Abstract
Most multicellular organisms use steroids as signalling molecules for physiological and developmental regulation. Two different modes of steroid action have been described in animal systems: the well-studied gene regulation response mediated by nuclear receptors1,2, and the rapid non-genomic responses mediated by proposed membrane-bound receptors3,4. Plant genomes do not seem to encode members of the nuclear receptor superfamily5. However, a transmembrane receptor kinase, brassinosteroid-insensitive1 (BRI1), has been implicated in brassinosteroid responses6,7. Here we show that BRI1 functions as a receptor of brassinolide, the most active brassinosteroid. The number of brassinolide-binding sites and the degree of response to brassinolide depend on the level of BRI1 protein. The brassinolide-binding activity co-immunoprecipitates with BRI1, and requires a functional BRI1 extracellular domain. Moreover, treatment of Arabidopsis seedlings with brassinolide induces autophosphorylation of BRI1, which, together with our binding studies, shows that BRI1 is a receptor kinase that transduces steroid signals across the plasma membrane.
Similar content being viewed by others
Main
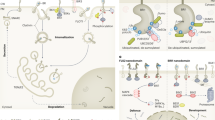
Brassinosteroids (BRs) are involved in a wide range of plant developmental processes8. Mutant plants deficient in BR biosynthesis, such as det2, show phenotypes of dwarfism, delayed senescence, reduced fertility, and light-independent development9,10. Mutations in the BRI1 gene cause BR-insensitivity and morphological phenotypes nearly identical to BR biosynthetic mutants, suggesting an important and specific role for BRI1 in BR perception or signal transduction11,12. The BRI1 gene encodes a receptor kinase that has an extracellular domain containing 25 leucine-rich repeats (LRRs), which are interrupted by a 70-amino-acid island, a transmembrane domain, and a cytoplasmic kinase domain with serine/threonine specificity6,13. The structure of BRI1 and its plasma membrane localization13 support the hypothesis that BRI1 interacts with an extracellular ligand, which is either BR itself or a secondary signal generated by BR perception, and the signal is transduced through the kinase. Consistent with this hypothesis, recent studies using a chimaeric receptor approach showed that the extracellular domain of BRI1 could confer brassinolide (BL) responsiveness to the intracellular kinase domain of a heterologous LRR kinase7.
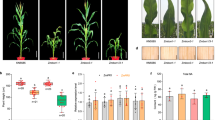
To test whether BL is the ligand that directly activates the BRI1 receptor kinase, we first analysed the effect of overexpression of BRI1 on BL binding activity in membrane fractions. Transgenic Arabidopsis plants overexpressing a BRI1–GFP fusion protein13 (GFP, green fluorescent protein) showed reduced inhibition of hypocotyl growth by a BR biosynthesis inhibitor14 (Fig. 1a, b). They also had longer petioles, similar to plants overexpressing the BR biosynthetic enzyme DWF415 (Z.W. and J.C., unpublished data) (Fig. 1c). These phenotypes are consistent with the interpretation that overexpression of the BRI1–GFP protein increases the response of Arabidopsis to BRs. We observed a dramatic increase of BL binding activity in the membrane fractions of the BRI1–GFP transgenic plants (Fig. 1d). The increase of binding was due to an increase of binding sites (maximum number of binding sites, Bmax = 2.66 pmol per mg membrane protein compared to 0.23 pmol per mg membrane protein), with similar binding affinities (dissociation constant Kd = 7.4 ± 0.9 nM compared to 10.8 ± 3.2 nM) (Fig. 1e). Such Kd values are consistent with physiological concentrations of BL16,17 and coincide with the BL concentration that induces 50% of the maximum growth response in BL-deficient mutants9.
a, Proteins from wild-type and BRI1–GFP transgenic plants probed with anti-BRI1N antibody after western blotting. N, a cleaved fragment of BRI1's extracellular domain. Asterisks, non-specific bands. b, Wild-type (WT) and BRI1–GFP plants grown on 2 µM brassinazole in the dark for 6 d. c, A wild-type plant, a BRI1–GFP transgenic plant, and a mutant plant overexpressing the DWF4 gene (35SE-DWF4) grown for 45 d in cycles of 9 h light/15 h dark. d, Specific [3H]-BL binding to microsomal fractions of wild-type (WT) and BRI1–GFP plants was determined by subtracting the binding in the presence of 100-fold unlabelled BL from the total binding in the absence of cold competitor. Representative data of one of three repeat experiments are shown. e, Scatchard plot of the binding data in d. The Kd values were calculated from data of three experiments, with correlation coefficient R2 = 0.998 for BRI1–GFP and 0.983 for wild-type samples. f, Competition for [3H]-BL binding to membrane fractions of BRI1–GFP plants by brassinolide, castasterone, ecdysone and 2,3,22,23-O-tetramethylbrassinolide. Structures of the competitors are shown.
We determined the specificity of the BL binding activity by comparing the relative binding affinity for several steroid compounds in binding competition assays (Fig. 1f). Binding of [3H]-BL to the membrane fraction of BRI1–GFP plants was effectively competed by unlabelled BL (50% inhibition concentration, IC50, 80 nM), less effectively by castasterone (IC50, 340 nM), and not competed by 2,3,22,23-O-tetramethylbrassinolide (Me-BL; IC50 > 10 µM) and ecdysone (IC50 > 10 µM). The relative binding affinity (the ratio between IC50 of a competitor and that of BL) of castasterone is about 4–5 times lower than BL, and this is consistent with castasterone being about 5 times less active than BL in bioassays18. The lack of competition by Me-BL and ecdysone is consistent with their lack of biological activity in plants19 (S.F., unpublished results). Such a specificity and high affinity for biologically active BRs indicate that this BL binding activity accounts for the BL-induced biological responses.
A specific BL binding activity was detected after the BRI1–GFP proteins were immunoprecipitated using anti-GFP antibodies (Fig. 2a, b). No specific BL binding activity was immunoprecipitated from wild-type Arabidopsis plants using the same antibodies (Fig. 2a), indicating that the BL binding activity is specific to the BRI1–GFP protein. The immunoprecipitated binding activity has a similar dissociation constant (Kd = 15.2 ± 5 nM) as determined for membrane fractions (Fig. 2b). These results show that BRI1 either binds BL directly or is a limiting component of a receptor complex for BL in plant cells.
a, Proteins immunoprecipitated with anti-GFP antibodies from extracts of wild-type or BRI1–GFP plants were assayed for [3H]-BL binding activity in the absence (open bars) or presence (filled bars) of 5 µM unlabelled BL. No protein, binding assays with protein A beads as control. b, Specific and saturation [3H]-BL binding to immunoprecipitated BRI1–GFP protein. c, Mutations in the extracellular domain of BRI1, but not the kinase domain, reduce BL binding. Specific [3H]-BL binding to microsomal fractions of wild type (WT), det2, bri1-6, bri1-116, bri1-104 and bri1-102 mutant plants. Data were normalized to the relative BRI1 protein levels determined by quantitative western blotting (d), except for bri1-116. d, Protein immunoblot showing the BRI1 protein levels in the membrane fractions used in the binding assays of c. Varying loading of the wild-type sample was used to generate a standard curve, which was used to determine the relative level of BRI1 in the bri1 mutant samples (6 µl per lane).
Mutations in large numbers of bri1 alleles implicate the functional importance of the cytoplasmic kinase domain and the 70-amino-acid island of BRI1's extracellular domain6,13,17. We found that bri1 mutants with missense mutations in the kinase domain (bri1-104, Ala1031Thr) or in a region of the extracellular domain near the transmembrane domain (bri1-102, Thr750Ile) have BL binding activities similar to wild type and the biosynthetic mutant det2 (Fig. 2c, d). In contrast, a missense mutation (bri1-6, Gly644Asp) and a nonsense mutation that results in a truncated protein at position Gln 583 (bri1-116), both in the 70-amino-acid island region, greatly reduced the BL binding activity (Fig. 2c, d). These results provide direct evidence that the 70-amino-acid island region of BRI1's extracellular domain is required for BL binding to the receptor on the cell membrane.
We tested whether BL-binding activates BRI1's kinase. Receptor activation often involves auto-phosphorylation, which can lead to a change of mobility in SDS–polyacrylamide gel electrophoresis (PAGE). Arabidopsis seedlings grown in the presence of the BR biosynthetic inhibitor brassinazole were treated with BL and analysed by immunoblotting (Fig. 3). Treatment of wild-type seedlings with 1 µM BL for 1 h caused a shift of BRI1 from a faster to a slower migrating band, compared with an untreated sample or a sample treated with mock solution (Fig. 3a). Phosphatase treatment of the BL-treated samples shifted the slower band back to the fast migrating band, suggesting that the shift of mobility represents BRI1 phosphorylation (Fig. 3b). We also observed such a BL-induced BRI1 mobility shift in the BL biosynthetic mutant det2, but not in the bri1-117 mutant (Fig. 3c), which contains a mutation that abolishes BRI1's in vitro kinase activity (data not shown). Our data indicate that BL induction of BRI1 phosphorylation requires the kinase activity of BRI1, suggesting that BL-binding induces autophosphorylation of BRI1.
a, Five-day-old wild-type seedlings grown in the dark on medium containing 1 µM brassinazole were untreated (-), treated with 1 µM BL in water (BL), or with water only (H2O) for 1 h. Proteins were analysed by 4% SDS–PAGE, blotted, and probed with anti-BRI1N antibody. b, Proteins of the BL-treated wild-type sample were treated with alkaline phosphatase (CIP), and analysed by western blotting as in a. c, A mutation in the kinase domain abolishes BL-activation of BRI1 phosphorylation. The det2 and bri1-117 mutant seedlings were treated and analysed as in a. N, cleavage product of BRI1's extracellular domain.
Our identification of the receptor kinase BRI1 as a plant steroid receptor illustrates the function of a member of the largest family of receptor kinases in Arabidopsis. The Arabidopsis genome sequence revealed 174 LRR-receptor kinases5, of which only a few are known for their biological functions20,21,22, and only one, CLV1, has been characterized at the biochemical level20. The mechanism by which BRI1 kinase is activated by ligand binding may be shared by other LRR-receptor kinases, as suggested by the BR activation of a BRI1-Xa21 chimaeric receptor7. However, BRI1 seems to differ from CLV1, which has recently been shown to require its own kinase activity for binding to its peptide ligand20.
Our results also reveal a new mechanism of steroid signalling. Steroid hormones are generally known to pass freely across plasma membranes into animal cells, where they bind to members of the nuclear receptor superfamily of ligand-dependent transcription factors1,2. In contrast, the Arabidopsis genome does not seem to encode members of this family of proteins5. The near identical phenotypes of bri1 to BR-biosynthetic mutants and our results presented here indicate that plants perceive steroids at the cell surface and that BRI1 is likely to be the primary BR receptor in Arabidopsis. A similar BL signalling mechanism is apparently conserved in other plants, as a homologue of BRI1 was shown recently to be required for BR responses in rice23. Such a cell-surface signalling mechanism may not be unique to plant steroids. In fact, membrane-initiated steroid responses have been observed in many animal systems, and signalling molecules such as calcium, inositol phosphates, cyclic AMP, G proteins and various kinases have been implicated3. However, little is known about the membrane-bound steroid receptors that initiate these signalling cascades in animal cells4. Considering the conservation of the steroid biosynthetic enzymes24 and similar roles of steroids in growth and differentiation in plants and animals, it will be interesting to see if a similar membrane-based steroid signalling mechanism exists in both the animal and plant kingdoms.
Methods
BL-binding assays
Tritium-labelled BL was made by American RadioChemicals using tritium reduction of 25,26-dehydrobrassinolide25. The specific activity of [3H]-BL was estimated to be 50 Ci mmol-1, and the correct structure was confirmed by tritium NMR analysis. Biological activity of the [3H]-BL was determined by rescue of the det2 mutant (data not shown). Plant microsomal fractions were prepared from Arabidopsis seedlings grown in a cycle of 9 h light/15 h dark for about 6 weeks, following a protocol described previously7. Membrane pellets were resuspended at a protein concentration of 2 mg ml-1 in BL-binding buffer (0.25 M mannitol, 10 mM Tris-2-[N-morpholino] ethanesulphonic acid (MES), pH 5.7, 5 mM MgCl2, 0.1 mM CaCl2, and protease inhibitor cocktail (Sigma). Each BL binding assay contains 50 µl membrane suspensions, 50 nM or indicated amount of [3H]-BL, without or with 100-fold excess unlabelled BL or indicated amount of unlabelled steroids, 1 mg ml-1 BSA and BL-binding buffer in 100 µl total volume. The binding reactions were incubated for 1 h or indicated time at 25 °C. The bound and free [3H]-BL were separated by filtering the mixture through a glass-fibre filter (Whatman, GF/F) and washing with 10 ml ice-cold BL-binding buffer, and were quantified by scintillation counting. Binding kinetic studies showed that the specific BL binding reaches equilibrium within 15 min of incubation; the binding is highly reversible and 50% competition by unlabelled BL was achieved in less than 1 min (data not shown). For binding assays with immunoprecipitated proteins, proteins were extracted with BRI1 extraction buffer (2 ml per g tissue) (50 mM Tris-HCl, pH 7.5, 10 mM NaCl, 5 mM EDTA, 1% Triton X-100, and protease inhibitor cocktail), and GFP-tagged proteins were immunoprecipitated using anti-GFP antibodies (Molecular Probes, 1 µl per ml extract) and protein A agarose beads (10 µl per ml extract, Pierce). BL binding assays with immunoprecipitant/agarose beads were under the same conditions as for membrane fractions. Specific binding was determined by subtracting the binding in the presence of 100 fold unlabelled BL from total binding. Binding data were analysed and plotted using KaleidaGraph software (Synergy Software).
Immunoblotting and assays for BL-induced BRI1 phosphorylation
A peptide containing the first 106 amino acids of BRI1 (excluding the signal peptide) was expressed in Escherichia coli and purified as a maltose binding protein (MBP) fusion. This MBP–BRI1N fusion protein was used as an antigen for generating the anti-BRI1N antibodies in rabbits, and for affinity purification of the antibodies. The identities of the BRI1-containing bands detected by the anti-BRI1N antibodies on immunoblots were determined by comparing wild type, BRI1–GFP and bri1-116 (a nonsense mutant) samples. To test BL induction of BRI1 phosphorylation, Arabidopsis seedlings were grown on MS medium plates containing 1 µM brassinazole in the dark for 4 d, submerged in water or in 1 µM BL solution for 1 min, then put back on MS plates without or with 1 µM BL for 1 h. Samples were analysed by SDS–PAGE using 4% gels and western blotting as described above7. For phosphatase treatment, SDS was removed from protein samples using the SDS-OUT kit (Pierce), and the proteins were then treated with alkaline phosphatase (Boehringer Mannheim) under conditions recommended by the manufacturer and described elsewhere26.
References
Beato, M., Herrlich, P. & Schutz, G. Steroid hormone receptors: many actors in search of a plot. Cell 83, 851–857 (1995).
Mangelsdorf, D. J. et al. The nuclear receptor superfamily: The second decade. Cell 83, 835–839 (1995).
Wehling, M. Specific, nongenomic actions of steroid hormones. Annu. Rev. Physiol. 59, 365–393 (1997).
Schmidt, B. M. et al. Rapid, nongenomic steroid actions: A new age? Front. Neuroendocrinol. 21, 57–94 (2000).
The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Li, J. & Chory, J. A putative leucine-rich repeat receptor kinase involved in brassinosteroid signal transduction. Cell 90, 929–938 (1997).
He, Z. et al. Perception of brassinosteroids by the extracellular domain of the receptor kinase BRI1. Science 288, 2360–2363 (2000).
Mandava, N. B. Plant growth-promoting brassinosteroids. Annu. Rev. Plant Physiol. Plant Mol. Biol. 39, 23–52 (1988).
Li, J., Nagpal, P., Vitart, V., McMorris, T. C. & Chory, J. A role for brassinosteroids in light-dependent development of Arabidopsis. Science 272, 398–401 (1996).
Wang, Z.-Y. & Chory, J. in Recent Advances in Phytochemistry Vol. 34, Evolution of Metabolic Pathways (eds Romeo, J. T., Ibrahim, R., Varin, L. & DeLuca, V.) 409–431 (Elsevier Science, Oxford, 2000).
Clouse, S. D., Langford, M. & McMorris, T. C. A brassinosteroid-insensitive mutant in Arabidopsis thaliana exhibits multiple defects in growth and development. Plant Physiol. 111, 671–678 (1996).
Schumacher, K. & Chory, J. Brassinosteroid signal transduction: still casting the actors. Curr. Opin. Plant Biol. 3, 79–84 (2000).
Friedrichsen, D. M., Joazeiro, C. A., Li, J., Hunter, T. & Chory, J. Brassinosteroid-insensitive-1 is a ubiquitously expressed leucine-rich repeat receptor Serine/Threonine kinase. Plant Physiol. 123, 1247–1256 (2000).
Asami, T. et al. Characterization of brassinazole, a triazole-type brassinosteroid biosynthesis inhibitor. Plant Physiol. 123, 93–100 (2000).
Choe, S. et al. The DWF4 gene of Arabidopsis encodes a cytochrome P450 that mediates multiple 22alpha-hydroxylation steps in brassinosteroid biosynthesis. Plant Cell 10, 231–243 (1998).
Clouse, S. & Sasse, J. Brassinosteroids: Essential regulators of plant growth and development. Annu. Rev. Plant Physiol. Plant Mol. Biol. 49, 427–451 (1998).
Noguchi, T. et al. Brassinosteroid-insensitive dwarf mutants of Arabidopsis accumulate brassinosteroids. Plant Physiol 121, 743–752 (1999).
Fujioka, S., Noguchib, T., Takatsutod, S. & Yoshida, S. Activity of brassinosteroids in the dwarf rice lamina inclination bioassay. Phytochemistry 49, 1841–1848 (1998).
Luo, W., Janzen, L., Pharis, R. P. & Back, T. G. Bioactivity of brassinolide methyl ethers. Phytochemistry 49, 637–642 (1998).
Trotochaud, A. E., Jeong, S. & Clark, S. E. CLAVATA3, a multimeric ligand for the CLAVATA1 receptor-kinase. Science 289, 613–617 (2000).
Torii, K. U. et al. The Arabidopsis ERECTA gene encodes a putative receptor protein kinase with extracellular leucine-rich repeats. Plant Cell 8, 735–746 (1996).
Jinn, T. L., Stone, J. M. & Walker, J. C. HAESA, an Arabidopsis leucine-rich repeat receptor kinase, controls floral organ abscission. Genes Dev. 14, 108–117 (2000).
Yamamuro, C. et al. Loss of function of a rice brassinosteroid insensitive1 homolog prevents internode elongation and bending of the lamina joint. Plant Cell 12, 1591–1605 (2000).
Li, J., Biswas, M. G., Chao, A., Russell, D. W. & Chory, J. Conservation of function between mammalian and plant steroid 5alpha- reductases. Proc. Natl Acad. Sci. USA 94, 3554–3559 (1997).
Seto, H. et al. A general approach to synthesis of labeled brassinosteroids: preparation of [25,26,27-2H7]brassinolide with 60% isotopic purity from the parent brassinolide. Tetrahedr. Lett. 39, 7525–7528 (1998).
Fankhauser, C. et al. PKS1, a substrate phosphorylated by phytochrome that modulates light signaling in Arabidopsis. Science 284, 1539–1541 (1999).
Acknowledgements
We thank M. Chen for comments and L. Barden for technical assistance on the manuscript; D. Vafeados for technical assistance; and D. Friedrichsen for providing the BRI1–GFP line. This work was supported by a grant from the USDA and the Howard Hughes Medical Institute to J.C., and by a Grant-in-Aid for Scientific Research from the Ministry of Education, Science, Sports, and Culture of Japan to S.F.. Z.W. is an NSF postdoctoral fellow and J.C. is an Associate Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, ZY., Seto, H., Fujioka, S. et al. BRI1 is a critical component of a plasma-membrane receptor for plant steroids. Nature 410, 380–383 (2001). https://doi.org/10.1038/35066597
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35066597
This article is cited by
-
An E2-E3 pair contributes to seed size control in grain crops
Nature Communications (2023)
-
Coordinated regulation of vegetative phase change by brassinosteroids and the age pathway in Arabidopsis
Nature Communications (2023)
-
Crosstalk between brassinosteroid signaling and variable nutrient environments
Science China Life Sciences (2023)
-
Physiological and Transcriptomic Analyses of the Effects of SlBRI1 Expression Levels on the Drought Tolerance of Tomato Seedlings
Journal of Plant Growth Regulation (2023)
-
Evolutionary analysis and functional characterization of BZR1 gene family in celery revealed their conserved roles in brassinosteroid signaling
BMC Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.