Abstract
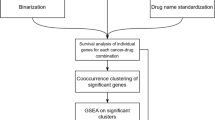
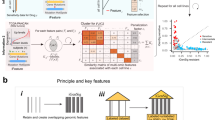
Genomic studies are producing large databases of molecular information on cancers and other cell and tissue types. Hence, we have the opportunity to link these accumulating data to the drug discovery processes. Our previous efforts at ‘information–intensive’ molecular pharmacology have focused on the relationship between patterns of gene expression and patterns of drug activity. In the present study, we take the process a step further—relating gene expression patterns, not just to the drugs as entities, but to ∼27 000 substructures and other chemical features within the drugs. This coupling of genomic information with structure-based data mining can be used to identify classes of compounds for which detailed experimental structure–activity studies may be fruitful. Using a systematic substructure analysis coupled with statistical correlations of compound activity with differential gene expression, we have identified two subclasses of quinones whose patterns of activity in the National Cancer Institute's 60-cell line screening panel (NCI-60) correlate strongly with the expression patterns of particular genes: (i) The growth inhibitory patterns of an electron-withdrawing subclass of benzodithiophenedione-containing compounds over the NCI-60 are highly correlated with the expression patterns of Rab7 and other melanoma-specific genes; (ii) the inhibitory patterns of indolonaphthoquinone-containing compounds are highly correlated with the expression patterns of the hematopoietic lineage-specific gene HS1 and other leukemia genes. As illustrated by these proof-of-principle examples, we introduce here a set of conceptual tools and fluent computational methods for projecting directly from gene expression patterns to drug substructures and vice versa. The analysis is presented in terms of the NCI-60 cell lines and microarray-based gene expression patterns, but the concept and methods are broadly applicable to other large-scale pharmacogenomic database sets as well. The approach (SAT for Structure-Activity-Target) provides a systematic way to mine databases for the design of further structure–activity studies, particularly to aid in target and lead identification.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ross DT, Scherf U, Eisen MB, Perou CM, Rees C, Spellman P et al . Systematic variation in gene expression patterns in human cancer cell lines Nat Genet 2000 24: 227–235
Scherf U, Ross DT, Waltham M, Smith LH, Lee JK, Tanabe L et al . A gene expression database for the molecular pharmacology of cancer Nat Genet 2000 24: 236–244
Boyd MR, Paull KD . Some practical consideration and applications of the National Cancer Institute in vitro anti-cancer drug discovery screen Drug Dev Des 1995 34: 91–109
Monks AP, Scudiero DA, Johnson GS, Paull KD, Sausville EA . The NCI anti-cancer drug screen: a smart screen to identify effectors of novel targets Anti-Cancer Drug Des 1997 12: 533–541
Paull KD, Shoemaker RH, Hodes L, Monks A, Scudiero DA, Rubinstein L et al . Display and analysis of patterns of differential activity of drugs against human tumor cell lines: development of mean graph and COMPARE algorithm J Natl Cancer Inst 1989 81: 1088–1092
Weinstein JN, Myers TG, O'Connor PM, Friend SH, Fornace AJ, Kohn KW et al . An information intensive approach to the molecular pharmacology of cancer Science 1997 275: 343–349
Weinstein JN, Kohn KW, Grever MR, Viswanadhan VN, Rubinstein LV, Monks AP et al . Neural computing in cancer drug development: predicting mechanism of action Science 1992 258: 447–451
Paull KD, Hamel E, Malspeis L . Prediction of biochemical mechanism of action from the in vitro antitumor screen of the National Cancer Institute In: Foye WE (ed) Cancer Chemotherapeutic Agents American Chemical Soc Books 1993 pp 1574–1581
Weinstein JN, Myers TG, Buolamwini JK, Raghavan K, van Osdol W, Licht J et al . Predictive statistics and artificial intelligence in the US National Cancer Institutes drug discovery program for cancer and AIDS Stem Cells 1994 12: 13–22
Shi LM, Myers TG, Fan Y, O'Connor PM, Paull KD, Friend SH et al . Mining the National Cancer Institute anticancer drug discovery database: cluster analysis of ellipticine analogs with p53-inverse and central nervous system-selective patterns of activity Mol Pharmacol 1998 53: 241–251
Shi LM, Fan Y, Myers TG, O'Connor PM, Paull KD, Friend SH et al . Mining the NCI anticancer drug discovery databases: genetic function approximation for the QSAR study of anticancer ellipticine analogues J Chem Inf Comput Sci 1998 38: 189–199
Wu L et al . Multidrug-resistant phenotype of disease-oriented panels of human tumor cell lines used for anticancer drug screening Cancer Res 1992 52: 3029–3034
Lee J-S et al . Rhodamine efflux patterns predict P-glycoprotein substrates in the National Cancer Institute drug screen Mol Pharmacol 1994 46: 627–638
Alvarez M et al . Generation of a drug resistance profile by quantitation of MDR-1/P-glycoprotein expression in the cell lines of the NCI anticancer drug screen J Clin Invest 1995 95: 2205–2214
Bates SE et al . Molecular targets in the National Cancer Institute drug screen J Cancer Res Clin Oncol 1995 121: 495–500
Izquierdo MA et al . Overlapping phenotypes of multidrug resistance among panels of human cancer-cell lines Int J Cancer 1996 65: 230–237
Koo H-M et al . Enhanced sensitivity to 1-beta-D-arabinofuranosylcytosine and topoisomerase II inhibitors in tumor cell lines harboring activated ras oncogenes J Natl Cancer Inst 1996 56: 5211–5216
O'Connor PM et al . Characterization of the p53-tumor suppressor pathway in cells of the National Cancer Institute anticancer drug screen and correlations with the growth-inhibitory potency of 123 anticancer agents Cancer Res 1997 57: 4285–4300
Freije JM et al . Identification of compounds with preferential inhibitory activity against low-Nm23-expressing human breast carcinoma and melanoma cell lines Nat Med 1997 3: 395–401
Wosikowski K et al . Identification of epidermal growth factor receptor and c-erbB2 pathway inhibitors by correlation with gene expression patterns J Natl Cancer Inst 1997 89: 1505–1513
Staunton JE, Slonim DK, Coller HA, Tamayo P, Angelo MJ, Park J et al . Chemosensitivity prediction by transcriptional profiling Proc Natl Acad Sci USA 2001 98: 10787–10792
Roberts G, Myatt GJ, Johnson WP, Cross KP, Blower PE . LeadScope: software for exploring large sets of screening data J Chem Inf Comput Sci 2000 40: 1302–1314
Fan Y, Weinstein JN, Kohn KW, Shi LM, Pommier Y . Molecular modeling studies of the DNA-topoisomerase I ternary cleavable complex with camptothecin J Med Chem 1998 41: 2216–2226
Cho SJ, Shen CF, Hermsmeier MA . Binary formal inference-based recursive modeling using multiple atom and physicochemical property class pair and torsion descriptors as decision criteria J Chem Inf Comput Sci 2000 40: 668–680
Klopman G, Shi LM, Ramu A . Quantitative structure-activity relationship of multi-drug resistance reversal agents Mol Pharmacol 1997 52: 323–334
Klopman G, Tu M . Diversity analysis of 14 156 molecules tested by the National Cancer Institute for anti-HIV activity using the quantitative structure-activity relational expert system MCASE J Med Chem 1999 42: 992–998
Weinstein JN . Fishing Expeditions Science 1998 282: 627–628
Weinstein JN . Pharmacogenomics: teaching old drugs new tricks N Eng J Med 2000 343: 1408–1409
Weinstein JN, Buolamwini JK . Molecular targets in cancer drug discovery: cell-based profiling Curr Pharm Des 2000 6: 473–483
Weinstein JN . Searching for pharmacogenomic markers: the synergy between omic and hypothesis-driven research Disease Markers 2001 17: 77–88
Bucci C, Thomsen P, Nicoziani P, McCarthy J, van Deurs B . Rab7: a key to lysosome biogenesis Mol Biol Cell 2000 11: 467–480
Meresse S, Steele-Mortimer O, Finlay BB, Gorvel JP . The rab7 GTPase controls the maturation of Salmonella typhimurium-containing vacuoles in HeLa cells EMBO J 1999 18: 4394–4403
Press B, Feng Y, Hoflack B, Wandinger-Ness A . Mutant. Rab7 causes the accumulation of cathepsin D and cation-independent mannose 6-phosphate receptor in an early endocytic compartment J Cell Biol 1998 140: 1075–1089
Hong SB, Li CM, Rhee HJ, Park JH, He X, Levy B et al . Molecular cloning and characterization of a human cDNA and gene encoding a novel acid ceramidase-like protein Genomics 1999 62: 232–241
Nagase T, Miyajima N, Tanaka A, Sazuka T, Seki N, Sato S et al . Prediction of the coding sequences of unidentified human genes III. The coding sequences of 40 new genes (KIAA0081-KIAA0120) deduced by analysis of cDNA clones from human cell line KG-1 DNA Res 1995 2: 37–43
Holmbeck K, Bianco P, Caterina J, Yamada S, Kromer M, Kuznetsov SA et al . MT1-MMP-deficient mice develop dwarfism, osteopenia, arthritis and connective tissue disease due to inadequate collagen turnover Cell 1999 99: 81–92
Apte SS, Fukai N, Beier DR, Olsen BR . The matrix metalloproteinase-14 (MMP-14) gene is structurally distinct from other MMP genes and is co-expressed with the TIMP-2 gene during mouse embryogenesis J Biol Chem 1997 272: 25511–25517
Shinmura K, Yamaguchi S, Saitoh T, Takeuchi-Sasaki M, Kim SR, Nohmi T et al . Adenine excisional repair function of MYH protein on the adenine:8-hydroxyguanine base pair in double-stranded DNA Nucleic Acids Res 2000 28: 4912–4918
Ohtsubo T, Nishioka K, Imaiso Y, Iwai S, Shimokawa H, Oda H et al . Identification of human MutY homolog (hMYH) as a repair enzyme for 2-hydroxyadenine in DNA and detection of multiple forms of hMYH located in nuclei and mitochondria Nucl Acids Res 2000 28: 1355–1364
Wang J, Brown EJ . Immune complex-induced integrin activation and L-plastin phosphorylation require protein kinase A J Biol Chem 1999 274: 24349–24356
Jones SL, Wang J, Turck CW, Brown EJ . A role for the actin-bundling protein L-plastin in the regulation of leukocyte integrin function Proc Natl Acad Sci 1998 95: 9331–9336
Ingley E, Sarna MK, Beaumont JG, Tilbrook PA, Tsai S, Takemoto Y et al . HS1 interacts with Lyn and is critical for erythropoietin-induced differentiation of erythroid cells J Biol Chem 2000 275: 7887–7893
Brunati AM, Donella-Deana A, James P, Quadroni M, Contri A, Marin O et al . Molecular features underlying the sequential phosphorylation of HS1 protein and its association with c-Fgr protein-tyrosine kinase J Biol Chem 1999 274: 7557–7564
Nestel FP, Colwill K, Harper S, Pawson T, Anderson SK . RS cyclophilins: identification of an NK-TR1-related cyclophilin Gene 1996 180: 151–155
Chao YH, Kuo SC, Ku K, Chiu I, Wu CH, Mauger A et al . Synthesis and cytotoxicity of Methyl-4,8-dihydrobenzo[1,2-b:5,4-b′]dithiophene-4,8-dione derivatives Bioorg Med Chem 1999 7: 1025–1031
Chao YH, Kuo SC, Wu CH, Lee CY, Mauger A, Sun IC et al . Synthesis and cytotoxicity of 2-acetyl-4,8-dihydrobenzodithiophene-4,8-dione derivatives J Med Chem 1998 41: 4658–4661
Kundel MW, Kirkpatrick DL, Johnson JI, Powis G . Cell line–directed screening assay for inhibitors of thioredoxin reductase signaling as potential anti-cancer drugs Anti Canc Drug Des 1997 12: 659–670
Rogge M, Fischer G, Pindur U, Schollmeyer D . α-Anellated carbazoles with anti-tumor activity: synthesis and cytotoxicity Monatsh Chem 1996 127: 97–102
Monks TJ, Hanzlik RP, Cohen GM, Ross D, Graham DG . Quinone chemistry and toxicity Toxicol Appl Pharmacol 1992 112: 2–16
O'Brien PJ . Molecular mechanisms of quinone cytotoxicity Chem Biol Interact 1991 80: 1–41
Bolton JL, Trush MA, Penning TM, Dryhurst G, Monks TJ . Role of quinones in toxicology Chem Res Toxicol 2000 13: 135–160
Phillips RM, Naylor MA, Jaffar M, Doughty SW, Everett SA, Breen AG et al . Bioreductive activation of a series of indolequinones by human DT-diaphorase: structure–activity relationships J Med Chem 1999 42: 4071–4080
Xing C, Wu P, Skibo EB, Dorr RT . Design of cancer-specific antitumor agents based on aziridinylcyclopent[b]indoloquinones J Med Chem 2000 43: 457–466
Beall HD, Hudnott AR, Winski S, Siegel D, Swann E, Ross D et al . Indolequinone antitumor agents: relationship between quinone structure and rate of metabolism by recombinant human NQO1 Bioorg Med Chem Lett 1998 8: 545–548
Fitzsimmons SA, Workman P, Grever M, Paull K, Camalier R, Lewis AD . Reductase enzyme expression across the National Cancer Institute tumor cell line panel: correlation with sensitivity to mitomycin C and EO9 J Natl Canc Inst 1996 88: 259–269
Sengupta SK . Inhibitors of DNA-transcribing enyzmes In Foye WE (ed) Cancer Chemotherapeutic Agents American Chemical Society: Washington, DC 1993 pp 205–260
Mitchell J, Marrian DH . Radiosensitization of cells by aderivative of2-methyl-1, 4-naphthoquinone In Morton RA (ed) Biochemistry of Quinones Academic Press: New York 1965 pp 503–541
Nesta P . Radiation chemistry of quinonoid compounds In Patai S, Rappoport S (eds) The Chemistry of the Quinonoid Compounds John Wiley & Sons: New York 1988 pp 879–898
Schaffer J . The Analysis of Incomplete Multivariate Data New York: Chapman and Hall 1996
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Blower, P., Yang, C., Fligner, M. et al. Pharmacogenomic analysis: correlating molecular substructure classes with microarray gene expression data. Pharmacogenomics J 2, 259–271 (2002). https://doi.org/10.1038/sj.tpj.6500116
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.tpj.6500116
Keywords
This article is cited by
-
Structure based classification for bile salt export pump (BSEP) inhibitors using comparative structural modeling of human BSEP
Journal of Computer-Aided Molecular Design (2017)
-
An integrative analysis of small molecule transcriptional responses in the human malaria parasite Plasmodium falciparum
BMC Genomics (2015)
-
Determination of minimal transcriptional signatures of compounds for target prediction
EURASIP Journal on Bioinformatics and Systems Biology (2012)
-
Systematically characterizing and prioritizing chemosensitivity related gene based on Gene Ontology and protein interaction network
BMC Medical Genomics (2012)
-
Comprehensive data-driven analysis of the impact of chemoinformatic structure on the genome-wide biological response profiles of cancer cells to 1159 drugs
BMC Bioinformatics (2012)