Abstract
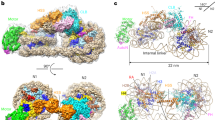
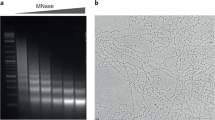
RECENT evidence indicates that a large portion of the DNA of higher organisms is organised in compact nucleoprotein structures. Initial studies on nuclease digestion of chroma-tin showed that about half the nuclear DNA is present in the form of small resistant chromatin fragments1–3. Subsequent studies have shown that the sites of nuclease digestion are regularly spaced, and that, at the limit of digestion, the majority of the DNA that remains is in the form of relatively homogeneous small fragments, which vary from about 120 to 200 nucleotide pairs in length4–7. This concept of small repeating resistant structures in chromatin is further supported by electron microscope studies on chromatin fibres spread from hypotonically treated nuclei8 and from isolated chromatin9.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Clark, R. J., and Felsenfeld, G., Nature new Biol., 229, 101–106 (1971).
Rill, R., and Van Holde, K. E., J. biol. Chem., 248, 1080–1083 (1973).
Hewish, D. R., and Burgoyne, L. A., Biochem. biophys. Res. Commun., 52, 504–510 (1973).
Oosterhof, D. K., Hozier, J. C., and Rill, R. L., Proc. natn. Acad. Sci. U.S.A., 72, 633–637 (1975).
Noll, M., Nature, 251, 249–251 (1974).
Lohr, D., and Van Holde, K. E., Science, 186, 165–166 (1975).
Shaw, B. R., Corden, J. L., Sahasrabuddhe, C. G., and Van Holde, K. E., Biochem. biophys. Res. Commun., 61, 1193–1198 (1974).
Olins, A. L., and Olins, D. E., Science, 183, 330–332 (1974).
Oudet, P., Gross-Bellard, A., and Chambon, P., Cell, 4, 282–290 (1975).
Mazrimas, J. A., and Hatch, F. T., Nature new Biol., 240, 102–105 (1972).
Mazrimas, J. A., and Hatch, F. T., Exp. Cell. Res., 63, 462–466 (1970).
Hatch, F. T., and Mazrimas, J. A., Nucleic Acids Res., 1, 559–575 (1974).
Bostock, C. J., and Christie, S., Chromosoma, 48, 73–87 (1974).
Wray, W., in Methods in Cell Biology, 6 (edit. by Prescott, D. M.), 283–306 and 307–315 (Academic, New York and London, 1973).
Maio, J. J., and Schildkraut, C. L., J. molec. Biol., 24, 29–39 (1967).
Bonner, W. M., and Laskey, R. A., Eur. J. Biochem., 46, 83–88 (1974).
Southern, E. M., J. molec. Biol., 94, 51–69 (1975).
Nathans, D., and Smith, H. O., A. Rev. Biochem., 44, 273–293 (1975).
Marmur, J. J., J. molec. Biol., 3, 208–218 (1961).
Crothers, D. M., Kallenbach, N. R., and Zimm, B. H., J. molec. Biol., 11, 802–820 (1965).
Wingert, L., and Von Hippel, P. H., Biochim. Biophys. Acta, 157, 114–126 (1968).
Van Holde, K. E., Sahasrabuddhe, C. G., and Shaw, B. R., Nucleic Acids Res., 1, 1579–1586 (1974).
Baldwin, J. P., Boseley, P. G., and Bradbury, E. M., Nature, 253, 245–249 (1975).
Hyde, J. E., and Walker, I. O., Nucleic Acid Res., 2, 405–421 (1975).
Gilmour, S., New Scient., 14, February, 390–392 (1975).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
BOSTOCK, C., CHRISTIE, S. & HATCH, F. Accessibility of DNA in condensed chromatin to nuclease digestion. Nature 262, 516–519 (1976). https://doi.org/10.1038/262516a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/262516a0
This article is cited by
-
The basis of chromatin fiber assembly within chromosomes studied by histone-DNA crosslinking followed by trypsin digestion
Chromosoma (1980)
-
Preferential occurrence of sister chromatid exchanges at heterochromatin-euchromatin junctions in the wallaby and hamster chromosomes
Chromosoma (1979)
-
The distribution of mitomycin C-induced sister chromatid exchanges in the euchromatin and heterochromatin of the Indian Muntjac
Chromosoma (1977)
-
Gene control in eukaryotes and thec-value paradox “Excess” DNA as an impediment to transcription of coding sequences
Journal of Molecular Evolution (1976)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.