdr45 knock-out mouse
Mammalian Genome (2021)
-
Wipi3 is essential for alternative autophagy and its loss causes neurodegeneration
Nature Communications (2020)
-
A single nucleotide polymorphism in the Plasmodium falciparum atg18 gene associates with artemisinin resistance and confers enhanced parasite survival under nutrient deprivation
Malaria Journal (2018)