Abstract
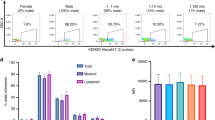
Chimerism analysis has become a routine method to document engraftment and also for detection of residual disease. PCR-based procedures using STR analysis, especially commercially available multiplex assays, are frequently used. However, these assays have been optimized for forensic purposes and do not necessarily fulfil all needs for chimerism analysis. To improve these analyses, data on the level of informativity of STR systems in the context of chimerism analysis would be helpful. We evaluated 27 STR markers for their informativity in 203 patients and their HLA-matched related donors. These STRs included 18 from different multiplex kits, whereas nine were selected from the literature or STR databases. The STR profiles were ranked from Type 1 (not informative) to Type 5 (best suited for chimerism analysis). According to this ranking, the informativity of the STR systems was found highly variable, ranging from 4.4 to 49.0% Type 5 constellations. Among the most informative STRs were Penta E, SE33, D2S1338 and D18S51. Informativity of an STR was correlated with the degree of heterozygosity (r=0.86; P=0.0001), but not with the total number of alleles present. These data indicate that selection of suitable STR markers is important to improve diagnostics based on STR analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bryant E, Martin PJ . Documentation of engraftment and characterization of chimerism following hematopoietic cell transplantation. In: Thomas ED, Blume KG, Forman SJ (eds). Hematopoietic Cell Transplantation. Oxford, London: Blackwell Science, 1999, pp 197–206.
Lawler M, Humphries P, McCann SR . Evaluation of mixed chimerism by in vitro amplification of dinucleotide repeat sequences using the polymerase chain reaction. Blood 1991; 77: 2504–2514.
Bader P, Holle W, Klingebiel T, Handgretinger R, Niethammer D, Beck J . Quantitative assessment of mixed hematopoietic chimerism by polymerase chain reaction after allogeneic BMT. Anticancer Res 1996; 16: 1759–1763.
Briones J, Urbano-Ispizua A, Rozman C, Marin P, Carreras E, Rovira M et al. Study of hematopoietic chimerism following allogeneic peripheral blood stem cell transplantation using PCR amplification of short tandem repeats. Ann Hematol 1996; 72: 265–268.
Scharf SJ, Smith AG, Hansen JA, McFarland C, Erlich HA . Quantitative determination of bone marrow transplant engraftment using fluorescent polymerase chain reaction primers for human identity markers. Blood 1995; 85: 1954–1963.
Dubovsky J, Daxberger H, Fritsch G, Printz D, Peters C, Matthes S et al. Kinetics of chimerism during the early post-transplant period in pediatric patients with malignant and non-malignant hematologic disorders: implications for timely detection of engraftment, graft failure and rejection. Leukemia 1999; 13: 2060–2069.
Thiede C, Florek M, Bornhauser M, Ritter M, Mohr B, Brendel C et al. Rapid quantification of mixed chimerism using multiplex amplification of short tandem repeat markers and fluorescence detection. Bone Marrow Transplant 1999; 23: 1055–1060.
Pindolia K, Janakiraman N, Kasten-Sportes C, Demanet C, Van Waeyenberge C, Pals G et al. Enhanced assessment of allogeneic bone marrow transplant engraftment using automated fluorescent-based typing. Bone Marrow Transplant 1999; 24: 1235–1241.
Nollet F, Billiet J, Selleslag D, Criel A . Standardisation of multiplex fluorescent short tandem repeat analysis for chimerism testing. Bone Marrow Transplant 2001; 28: 511–518.
Butler JM . Commonly used short tandem repeat markers. In: Butler JM (ed). Forensic DNA typing- Biology & Technology behind STR markers. London: Academic Press, 2001, pp 53–80.
Walsh PS, Fildes NJ, Reynolds R . Sequence analysis and characterization of stutter products at the tetranucleotide repeat locus vWA. Nucleic Acids Res 1996; 24: 2807–2812.
Butler JM . Biology of STRs: stutter products, non-template addition, microvariants, null alleles, and mutation rates. In: Butler JM (ed). Forensic DNA typing- Biology & Technology behind STR markers. London: Academic Press, 2001, pp 81–98.
Lazaruk K, Wallin J, Holt C, Nguyen T, Walsh PS . Sequence variation in humans and other primates at six short tandem repeat loci used in forensic identity testing. Forensic Sci Int 2001; 119: 1–10.
Bacher J, Schumm JW . Development of highly polymorphic pentanucleotide tandem repeat loci with low stutter. Profiles DNA 1998; 2: 3–6.
Nollet F, Billiet J, Selleslag D, Criel A . Standardisation of multiplex fluorescent short tandem repeat analysis for chimerism testing. Bone Marrow Transplant 2001; 28: 511–518.
Thiede C, Lion T . Quantitative analysis of chimerism after allogeneic stem cell transplantation using multiplex PCR amplification of short tandem repeat markers and fluorescence detection. Leukemia 2001; 15: 303–306.
Powerplex 16 System. Technical Manual No. D012. Madison, WI: Promega Corporation, 2002, pp. 3–7.
AmpFlSTR Profiler SGM kit. AmpFlSTR SGM Plus PCR Amplification Kit User's Manual. Foster City, CA: Applied Biosystems, 1999, pp. 1–16.
AFS. Mentype Triplex AFS User Manual. Dresden: Biotype AG, 2002, pp. 1–2.
Dubovsky J, Daxberger H, Fritsch G, Printz D, Peters C, Matthes S et al. Kinetics of chimerism during the early post-transplant period in pediatric patients with malignant and non-malignant hematologic disorders: implications for timely detection of engraftment, graft failure and rejection. Leukemia 1999; 13: 2060–2069.
Ruitberg CM, Reeder DJ, Butler JM . STRBase: a short tandem repeat DNA database for the human identity testing community. Nucleic Acids Res 2001; 29: 320–322.
Thiede C, Bornhauser M, Oelschlagel U, Brendel C, Leo R, Daxberger H et al. Sequential monitoring of chimerism and detection of minimal residual disease after allogeneic blood stem cell transplantation (BSCT) using multiplex PCR amplification of short tandem repeat-markers. Leukemia 2001; 15: 293–302.
Thiede C, Florek M, Bornhauser M, Ritter M, Mohr B, Brendel C et al. Rapid quantification of mixed chimerism using multiplex amplification of short tandem repeat markers and fluorescence detection. Bone Marrow Transplant 1999; 23: 1055–1060.
Nollet F, Billiet J, Selleslag D, Criel A . Standardisation of multiplex fluorescent short tandem repeat analysis for chimerism testing. Bone Marrow Transplant 2001; 28: 511–518.
Bar W, Brinkmann B, Budowle B, Carracedo A, Gill P, Lincoln P, Mayr W, Olaisen B . DNA recommendations. Further report of the DNA Commission of the ISFH regarding the use of short tandem repeat systems. International Society for Forensic Haemogenetics. Int J Legal Med 1997; 110: 175–176.
Piccinini A, Waterkamp K, Meyer E . Short tandem repeat HumACTBP2 (SE33) and HumVWA: population genetic study on a north Italian population. Int J Legal Med 1997; 110: 292–294.
Hou YP, Tang JP, Dong JG, Ji Q, Li YB, Wu J et al. Further characterization and population data for the pentanucleotide STR polymorphism D10S2325. Forensic Sci Int 2001; 123: 107–110.
Lion T . Summary: reports on quantitative analysis of chimerism after allogeneic stem cell transplantation by PCR amplification of microsatellite markers and capillary electrophoresis with fluorescence detection. Leukemia 2003; 17: 252–254.
Hancock JP, Goulden NJ, Oakhill A, Steward CG . Quantitative analysis of chimerism after allogeneic bone marrow transplantation using immunomagnetic selection and fluorescent microsatellite PCR. Leukemia 2003; 17: 247–251.
Acquaviva C, Duval M, Mirebeau D, Bertin R, Cave H . Quantitative analysis of chimerism after allogeneic stem cell transplantation by PCR amplification of microsatellite markers and capillary electrophoresis with fluorescence detection: the Paris-Robert Debre experience. Leukemia 2003; 17: 241–246.
Kreyenberg H, Holle W, Mohrle S, Niethammer D, Bader P . Quantitative analysis of chimerism after allogeneic stem cell transplantation by PCR amplification of microsatellite markers and capillary electrophoresis with fluorescence detection: the Tuebingen experience. Leukemia 2003; 17: 237–240.
Chalandon Y, Vischer S, Helg C, Chapuis B, Roosnek E . Quantitative analysis of chimerism after allogeneic stem cell transplantation by PCR amplification of microsatellite markers and capillary electrophoresis with fluorescence detection: the Geneva experience. Leukemia 2003; 17: 228–231.
Schraml E, Daxberger H, Watzinger F, Lion T . Quantitative analysis of chimerism after allogeneic stem cell transplantation by PCR amplification of microsatellite markers and capillary electrophoresis with fluorescence detection: the Vienna experience. Leukemia 2003; 17: 224–227.
Acknowledgements
This study was supported in part by the Deutsche Krebshilfe, Bonn (Grant number 70-2755 to CT and MB).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Thiede, C., Bornhäuser, M. & Ehninger, G. Evaluation of STR informativity for chimerism testing – comparative analysis of 27 STR systems in 203 matched related donor recipient pairs. Leukemia 18, 248–254 (2004). https://doi.org/10.1038/sj.leu.2403212
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2403212
Keywords
This article is cited by
-
Indel analysis by droplet digital PCR: a sensitive method for DNA mixture detection and chimerism analysis
International Journal of Legal Medicine (2017)
-
Sex chromosome loss after allogeneic hematopoietic stem cell transplant in patients with hematologic neoplasms: a diagnostic dilemma for clinical cytogeneticists
Molecular Cytogenetics (2016)
-
Tri-allelic patterns at the D7S820 locus detected in two generations of a Chinese family
International Journal of Legal Medicine (2016)
-
Quantitative monitoring of multi-donor chimerism: a systematic, validated framework for routine analysis
Bone Marrow Transplantation (2010)