Abstract
The prognostic value of the detection of peripheral blood (PB) and/or bone marrow (BM) involvement by polymerase chain reaction (PCR) amplification of rearranged immunoglobulin heavy chain (IgH) and immunoglobulin kappa light chain (Igκ) genes was evaluated in 155 patients with diffuse large B-cell lymphomas (DLBCL). Immunoglobulin gene rearrangements (IgR) were detected in 35/155 (23%) patients. The presence of IgR in PB/BM was related to clinical stage (CS I–III vs CS IV; P<0.001), histopathological detection of BM involvement (P<0.001), and the International Prognostic Index (P<0.001). IgR-positive cases had a significantly lower complete remission (CR) rate (18/35, 51%) than IgR-negative patients (85/120, 71%; P=0.042), and a significantly poorer overall survival (OAS) at 5 years (25 vs 66%; P<0.001). There was a significant difference in the estimated OAS at 5 years between patients with negative BM histology and negative PCR results (66%), patients with negative BM histology but positive IgR (37%), and patients with positive BM histology (12%). Our results indicate that molecular methods improve the accuracy of staging in patients with DLBCL and define a group of patients with normal bone marrow histology who have a significantly poorer OAS due to molecular detection of PB/BM involvement.
Similar content being viewed by others
Introduction
Diffuse large B-cell lymphomas (DLBCLs) are diagnosed in 30–40% of patients with non-Hodgkin's lymphomas (NHL). Diagnosis and classification are mainly based on morphologic and immunophenotypic analyses. More recently, cytogenetic and molecular techniques have provided additional information on the biology of DLBCLs. Up to one-third of cases have abnormalities of BCL6, and about 20% of cases have translocations of BCL2. Staging is based on clinical, radiological, and histological findings including bone marrow histology. Approximately 70% of early stages and 50% of advanced-stage DLBCL can be cured by chemotherapy with or without radiotherapy.1,2 The clinical stage is an important prognostic factor and was therefore included in the International Prognostic Index.3 Systemic dissemination to the peripheral blood and bone marrow occurs with advanced tumor stages and is associated with worse prognosis, but minimal morphologic changes may not be diagnosed by conventional methods in early stages of the disease.
The majority of NHL have undergone physiological, clonal immunoglobulin (Ig), or T-cell receptor (TCR) rearrangements. The identification of lymphoid clonality by detection of a clonal Ig/TCR rearrangement is therefore used at diagnosis and increasingly for follow-up studies.4,5 PCR detection of Ig/TCR V(D)J gene rearrangements exploits the fact that the junctional regions are clone specific and highly variable, both with regard to length and nucleotide content. The most sensitive and specific detection techniques involve sequencing of the clonal V-J junction and synthesis of an antijunctional or allele-specific oligonucleotide, which is then used as a hybridization probe or clone-specific primer.6,7 While these strategies represent optimal specificity and sensitivity (approximately 10−5), they are labor intensive and do not permit detection of clonal evolution. Simpler strategies as consensus PCR that provide an intermediate level of detection (approximately 1%) may be more easily applicable to large-scale clinical studies. The immunoglobulin heavy chain (IgH) gene represents the most useful gene target for detecting B-cell clonality since it rearranges early during B-lymphoid development and demonstrates extensive junctional diversity.8,9 The incidence of informativity of IgH PCR in B-cell lymphomas varies with the IgH PCR strategy and the pathological subtype.10,11 The rearranged genomic products of the immunolobulin κ light chain (Igκ) gene also represent an excellent marker for B-cell clonal analysis. Analysis of Igκ in conjunction with IgH rearrangements substantially improves the detection of lymphoid clonality in B-cell neoplasms.12,13
The relatively insensitive Southern blot technique has been used to study the involvement of peripheral blood or bone marrow in a variety of NHL subtypes.14,15 However, there are no large clinical studies evaluating the use of PCR staging in DLBCL.
The aim of this study was to determine the prognostic value of rearranged Ig genes in the PB and BM of patients with DLBCL at the time of diagnosis. In particular, we wished to assess the additional information provided by molecular staging in patients with localized clinical stages and in patients with advanced stages but negative bone marrow histology.
Patients and methods
Patients
A total of 155 adult patients with DLBCL were included in this study. The median age of the patients was 59 years (range, 17–88 years), 84 were male, 71 were female subjects. They had all been diagnosed between March 1991 and November 2002 at the Department of Internal Medicine I, Division of Hematology, University of Vienna. Immunocompromised patients were not included. Diagnosis was based on histopathologic and immunophenotypic assessment of formaldehyde-fixed, paraffin-embedded tumor biopsy specimen according to the Revised European–American Classification.16,17 For adequate clinical staging, computed tomographic scans of the thorax and abdomen as well as a bone marrow biopsy were required. In all, 38 patients (25%) had clinical stage (CS) I, 47 (30%) CS II, 34 (22%) CS III, and 36 (23%) CS IV disease. Bone marrow involvement was detected histologically in 19 patients with CS IV disease. In two of these patients, circulating tumor cells also were detected by the differential blood count. In 130 patients, lymphoma cells were not found in the BM biopsies. In six patients, the BM biopsies were inconclusive. Patients were assigned to specific risk groups according to the International Prognostic Index (IPI): 71 (46%) patients had IPI 0–1, 67 (43%) patients IPI 2–3, 17 (11%) patients IPI 4–5.
Treatment
The following chemotherapeutic regimen were used: CHOP in 125 patients,18,19 ProMACE-CytaBOM in 25 patients,20 CNOP in three patients,21 P-DOCE in one patient,22 and CEP in one patient.23 In CS I/II patients without bulky disease, treatment consisted of 3–4 cycles of chemotherapy followed by involved field radiotherapy. CS IIbulky to CS IV patients received six cycles of chemotherapy. Bulky disease was treated with involved field radiotherapy. A total of 20 patients were treated with an autologous stem cell transplant for consolidation or salvage therapy, one patient received an allogeneic transplant from a sibling, and four patients from an unrelated donor. For assessing response to therapy, standardized criteria were followed.24 Response to therapy was evaluated 1–2 months after the end of treatment.
Detection of clonal gene rearrangements by PCR analysis of genomic DNA
Peripheral blood samples (25 patients), BM aspirates (18 patients), or both (112 patients) were available for extraction of genomic DNA and subsequent PCR analysis. In addition, diagnostic material of the initial tumor biopsy of 108 patients was analyzed by molecular methods.
DNA was prepared from mononuclear cells of PB, BM, and native tissue using cell lysis, proteinase K (Boehringer Mannheim, Mannheim, Germany) digestion, phenol extraction, and ethanol precipitation according to standard protocols.25 For preparation of DNA from paraffin blocks, two 10 μm thick paraffin sections were dewaxed with xylene, digested with proteinase K without ionic detergents followed by boiling without phenol extraction as previously described.26
PCR amplification of IgH gene V-D-J junctional variability was performed using consensus primers against the four framework regions FR1 (FR1c), FR2 (FR2), FR3 (VH26, V670, FR3A), and FR4 (OL-4, VLJH) as previously described.11,27,28,29,30 For PCR analysis of Igκ gene rearrangements, oligonucleotide primers recognizing the FR3 (FR3κ) and the joint region (Jκ) of the Igκ gene were used.12 Analysis of immunoglobulin gene rearrangements (IgR) was performed by amplification of 300 ng of genomic DNA with GeneAmp PCR Core Reagents (Perkin Elmer Cetus, Norwalk, CT, USA) using 0.6 μ M primers, 200 μ M GeneAmp dNTPs, 2.5 mM MgCl2, and 0.9 U AmpliTaq Gold DNA Polymerase in a total reaction volume of 30 μl. After an initial denaturation of 10 min at 95°C, 44 cycles of 45 s at 95°C, 1 min at 58°C, and 1 min at 72°C, followed by 10 min at 95°C and rapid cooling to 4°C were performed in a Perkin-Elmer Cetus DNA thermal cycler (Emeryville, CA, USA). PCR products (6 μl) were electrophoresed through Spreadex EL 400 (FR3 and Igκ PCR products), EL 600 (FR2 PCR products), and EL 800 (FR1 PCR products) mini gels (Elchrom Scientific, Cham, Switzerland) at 125 V and 30°C for 2.5, 3.5, and 5 h, respectively. Following electrophoresis, the gels were stained with SYBR Green I (Molecular Probes, Leiden, The Netherlands) for 25 min and destained in distilled water for 24 h prior to photography.
The sensitivity of the IgH and Igκ PCR assays to detect a clonal lymphoid population within a polyclonal cell population was evaluated by mixing DNA of monoclonal cell population from a patient at the time of diagnosis (100% clonal cells) with different amounts of a mixture of polyclonal DNAs obtained from polyclonal mononuclear cells of 32 healthy individuals. The sensitivity was at least 1%.
The quality of IgR-negative samples was assessed by amplification of a polyclonal background smear, which indicates amplifiable DNA from polyclonal B lymphocytes, and the amplification of specific predictable bands, which are seen in all cases and represent nonspecific amplifications of sequences homologous to the JH locus.
Precautions to prevent crosscontamination of the amplified material were taken according to published recommendations.31,32
In each experiment, a reagent control (one aliquot of PCR reagents without genomic DNA), a negative control (mixture of genomic DNA from healthy individuals), and positive controls (genomic DNA from monoclonal neoplastic B-cells, undiluted and diluted 1:100) were coamplified with the samples.
Statistical analysis
Categorical data was compared between groups by means of Fisheŕs exact test in the case of dichotomous variables and using the Cochran–Armitage trend test in the case of IPI risk groups (low/intermediate/high). The dependency of the complete remission (CR) rate and (overall or progression-free) survival on IgR was investigated using a univariate logistic and a Cox proportional hazards regression model, respectively, and is described via odds ratios (OR) and hazard ratios (HR), respectively, with 95% confidence intervals (CI). The 5-year survival and 95% confidence intervals are given for various groupings of patients. In order to assess the additional prognostic impact of IgR in the case of negative BM histology, a multiple model was used where, in addition to BM histology, a categorical variable entered which discriminates between negative and positive IgR only in the case of negative BM histology. For further adjustment, a third model was used with the IPI score (levels 0–5, taken as continuous variable) as additional independent variable. IPI was tested for possible nonlinear effects by adding a quadratic term to the model equation. In order to test for potential nonproportionality of the hazards with respect to IgR, a term has been added for the interaction between IgR and the logarithm of survival time. Overall survival (OAS) time was calculated from the date of beginning of chemotherapy until the patient's death or last follow-up examination. Progression-free survival time was calculated from the date of achievement of CR until relapse, death, or last follow-up examination. Kaplan–Meier survival curve estimates are given for various groupings of patients, together with P-values of the log-rank statistic for equality over the two or three groups, respectively.33 Patients who received a stem cell transplant were treated as censored observations. The P-values ⩽0.05 were considered to be statistically significant. All computations have been performed using WinSTAT Version 3.0 and SAS software Version 8.2 (SAS Institute Inc., Cary, NC, USA, 2001).
Results
Detection of IgR in PB/BM and in tumor biopsies
In 155 patients with DLBCL, PB (n=25), BM (n=18), or both (n=112) were analyzed at the time of diagnosis for the presence of clonal IgH and Igκ gene rearrangements. In total, clonal IgR were found in 35 of 155 (23%) patients: IgH in 19, IgH+Igκ in 16 cases. Of the 112 patients in whom BM and PB was tested, 19 showed Ig rearrangements in BM and PB, 90 had no rearrangements in both materials, and three showed positive results in BM, but not in PB. In the remaining 13 patients with clonal IgR, positive PCR results were detected either in BM (n=6) or in PB (n=7), as the corresponding material was not available for analysis.
Material of the initial tumor biopsy was available for analysis in 108 patients. Clonal IgR were detected in 93 of 108 (86%) tumor specimens. The tumor biopsies exhibited rearrangements of IgH in 45, Igκ in eight, and IgH+Igκ in 40 cases. PCR results of tumor biopsies were compared with results obtained from PB samples and/or BM aspirates. In 23 (21%) patients, identical clonal IgRs were demonstrated in tumor biopsies and PB/BM. In 70 (65%) patients, clonal Ig rearrangements could only be detected in the tumor biopsy, but not in PB/BM. In 15 (14%) patients, neither tumor nor PB/BM exhibited IgR.
Clonal IgR in PB/BM correlate with clinical stage, BM involvement, and IPI (Table 1)
The percentage of PCR-positive samples at various clinical stages is shown in Table 1. The data show a significant difference between stages CS I–III (16 of 119 patients; 13%) and CS IV (19 of 36 patients; 53%; P<0.001).
By histopathologic (morphologic and immunhistochemical) examination of BM biopsies, BM infiltration with lymphoma cells was seen in 19 patients (CS IV disease by definition). Clonal IgR were found in PB/BM of 14 (74%) of these cases. In two patients, Ig rearrangements were also undetectable in the corresponding tumor biopsy probably due to a methodical problem; in one patient, the tumor biopsy was not available for analysis. IgR were found in additional 21 of 130 (16%; BM involvement vs no BM involvement, P<0.001) patients who showed no evidence of lymphoma infiltration in good quality-bone marrow biopsies. In six patients in whom BM histology was inconclusive, IgR were not detected. IgR were detected in five of 17 (29%) patients with CS IV but without BM involvement.
There was also a significant association between the detection of Ig rearrangements in PB/BM and the IPI risk groups (P<0.001).
Correlation of clonal immunoglobulin rearrangements in PB/BM with clinical outcome (Table 2)
The overall CR rate was 66% (103 of 155). IgR-positive cases had a significantly lower CR rate (18 of 35, 51%) than IgR-negative patients (85 of 120, 71%; P=0.042). Applying a logistic regression model, the odds to achieve a CR was 0.44 (CI 0.20–0.94, P=0.035) for the former compared to the latter group. However, the additional prognostic impact of IgR in a multiple model with BM histology does not reach statistical significance (OR 0.99, CI 0.35–2.80, P=0.990), whereas BM histology showed a significant influence on the CR rate (OR 0.14, CI 0.05–0.43, P<0.001). In a multiple model adjusting for IPI score, both IgR and BM histology are not significant (P=0.181 and P=0.417, respectively). There was no statistical difference in the relapse rate between patients with and without Ig rearrangements (P=0.384).
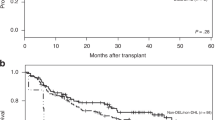
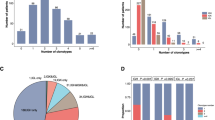
The estimated OAS at 5 years in the group with and without IgR in PB/BM is 25% (CI 7–43%) and 66% (CI 54–78%), respectively (Figure 1). The estimated OAS at 5 years for patients with negative BM histology and negative PCR results is 66% (CI 53–79%), for patients with negative BM histology but positive PCR results 37% (CI 12–63%), and for patients with positive BM histology 12% (CI 0–33%; Figure 2a). The hazard ratio for IgR is 3.78 (CI 2.11–6.77, P<0.001) in the univariate Cox model and 2.39 (CI 1.15–4.99, P=0.020) when used as a prognostic factor in addition to BM histology, the latter showing a hazard ratio of 5.39 (CI 2.60–11.20, P<0.001). However, both markers do not remain significant when adjusted for IPI score (P=0.357 and P=0.579, respectively).
Kaplan–Meier curves of overall survival of patients with diffuse large B-cell lymphoma according to bone marrow histology and the detection of IgR in PB/BM. (a) OAS analysis of all patients with successful histopathologic analysis of BM biopsies. (b) OAS analysis after exclusion of patients in whom diagnostic tumor samples were not available or not informative. BM, bone marrow; IgR, immunoglobulin rearrangement; neg, negative; pos, positive.
The progression-free survival at 5 years was 55% (CI 26–83%) in the IgR-positive group and 63% (CI 48–79%) in the IgR-negative group, the OR in the univariate Cox model was 1.76 (CI 0.71–4.39, P=0.225). Both, IgR and BM histology are not significant, whether or not adjusted for IPI scores (all Ps >0.400).
When the OAS of patients without IgR in PB/BM was investigated for only those patients who showed IgR in the diagnostic tumor samples (exclusion of patients in whom diagnostic tumor samples were not available (n=35) or not informative (n=15), Figure 2b), the OAS at 5 years in the three groups is estimated as 67% (CI 53–81%), 37% (CI 12–63%), and 0%, respectively. The hazard ratio for IgR in this case is 3.56 (CI 1.83–6.91, P<0.001) in the univariate model and 2.28 (CI 1.03–5.03, P=0.041) when used as a prognostic factor in addition to BM histology, the latter showing a hazard ratio of 7.97 (CI 3.37–18.86, P<0.001). Again, both markers do not remain significant when adjusted for IPI score (P=0.467 and P=0.158, respectively).
Discussion
We report on the contribution of PCR amplification of IgR to staging in DLBCL. Two previous reports have used Southern blotting for stage determination in small (11 high grade) or less well defined series (41 intermediate and high grade) of lymphoma patients.14,15 Both studies suggested that the molecular detection of circulating lymphoma cells may be of prognostic value. This is the first large study investigating the more sensitive and widely applicable PCR amplification of IgR, which was also controlled by analysis of diagnostic tumor biopsies in the majority of patients.
Clonal IgR were detected in 86% of tumor specimen which is within the range of published results.11,12,30 None of the patients who could be analyzed in the diagnostic tumor sample had an additional IgR in the PB or BM, indicating the specificity of the method. This is in contrast to the poor reliability of PCR amplification of TCR genes in mature B-cell lymphomas.34 However, the detection rate of clonal immunoglobulin rearrangements might be further increased by PCR analysis of additional PCR targets in future studies. The junctional diversity of immunoglobulin kappa deleting element (Kde) rearrangements, which are ‘end-stage’ rearrangements and cannot undergo further rearrangements, is a stable target for reliable detection of clonal cell growth and minimal residual disease.35,36 In addition, incomplete DH-JH gene rearrangements can be perceived as a supplementary PCR target particularly for patients who do not have other clone-specific rearrangements.37
The major goal of our study was to determine the additional information on the extent of lymphoma dissemination provided by PCR. We found clonal IgR in 21 of 130 (16%) patients with normal histology of good quality BM biopsies, suggesting an improved staging by PCR analysis. Overall, patients with positive PCR results had a significantly poorer CR rate and OAS.
Another key questions was whether molecular staging could increase the prognostic value of histopathological staging. Patients with positive IgR in PB/BM but without morphologically detectable BM involvement indeed had a significantly poorer OAS than patients with positive BM histology (Figure 2). The difference in OAS was 29% at 5 years (66 vs 37%). This was still true when the survival analysis was corrected by the exclusion of those patients in whom diagnostic tumor samples were not available or not informative. Our results also indicate that patients with morphologically detectable infiltration of the bone marrow have a significantly poorer OAS than patients with positive PCR but normal histology (0 vs 37%). Therefore, molecular analysis of BM or PB cannot replace histologic examination of the BM. This is supported by the fact that three patients had a positive PCR result in the BM but not in the PB.
Recent gene-expression profiling of DLBCL has shown that this single diagnostic category includes more than one molecularly and clinically distinct disease. DLBCL consists of at least three different gene-expression subgroups, known as germinal-center B-cell-like (GCB), activated B-cell-like (ABC) and type 3.38,39 These gene-expression subgroups differ by the expression of more than 1000 genes and seem to be derived from different stages of normal B-cell differentiation. The molecular distinction between subgroups of DLBCL is important because the subgroups have distinct mechanisms of malignant transformation and differ in their ability to be cured by the multiagent chemotherapy that is used at present. Patients with GCB DLBCL have the most favorable cure rate (60% 5-year survival), whereas patients with ABC and type 3 DLBCL have 5-year survival rates of only 36 and 39%, respectively. The improvement of staging by molecular methods, which results in a shift of patients from localized CS I-III to CS IV with diffuse or disseminated involvement of bone marrow and therefore worsening of prognosis, might affect particularly patients with expression profiles that are associated with poor response to chemotherapy. This effect seems to be noticed primarily in the OAS analysis which includes the whole group of DLBCL patients, whereas the affect of an improved staging on progression-free survival analysis which only comprises patients with a relatively good response to chemotherapy might not be seen until more patients with DLBCL can be studied in consideration of different gene-expression profiles. Altogether, our data demonstrate an important contribution of molecular staging in DLBCL. The results of OAS analysis indicate that patients without histopathological BM involvement can be subdivided into two prognostic groups by molecular analysis of IgR in circulating blood and/or BM cells and that there is also a prognostic threshold between histologically verified and ‘molecular’ stage IV disease.
References
Fisher RI, Gaynor ER, Dahlberg S, Oken MM, Grogan TM, Mize EM et al. Comparison of a standard regimen (CHOP) with three intensive chemotherapy regimens for advanced non-Hodgkin's lymphoma. N Engl J Med 1993; 328: 1002–1006.
Miller TP, Dahlberg S, Cassady JR, Adelstein DJ, Spier CM, Grogan TM et al. Chemotherapy alone compared with chemotherapy plus radiotherapy for localized intermediate- and high-grade non-Hodgkin's lymphoma. N Engl J Med 1998; 339: 21–26.
A predictive model for aggressive non-Hodgkin's lymphoma. The International Non-Hodgkin's Lymphoma Prognostic Factors Project. N Engl J Med 1993; 329: 987–994.
Langerak AW, van Krieken JH, Wolvers-Tettero IL, Kerkhof E, Mulder AH, Vrints LW et al. The role of molecular analysis of immunoglobulin and T cell receptor gene rearrangements in the diagnosis of lymphoproliferative disorders. J Clin Pathol 2001; 54: 565–567.
Pongers-Willemse MJ, Seriu T, Stolz F, d'Aniello E, Gameiro P, Pisa P et al. Primers and protocols for standardized detection of minimal residual disease in acute lymphoblastic leukemia using immunoglobulin and T cell receptor gene rearrangements and TAL1 deletions as PCR targets: report of the BIOMED-1 CONCERTED ACTION: investigation of minimal residual disease in acute leukemia. Leukemia 1999; 13: 110–118.
van Dongen JJ, Seriu T, Panzer-Grumayer ER, Biondi A, Pongers-Willemse MJ, Corral L et al. Prognostic value of minimal residual disease in acute lymphoblastic leukaemia in childhood. Lancet 1998; 352: 1731–1738.
Cave H, van der Werff ten Bosch J, Suciu S, Guidal C, Waterkeyn C, Otten J et al. Clinical significance of minimal residual disease in childhood acute lymphoblastic leukemia. European Organization for Research and Treatment of Cancer – Childhood Leukemia Cooperative Group. N Engl J Med 1998; 339: 591–598.
Kuppers R, Klein U, Hansmann ML, Rajewsky K . Cellular origin of human B-cell lymphomas. N Engl J Med 1999; 341: 1520–1529.
Stevenson F, Sahota S, Zhu D, Ottensmeier C, Chapman C, Oscier D et al. Insight into the origin and clonal history of B-cell tumors as revealed by analysis of immunoglobulin variable region genes. Immunol Rev 1998; 162: 247–259.
Derksen PW, Langerak AW, Kerkhof E, Wolvers-Tettero IL, Boor PP, Mulder AH et al. Comparison of different polymerase chain reaction-based approaches for clonality assessment of immunoglobulin heavy-chain gene rearrangements in B-cell neoplasia. Mod Pathol 1999; 12: 794–805.
Aubin J, Davi F, Nguyen-Salomon F, Leboeuf D, Debert C, Taher M et al. Description of a novel FR1 IgH PCR strategy and its comparison with three other strategies for the detection of clonality in B cell malignancies. Leukemia 1995; 9: 471–479.
Gong JZ, Zheng S, Chiarle R, Wolf-Peeters C, Palestro G, Frizzera G et al. Detection of immunoglobulin kappa light chain rearrangements by polymerase chain reaction. An improved method for detecting clonal B-cell lymphoproliferative disorders. Am J Pathol 1999; 155: 355–363.
Seriu T, Hansen-Hagge TE, Stark Y, Bartram CR . Immunoglobulin kappa gene rearrangements between the kappa deleting element and Jkappa recombination signal sequences in acute lymphoblastic leukemia and normal hematopoiesis. Leukemia 2000; 14: 671–674.
Hiorns LR, Nicholls J, Sloane JP, Horwich A, Ashley S, Brada M . Peripheral blood involvement in non-Hodgkin's lymphoma detected by clonal gene rearrangement as a biological prognostic marker. Br J Cancer 1994; 69: 347–351.
Horning SJ, Galili N, Cleary M, Sklar J . Detection of non-Hodgkin's lymphoma in the peripheral blood by analysis of antigen receptor gene rearrangements: results of a prospective study. Blood 1990; 75: 1139–1145.
Harris NL, Jaffe ES, Stein H, Banks PM, Chan JK, Cleary ML et al. A revised European–American classification of lymphoid neoplasms: a proposal from the International Lymphoma Study Group. Blood 1994; 84: 1361–1392.
Harris NL, Jaffe ES, Diebold J, Flandrin G, Muller-Hermelink HK, Vardiman J et al. The World Health Organization classification of neoplasms of the hematopoietic and lymphoid tissues: report of the Clinical Advisory Committee meeting – Airlie House, Virginia, November, 1997. Hematol J 2000; 1: 53–66.
Armitage JO, Dick FR, Corder MP, Garneau SC, Platz CE, Slymen DJ . Predicting therapeutic outcome in patients with diffuse histiocytic lymphoma treated with cyclophosphamide, adriamycin, vincristine and prednisone (CHOP). Cancer 1982; 50: 1695–1702.
Fisher RI, DeVita Jr VT, Hubbard SM, Longo DL, Wesley R, Chabner BA et al. Diffuse aggressive lymphomas: increased survival after alternating flexible sequences of proMACE and MOPP chemotherapy. Ann Intern Med 1983; 98: 304–309.
Longo DL, DeVita Jr VT, Duffey PL, Wesley MN, Ihde DC, Hubbard SM et al. Superiority of ProMACE-CytaBOM over ProMACE-MOPP in the treatment of advanced diffuse aggressive lymphoma: results of a prospective randomized trial. J Clin Oncol 1991; 9: 25–38.
Pavlovsky S, Santarelli MT, Erazo A, Diaz Maqueo JC, Somoza N, Lluesma GM et al. Results of a randomized study of previously-untreated intermediate and high grade lymphoma using CHOP versus CNOP. Ann Oncol 1992; 3: 205–209.
O'Reilly SE, Connors JM, Howdle S, Hoskins P, Klasa R, Klimo P et al. In search of an optimal regimen for elderly patients with advanced-stage diffuse large-cell lymphoma: results of a phase II study of P/DOCE chemotherapy. J Clin Oncol 1993; 11: 2250–2257.
Szanto I, Fleischmann T, Eckhardt S . Treatment of resistant Hodgkin's disease with CCNU, etoposide and prednimustine (CEP). Oncology 1991; 48: 456–458.
Cheson BD, Horning SJ, Coiffier B, Shipp MA, Fisher RI, Connors JM et al. Report of an international workshop to standardize response criteria for non-Hodgkin's lymphomas. NCI Sponsored International Working Group. J Clin Oncol 1999; 17: 1244.
Sambrook J, Fritsch EF, Maniatis T . Molecular Cloning. A Laboratory Manual. New York: Cold Spring Harbor Laboratory Press, 1989.
Frank TS, Svoboda-Newman SM, Hsi ED . Comparison of methods for extracting DNA from formalin-fixed paraffin sections for nonisotopic PCR. Diagn Mol Pathol 1996; 5: 220–224.
Sioutos N, Bagg A, Michaud GY, Irving SG, Hartmann DP, Siragy H et al. Polymerase chain reaction versus Southern blot hybridization. Detection of immunoglobulin heavy-chain gene rearrangements. Diagn Mol Pathol 1995; 4: 8–13.
Liang R, Chan V, Chan TK, Wong T, Chiu E, Lie A et al. Detection of immunoglobulin gene rearrangement in lymphoid malignancies of B-cell lineage by seminested polymerase chain reaction gene amplification. Am J Hematol 1993; 43: 24–28.
Trainor KJ, Brisco MJ, Wan JH, Neoh S, Grist S, Morley AA . Gene rearrangement in B- and T-lymphoproliferative disease detected by the polymerase chain reaction. Blood 1991; 78: 192–196.
Fodinger M, Winkler K, Mannhalter C, Chott A . Combined polymerase chain reaction approach for clonality detection in lymphoid neoplasms. Diagn Mol Pathol 1999; 8: 80–91.
Kwok S, Higuchi R . Avoiding false positives with PCR. Nature 1989; 339: 237–238.
Lo YM, Mehal WZ, Fleming KA . False-positive results and the polymerase chain reaction. Lancet 1988; 2: 679.
Kaplan EL, Meier P . Nonparametric estimation from incomplete observations. J Am Stat Assoc 1958; 53: 457–481.
Macintyre EA, Delabesse E . Molecular approaches to the diagnosis and evaluation of lymphoid malignancies. Semin Hematol 1999; 36: 373–389.
Beishuizen A, de Bruijn MAC, Pongers-Willemse MJ, Verhoeven M-AJ, van Wering ER, Hählen K et al. Heterogeneity in junctional regions of immunoglobulin kappa deleting element rearrangements in B cell leukemias: a new molecular target for detection of minimal residual disease. Leukemia 1997; 11: 2200–2207.
Van der Velden VHJ, Willemse MJ, van der Schoot CE, Hählen K, van Wering ER, van Dongen JJM . Immunoglobulin kappa deleting element rearrangements in precursor-B acute lymphoblastic leukemia are stable targets for detection of minimal residual disease by real-time quantitative PCR. Leukemia 2002; 16: 928–936.
Szczepanski T, Willemse MJ, van Wering ER, van Weerden JF, Kamps WA, van Dongen JJM . Precursor-B-ALL with DH-JH gene rearrangements have an immature immunophenotype with a high frequency of oligoclonality and hyperdiploidy of chromosome 14. Leukemia 2001; 15: 1415–1423.
Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 2000; 403: 503–511.
Rosenwald A, Wright G, Chan WC, Connors JM, Campo E, Fisher RI et al. Lymphoma/Leukemia Molecular Profiling Project. The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N Engl J Med 2002; 346: 1937–1947.
Acknowledgements
We are indebted to Susanne Hagmann for expert assistance with data management, and to Monika Heimbach and Helmut Sommer for excellent technical assistance. This study was supported by the FWF Grant P 13984-GEN and the Center of Molecular Medicine of the Austrian Academy of Sciences, and by Grant NB 9964 of the Austrian National Bank.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mitterbauer-Hohendanner, G., Mannhalter, C., Winkler, K. et al. Prognostic significance of molecular staging by PCR-amplification of immunoglobulin gene rearrangements in diffuse large B-cell lymphoma (DLBCL). Leukemia 18, 1102–1107 (2004). https://doi.org/10.1038/sj.leu.2403376
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2403376
Keywords
This article is cited by
-
Prognostic markers for immunodeficiency-associated primary central nervous system lymphoma
Journal of Neuro-Oncology (2019)
-
Clinical significance of cytogenetic aberrations in bone marrow of patients with diffuse large B-cell lymphoma: prognostic significance and relevance to histologic involvement
Journal of Hematology & Oncology (2013)
-
Bone marrow involvement is predictive of infusion-related reaction during rituximab administration in patients with B cell lymphoma
Supportive Care in Cancer (2013)
-
Routine use of ancillary investigations in staging diffuse large B-cell lymphoma improves the International Prognostic Index (IPI)
Journal of Hematology & Oncology (2009)