Abstract
Background:
MicroRNA (miRNA, miR)-18a is a member of the miR-17–92 cluster, an important locus that is markedly overexpressed in several cancers and associated with cancer development and progression. However, the mechanism of action of the miR-17–92 cluster and its individual miRNAs are largely unknown.
Methods and Results:
In this study, we investigated the expression of the miR-17–92 cluster by in situ hybridisation (ISH) assay and copy-number analysis in gastric tissue microarray (TMA) specimens. We determined that miR-18a was present at higher levels than the other five miRNAs in the cluster. In addition, we identified Protein Inhibitor of Activated Signal Transducer and Activator of Transcription 3 (PIAS3) as a direct target of miR-18a in gastric cancer. miR-18a level was positively correlated with levels of Survivin, Bcl-xL, and c-Myc, which are downstream transcriptional targets of Signal Transducer and Activator of Transcription 3 (STAT3). STAT3-induced transcription can be negatively regulated by PIAS3; consistent with this, PIAS3 level was negatively correlated with levels of Survivin, Bcl-xL, and c-Myc.
Conclusion:
Our findings indicate that miR-18a acts as an oncogene and plays a role in gastric adenocarcinogenesis, at least in part by negatively regulating PIAS3 and thereby modulating expression of STAT3 target genes.
Similar content being viewed by others
Main
Gastric cancer is the fourth most common cancer and the second leading cause of cancer death in the world. More than 980 000 new gastric cancer cases are diagnosed annually, and the disease causes ∼730 000 deaths per year. The highest incidence rates of gastric cancer are in Eastern Asia, Eastern Europe, and South America (Jemal et al, 2011). Helicobacter pylori (H. pylori) is an important environmental cause of gastric cancer and has a high infection rate in the general population. However, only a small number of H. pylori-infected individuals suffer from gastric cancer, suggesting that genetic polymorphism plays an important role in gastric cancer (Wang et al, 2010).
MicroRNAs are endogenous 18–24-nucleotide (nt) single-stranded RNA molecules that act as posttranscriptional regulators of gene expression (Poliseno et al, 2010). They bind to sequences in the 3′ untranslated regions (3′UTRs) of target genes; base pairing between nucleotides 2–7 of the miRNA (the miRNA seed sequence) and the corresponding sequence in the target 3′UTR (the seed match) is necessary for target recognition (Grimson et al, 2007). Binding of an mRNA to a 3′UTR decreases protein translation, the stability of nascent mRNA strands, or both, thereby decreasing production of the target protein (Filipowicz et al, 2008). miRNAs sharing the same seed sequence are grouped into families and are theorised to target overlapping sets of genes (Landgraf et al, 2007). MicroRNAs play diverse roles in numerous cellular processes; in particular, their abundance is altered during tumourigenesis, and they can act as tumour suppressors or oncogenes (Ventura and Jacks, 2009).
The miR-17–92 cluster encodes six miRNAs (miR-17, miR-18a, miR-19a, miR-20a, miR-19b, and miR-92a) (Mendell, 2008). The promoter region of miR-17–92 bears a functional binding site for STAT3, which transcriptionally activates the miR-17–92 cluster (Brock et al, 2009). A recent study identified the members of the miR-17–92 cluster that are most highly expressed in gastric cancer tissues (Guo et al, 2009). In addition, Pichiorri et al (2008) showed that miRNAs derived from the miR-17–92 cluster modulate STAT3 phosphorylation in multiple myeloma cells.
Recent studies have also revealed that altered STAT3 activity can contribute to oncogenesis (Yu et al, 1995). Regulators of STAT3 include PIAS3, which was originally identified as a specific inhibitor of STAT3 (Junicho et al, 2000). Under physiological conditions in normal cells, the activation of STAT proteins is rapid and transient because of negative regulation by proteins such as SOCS and PIAS (Alexander, 2002; Kubo et al, 2003; Shuai and Liu, 2005). Activated STAT3 protects tumour cells from apoptosis and promotes the expression of cell proliferation-associated proteins, such as Bcl-xL, Mcl-1, Bcl-2, Fas, cyclin D1, Survivin, and c-Myc (Bromberg et al, 1999; Catlett-Falcone et al, 1999; Epling-Burnette et al, 2001; Ivanov et al, 2001; Bromberg, 2002; Yu and Jove, 2004).
In this study, we examined the expression level of miR-18a in gastric adenocarcinoma (GAC) tissues and adjacent non-tumour tissues. In addition, we investigated the underlying mechanisms of miR-18a function in GAC. Our results may lead to identification of therapeutic candidates for GAC.
Materials and methods
Human tissue specimens and tissue microarrays
We used tissue samples obtained from surgically resected GAC specimens generated at Tokyo Medical University Hospital, from 2003 to 2009. The institutional review board approved this study, and all patients provided written informed consent. Tissue specimens were fixed in 10% formalin solution and embedded in paraffin; clinicopathological characteristics are listed in Table 1. Tissue microarrays (TMAs) allow the relocation of multiple tissue samples from conventional histologic paraffin blocks so that tissues from multiple patients can be seen on the same slide. We generated GAC TMAs, each consisting of 12 specimens of 6-mm diameter on one slide.
In situ hybridisation
In situ hybridisation (ISH) was performed on TMAs containing 3-μm consecutive sections of formalin-fixed, paraffin-embedded (FFPE) samples. Probes were 5′ digoxigenin (DIG)-labelled ISH LNA (locked nucleic acid) (miRCURY-LNA detection probe; Exiqon). Slides were prepared using the Ventana HX System BenchMark (Ventana Medical Systems Inc., Tucson, AZ, USA). We used the following 5′ DIG-labelled LNA probes for ISH:
hsa-miR-17, 5′-CTACCTGCACTGTAAGCACTTTG-3′;
hsa-miR-18a, 5′-CTATCTGCACTAGATGCACCTTA-3′; hsa-miR-19a, 5′-TCAGTTTTGCATAGATTTGCACA-3′; hsa-miR-20a, 5′-CTACCTGCACTATAAGCACTTTA-3′; hsa-miR-19b, 5′-TCAGTTTTGCATGGATTTGCACA-3′; hsa-miR-92a, 5′-ACAGGCCGGGACAAGTGCAATA-3′; and non-target negative control sequence, 5′-GTGTAACACGTCTATACGCCCA-3′.
RNA isolation and quantitative real-time reverse transcription polymerase chain reaction (qRT–PCR) analysis
Total RNA was extracted from FFPE human gastric tissues (10 non-tumour samples and 15 GAC samples) using the High Pure FFPE RNA miRNA Isolation Kit (Roche Diagnostic, Basel, Switzerland); clinicopathological characteristics of these samples are listed in Table 2. The RNA samples were suspended in 20 μl nuclease-free water, and miRNAs were quantified using TaqMan MicroRNA Assays (Applied Biosystems; Life Technologies, Carlsbad, CA, USA), as described previously (Tanaka et al, 2009). The miRNA levels were normalised against levels of RNU6B.
Cell culture and transfection
The human GAC cell lines MKN28 and MKN1 were cultured in RPMI-1640 medium (Gibco BRL, Rockville, MD, USA) supplemented with 10% fetal calf serum (FCS) in 5% CO2 at 37°C. MiRNA was transfected at the concentration of 100 nM using the HiPerFect Transfection Reagent (QIAGEN, Hilden, Germany). The hsa-miR-18a oligonucleotide was a synthetic double-stranded 23-nt RNA, 5′-UAAGGUGCAUCUAGUGCAGAUAG-3′, purchased from BONAC Corporation (Fukuoka, Japan). The micRCURY LNA Power Inhibitor hsa-miR-18a oligonucleotide sequence, 5′-ATCTGCACTAGATGCACCTT-3′, was purchased from Exiqon.
qRT–PCR analysis of PIAS3, Bcl-xL, c-Myc, and Survivin
Total RNA was isolated from MKN28 and MKN1 cells with the use of an ISOGEN reagent (Nippon Gene, Osaka, Japan). To measure the mRNA levels of PIAS3 and STAT3 target genes (Bcl-xL, c-Myc, Survivin), reverse-transcription reactions were performed using M-MLV reverse transcriptase (Invitrogen, Carlsbad, CA, USA). Quantitative PCR (FastStart Universal SYBR Green Master; Roche) reactions were run on an MX3005P thermocycler (Stratagene, La Jolla, CA, USA) and analysed using the MxPro QPCR software, version 4.01 (Stratagene, Agilent Technologies, Santa Clara, CA, USA). The level of gene expression relative to Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was determined. The primer sequence were as follows: The PIAS3 primers: 5′-TGTCACCATGAAACCATTGC-3′ (forward) and 5′-AGGTAAAGTGCGCTTCCTCA-3′ (reverse). Bcl-xL primers: 5′-GGGCATTCAGTGACCTGACA-3′ (forward) and 5′-GCATTGTTCCCATAGAGTTC-3′ (reverse). c-Myc primers: 5′-TCCTCGGATTCTCTGCTCTC-3′ (forward) and 5′-CTCTGACCTTTTGCCAGGAG-3′ (reverse). Survivin primers: 5′-CTGGCAGCCCTTTCTCAA-3′ (forward) and 5′-CAGCCTTCCAGCTCCTTG-3′ (reverse). GAPDH primers: 5′-ATGGGGAAGGTGAAGGTCG-3′ (forward) and 5′-GGGTCATTGATGGCAACAATATC-3′ (reverse).
Plasmid construction
A 924-base pair (bp) fragment of PIAS3 3′UTR (nt 2079–3002, WT) was amplified by PCR and cloned into the BamHI and EcoRI sites of the pcDNA-GL3 reporter vector (Promega, Madison, WI, USA), using the PIAS3 3′UTR cloning primers 5′-AAAGAATTCGTTCCCTGGATTATGGAAAC-3′ (forward) and 5′-AAAGGATCCGAACATTCACAACCTTTATT-3′ (reverse). A deletion mutant of the miR-18a-binding site of the PIAS3 3′UTR (Mut) was generated using primers 5′-AAAGGGCCCTCCAGGGAGGTGGAAGGA-3′ (forward) and 5′-AAAGGGCCCGAATGAGCCAGGCCTAAC-3′ (reverse).
Analysis of Argonaute 2-associated mRNA
MKN28 and MKN1 cells were transfected with 100 nM of miR-18a oligonucleotide. Co-immunoprecipitation experiments were performed using an anti-Argonaute 2 (Ago2) antibody (Wako Pure Chemical Industries, Tokyo, Japan), as described previously (Tanaka et al, 2009). The mRNAs that were associated with Ago2 were analysed by qRT–PCR. The data were normalised against the PIAS3 mRNA expression levels in the input samples.
Immunohistochemistry
Immunohistochemistry (IHC) was performed on TMAs containing 4-μm consecutive sections of FFPE gastric tissue samples. After deparaffinisation and rehydration, endogenous peroxidases were blocked by incubation in 0.3% hydrogen peroxide solution for 20 min. Antigen-retrieval treatment was performed by heating in a microwave oven for 20 min in 0.01 M sodium citrate buffer (pH 6.0). For detection of PIAS3, Survivin, or c-Myc, cells were incubated overnight at room temperature (RT) with PIAS3 antibody (rabbit anti-human PIAS3 polyclonal antibody (N-term) at 1 : 100 dilution; Abgent, San Diego, CA, USA), Survivin antibody (rabbit anti-human Survivin polyclonal antibody at 1 : 1000 dilution; Novus Biologicals, Littleton, CO, USA), or c-Myc antibody (mouse monoclonal [9E10] at 1 : 300 dilution; Abcam, Cambridge, UK), respectively. Samples were then washed three times in PBS and incubated with Dako EnVision Labelled Polymer Peroxidase secondary antibody for 30 min at RT. For detection of Bcl-xL, STAT3, or pSTAT3, samples were incubated with Bcl-xL antibody (rabbit anti-human Bcl-xL polyclonal antibody at 1 : 100 dilution, Cell Signaling Technology, Danvers, MA, USA), STAT3 antibody (rabbit anti-human STAT3 polyclonal antibody at 1 : 400 dilution, Cell Signaling Technology), or pSTAT3 antibody (rabbit anti-human pSTAT3 polyclonal antibody at 1 : 100 dilution, Abgent), respectively, for 3 h at RT. Samples were then washed three times in PBS and incubated with Dako anti-rabbit secondary antibody for 15 min at RT. For all immunohistochemistry experiments, 3, 3′-Diaminobenzidine tetrachloride (DAB) was used for colour development, and sections were counterstained with haematoxylin. Brown colour in the nucleus or cytoplasm was scored as positive staining.
Reporter gene assay
MKN28 and MKN1 cells were seeded in 24-well plate at 8 × 104 cells per well and incubated overnight. miR-18a was transfected into MKN28 and MKN1 cells using the HiPerFect transfection reagent (QIAGEN). One day (24 h) after transfection, the cells were transfected with pRL-TK Renilla luciferase plasmid in combination with PIAS3 3′UTR expression plasmid or STAT3 reporter plasmid (Cignal Reporter Assay Kit, QIAGEN), using the Fugene HD reagent (Roche Applied Sciences, Tokyo, Japan). Cells were harvested 24 or 48 h after transfection and assayed for luciferase activity using the Dual-Luciferase Reporter Assay System (Promega); luciferase activity was normalised against the activity of Renilla luciferase. Luminescence was quantified with a luminometer (Lumat CB 9507, Belthold, Wildbad, Germany).
Image analysis
IHC- or ISH-stained slides were examined by light microscopy using an Olympus BX50 (Olympus, Tokyo, Japan). Positive IHC results appeared light brown, brown, or dark brown, whereas positive ISH results caused the cytoplasm to appear blue. Slides were interpreted semi-quantitatively by assessing the intensity and extent of staining on the entirety of each tissue section present on the slides, as described previously (Acs et al, 2003).
Statistical analysis
The comparison of GAC with adjacent non-tumour tissues for each member of the miR-17–92 cluster, STAT3, and pSTAT3 were performed using Welch’s t-test. The correlation between the miR-18a ISH staining score and the PIAS3, Survivin, Bcl-xL, and c-Myc staining scores were estimated using Spearman’s correlation coefficient by rank test. P-value<0.05 was considered to be statistically significant (*P<0.05, **P<0.01, ***P<0.001).
Results
Upregulation of miR-18a in human GAC tissues in comparison with adjacent non-tumour tissues
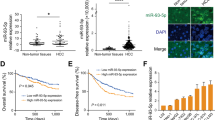
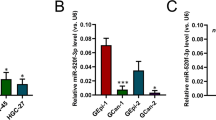
The miR-17–92 cluster (C13orf25) produces a single polycistronic primary transcript that yields six mature miRNAs: miR-17, miR-18a, miR-19a, miR-20a, miR-19b, and miR-92a (Figure 1A). To determine the expression patterns of these six miRNAs, we quantified their expression levels by performing ISH assays on gastric TMA specimens. Our TMAs contained 73 GAC specimens and 10 adjacent non-tumour tissue specimens (Table 1). MicroRNA expression was defined by ISH staining scores, as described previously (Acs et al, 2003). In this study, we found that expression levels of all miRNAs were increased in GAC tissues in comparison with adjacent non-tumour tissues: miR-17 (P<0.001), miR-18a (P<0.001), miR-19a (P=0.219), miR-20a (P<0.001), miR-19b (P<0.001), and miR-92a (P<0.001). MicroRNA-19a expression was the only member of the miR-17–92 cluster that was not significantly differentially expressed between GAC tissues and adjacent non-tumour tissues (Figure 1B). We suspect that low stability, rather than differences in maturation or processing, caused the low abundance of miR-19a. The expression level of miR-92a was the highest among the members of the miR-17–92 cluster, but the expression ratio of miR-18a in GAC vs adjacent non-tumour tissue was the highest (>20-fold) (Figure 1B and C). We also quantified the miR-18a levels in gastric FFPE samples by qRT–PCR analysis. In qRT–PCR analysis, the efficiency of primer annealing influences the amplification of the product, making it difficult to compare expression levels of amplified products that use different primers. To allow us to compare the six miRNAs produced from a single polycistronic transcript, we calculated the copy number for each of these six miRNAs, and normalised the values against a standard curve generated using individual synthetic miRNAs. We examined 10 samples of non-tumour tissues and 15 samples of GAC tissues; clinicopathological characteristics are listed in Table 2. There were significant increases in the expression levels of all six miRNAs in cancer tissue in compared with adjacent non-tumour tissues. These results were consistent with the ISH results, in that the copy number ratio of miR-18a in GAC vs adjacent non-tumour tissue was the highest (P<0.01) (Figure 1D and data not shown). These results confirm that the expression of miR-18a, a member of the miR-17–92 cluster, is significantly elevated in human gastric cancer tissues.
Expression ratios for members of the miR-17–92 cluster in human GAC specimens on TMAs. (A) Genomic structure of the miR-17–92 polycistronic cluster. (B) Representative ISH micrographs ( × 400) for members of the miR-17–92 cluster. (C) Expression ratios for members of the miR-17–92 cluster (gastric adenocarcinoma vs adjacent non-tumour tissue). (D) Expression levels of miR-18a, as determined by qRT–PCR, in GAC vs adjacent non-tumour tissue (**P<0.01, Welch’s t-test).
PIAS3 is a direct target of miR-18a
Recent studies have also indicated that miR-18a is significantly upregulated in gastric cancer compared with adjacent non-tumour tissue (Guo et al, 2009; Yao et al, 2009). Overexpression of miR-18a does not directly affect the proliferation of tumour cells (Hayashita et al, 2005; Dews et al, 2006; Matsubara et al, 2007). On this basis, we attempted to dissect the potential mechanism of miR-18a in the development of GAC. The importance of miRNAs in IL-6 signal transduction has been highlighted by a recent study showing that miRNAs derived from the miR-17–92 cluster modulate STAT3 phosphorylation in multiple myeloma cells (Pichiorri et al, 2008). Since miRNAs have been implicated in the repression of gene expression, we speculated that miR-18a might target a negative regulator of the STAT3 signalling pathway. Utilising computational screening (TargetScan, Whitehead institute for Biomedical Research, www.targetscan.org), we determined that the 3′UTR of PIAS3 contains a potential binding site for miR-18a (Figure 2A). We evaluated the transfection efficiency of synthetic miR-18a in the gastric cancer cell lines MKN28 and MKN1. As shown in Figure 2B, qRT–PCR analysis revealed that miRNA-18a levels in miR-18a-transfected cells were significantly higher than those in untransfected or control-transfected cells. Using these experimental conditions, we conducted further investigations. To determine whether miR-18a could directly target PIAS3 in a GAC cell line, we performed immunoprecipitation assays using an anti-Ago2 antibody. We analysed the expression levels of PIAS3 mRNA in Ago2 immunoprecipitates from cell lysates of MKN28 and MKN1 cells transfected with miR-18a. These two cell lines ordinarily exhibit low levels of miR-18a and high levels of PIAS3 mRNA (data not shown). We observed increases in PIAS3 mRNA associated with Ago2 in miR-18a-transfected cells (Figure 2C) (MKN28 P<0.01, MKN1 P<0.01). To investigate whether miR-18a influences PIAS3 expression, we measured mRNA levels of PIAS3 in miR-18a-transfected MKN28 and MKN1 cells. As shown in Figure 2D, miR-18a transfection significantly reduced the level of PIAS3 mRNA relative to the levels in untransfected and control-transfected cells, in both MKN28 (vs untransfected: P<0.01; vs control-transfected: P<0.05) and MKN1 cells (P<0.05 for both cases).
Protein inhibitor of activated signal transducer and activator of transcription 3 is a direct target of miR-18a. (A) TargetScan prediction software identified one seed match for miR-18a in the 3′UTR of PIAS3; the predicted base pairing of PIAS3 mRNA and miR-18a is shown. (B) miR-18a-transfected MKN28 and MKN1 cells revealed a significant upregulation of miR-18a compared with untransfected and control-transfected cells (P<0.01 and P<0.01). Data are mean values±s.d. of values from three independent experiments. (C) Using an anti-Ago2 antibody, RNA was co-immunoprecipitated from MKN28 and MKN1 cells transfected with synthetic miR-18a or a control oligonucleotide. RNA was isolated, reverse transcribed, and PCR amplified using primers specific for the PIAS3 mRNA. Data are mean values±s.d. of values from three independent experiments. (D) MicroRNA-18a-transfected MKN28 and MKN1 cells revealed a significant downregulation of PIAS3 mRNA levels compared with untransfected and control-transfected cells (P<0.05 and P<0.05). Data are mean values±s.d. of values from three independent experiments. (E) Reporter constructs were generated containing the seed-match fragment of the PIAS3 3′UTR with either a wild-type (WT) or mutated (Mut) miR-18a complementary site. Black rectangle indicates the miR-18a seed match; black rectangle with an X reflects the mutation sites. (F) Reporter gene studies in MKN28 and MKN1 cells revealed that co-transfection of miR-18a significantly decreased relative luciferase activity compared with untransfected and control-transfected cells. Neither the pcDNA-GL3 construct (GL3) nor the mutated construct (Mut) was affected by overexpression of miR-18a. Data are mean values±s.d. of values from three independent experiments. * and **, statistically significant (P<0.05 and P<0.01, respectively; post-hoc test). GL3, no insertion of 3′UTR seed sequence; WT, wild-type 3′UTR seed; Mut, mutant 3′UTR seed. (G) Anti-miR-18a-transfected MKN28 and MKN1 cells revealed a significant upregulation of PIAS3 mRNA levels compared with untransfected and control-transfected cells (P<0.05 and P<0.05, respectively). Data are means±s.d. of values from three independent experiments.
To determine whether the observed reduction of PIAS3 expression is directly driven by miR-18a, we performed reporter gene assays in MKN28 and MKN1 cells. The entire 3′UTR of PIAS3 was cloned into the pcDNA-GL3 control vector, creating a luciferase reporter gene containing the seed match for miR-18a. In addition, we generated a mutated reporter construct in which the miR-18a seed match sequence was deleted from the 3′UTR of PIAS3. Co-transfection of the PIAS3 3′UTR construct and miR-18a yielded a significantly reduced relative luciferase activity (MKN28, P<0.01; MKN1, P<0.05; Figure 2F). Neither the pcDNA-GL3 construct nor the mutated construct was affected by overexpression of miR-18a. The luciferase assays were performed in triplicate, and the data were observed three times after transfected 24 h. Moreover, anti–miR-18a transfection significantly increased the expression levels of PIAS3 mRNA compared to untransfected and control-transfected cells, in both MKN28 and MKN1 cells (P<0.01 for all comparisons) (Figure 2G). These data demonstrate a direct interaction between PIAS3 mRNA and miR-18a.
Downregulation of PIAS3 expression was inversely correlated with miR-18a expression in GAC TMA specimens
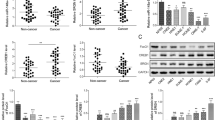
Next, we used the same TMA specimens to examine the expression levels of PIAS3 by immunohistochemical assay using anti-PIAS3 antibody. In GAC, PIAS3 expression was lower (P<0.01), whereas miR-18a expression was higher (P<0.01), than in adjacent non-tumour tissues (Figure 3B). We also observed a significant inverse correlation between the expression of miR-18a and PIAS3 in the same TMA specimens (r=−0.42, P<0.001) (Figure 3C). These data support the hypothesis that PIAS3 is a direct target of miR-18a.
Protein inhibitor of activated signal transducer and activator of transcription 3 was downregulated in GAC TMA specimens and inversely correlated with miR-18a expression levels. (A) Representative visualisations of miR-18a and PIAS3. A tumour specimen with high expression of miR-18a and low expression of PIAS3 (right), and adjacent non-tumour tissue with high expression of PIAS3 and low expression of miR-18a (left), are shown. (B) miR-18a ISH staining score and PIAS3 staining score in GAC and adjacent non-tumour tissues (**P<0.01, Welch’s t-test). (C) The level of miR-18a exhibited a statistically significant inverse correlation with the level of PIAS3 (r=−0.42, P<0.001, Spearman’s correlation coefficient rank test).
Overexpression of miR-18a and downregulation of PIAS3 enhance STAT3-mediated gene expression
The data presented so far demonstrate that miR-18a directly inhibits PIAS3, a known repressor of STAT3 activity. To demonstrate that miR-18a affects STAT3 signalling, we performed STAT3 reporter assays in miR-18a-transfected MKN28 and MKN1 cells. As shown in Figure 4A and B, the transcriptional activity of STAT3 was more pronounced in miR-18a-transfected MKN28 and MKN1 cells than in cells transfected with a control oligonucleotide (P<0.01). Next, we evaluated whether overexpression of miR-18a could increase STAT3-mediated gene expression. MKN28 and MKN1 cells were transfected with miR-18a for 48 h, and then the cells were harvested and processed for qRT–PCR. miR-18a increased Bcl-xL and c-Myc mRNA levels in MKN28 and MKN1 cells (P<0.01) (Figure 4C–F). However, the transcript levels of Survivin were not increased by transfection of miR-18a (data not shown). In addition, we performed IHC on the same gastric TMA specimens using antibodies against Survivin, STAT3, pSTAT3, Bcl-xL, and c-Myc. The expression levels of miR-18a correlated positively, and the expression levels of PIAS3 correlated negatively, with the expression levels of Survivin, Bcl-xL, and c-Myc (Figure 4G × 400 and 4H). Moreover, the expression levels of STAT3 and pSTAT3 were significantly increased in GAC compare to non-tumour tissues (Figure 4I × 400 and 4J). STAT3 has a broad range of biological functions, including cell activation, cell proliferation, and apoptosis. Therefore, activated STAT3 can protect tumour cells from apoptosis and promote cell proliferation by regulating genes encoding anti-apoptotic and proliferation-associated proteins, such as Bcl-xL, Mcl-1, Bcl-2, Fas, cyclin D1, Survivin, and c-Myc (Bromberg et al, 1999; Catlett-Falcone et al, 1999; Epling-Burnette et al, 2001; Ivanov et al, 2001; Bromberg, 2002; Yu and Jove, 2004). Thus, these findings demonstrate that expression of miR-18a enhances STAT3-mediated gene expression and promotes development of adenocarcinoma.
Overexpression of miR-18a downregulation of PIAS3 enhanced STAT3-mediated gene expression. (A, B) Reporter gene studies in MKN28 and MKN1 cells revealed that co-transfection of miR-18a significantly increased the relative STAT3 luciferase activity compared with untransfected and control-transfected cells (**P<0.01, post-hoc test). Exogenous overexpression of miR-18a enhanced the expression levels of Bcl-xL (C, D) and c-Myc (E, F). (G) Representative pictures of GAC specimen positive for miR-18a and Survivin, Bcl-xL, and c-Myc and negative for PIAS3. (H) A statistically significant positive association between miR-18a and Survivin, Bcl-xL, and c-Myc, and a statistically significant negative association between PIAS3 and these proteins. (I) Representative pictures of a GAC specimen positive for STAT3 and pSTAT3. (J) The expression levels of STAT3 and pSTAT3 in GAC and adjacent non-tumour tissues (**P<0.01, Welch’s t-test).
Discussion
Recently, a large-scale analysis of the miRNA profiles of solid tumours detected upregulation of the human miR-17–92 cluster in many cancers, including lung (Yanaihara et al, 2006) and gastrointestinal cancers (Valladares-Ayerbes et al, 2011). Moreover, miR-18a is significantly upregulated in gastric cancer compared with adjacent non-tumour tissue (Guo et al, 2009; Yao et al, 2009), but its roles and regulatory mechanisms in gastric cancer remain unknown. In this study, we used ISH to show that miR-18a was the most upregulated miRNA from the miR-17–92 cluster in GAC relative to adjacent non-tumour tissue (Figure 1C); this finding was confirmed by qRT–PCR assays (Figure 1D). It is therefore conceivable that miR-18a functions as an oncogenic miRNA.
MicroRNAs have primarily been associated with the repression of gene expression (Bartel, 2004). Using a computational approach, we identified a potential binding site for miR-18a in the 3′UTR of the PIAS3 gene, which encodes a protein that interferes with the DNA-binding activity of STAT3 (Chung et al, 1997). We observed increased levels of PIAS3 mRNA in association with Ago2 in miR-18a-transfected MKN28 and MKN1 cells (Figure 2C). Overexpression of miR-18a decreased the PIAS3 mRNA level, as determined by qRT–PCR (Figure 2D). Furthermore, using a luciferase reporter gene containing the 3′UTR of PIAS3, we confirmed direct repression of PIAS3 by miR-18.
Protein inhibitor of activated signal transducer and activator of transcription 3 is a specific inhibitor of STAT3 (Chung et al, 1997). STAT3 has a broad range of biological functions, including cell activation, cell proliferation, and apoptosis. Activated STAT3 protects tumour cells from apoptosis and promote cell proliferation by regulating genes encoding antiapoptotic and proliferation-associated proteins, such as Bcl-xL, Mcl-1, Bcl-2, Fas, cyclin D1, Survivin, and c-Myc (Bromberg et al, 1999; Catlett-Falcone et al, 1999; Epling-Burnette et al, 2001; Ivanov et al, 2001; Bromberg, 2002; Yu and Jove, 2004). Inhibition of apoptosis by Survivin is important in the pathogenesis of gastric cancer (Yu et al, 2002). Bcl-xL is overexpressed in gastric carcinomas at both the RNA and protein levels, suggesting that overexpression of Bcl-xL may also play a role in gastric carcinogenesis (Kondo et al, 1996). c-Myc is a strong inducer of proliferation and is believed to be critical for oncogenesis; aberrant c-Myc expression is widespread in tumour cells and important for multistep carcinogenesis (Yokozaki, 2000; Chen et al, 2001). We performed STAT3 reporter assays in miR-18a-transfected MKN28 and MKN1 cells; the results demonstrated that miR-18a affects STAT3 signalling. Furthermore, qRT–PCR revealed that levels of Bcl-xL and c-Myc were significantly increased in MKN28 and MKN1 cells transfected with miR-18a (Figure 4C–F). In addition, our IHC assays revealed that miR-18a level correlated positively, whereas PIAS3 mRNA level correlated negatively, with Survivin, Bcl-xL, and c-Myc expression in GAC TMA specimens (Figure 4G and H). These findings indicate that miR-18a induces anti-apoptotic and cell-proliferation gene in GAC, suggesting that inhibitors of miR-18a or activators of PIAS3 could potentially act as adenocarcinostatic drugs.
Interestingly, Brock et al (2011) demonstrated that miR-18a activates the IL-6/STAT3 signalling pathway by repressing the expression of PIAS3 in human hepatocytes. This study supported our findings and suggested that miR-18a plays the role of STAT3 signalling pathway not only in tumour but also in non-tumour.
We propose a model for mechanisms of gastric adenocarcinogenesis by miR-18a, shown as Figure 5. In this model, overexpression of miR-18a in MKN28 and MKN1 cells decreases PIAS3 expression, thereby increasing STAT3 activity and leading to enhanced activation of genes downstream of STAT3, such as the antiapoptotic gene Bcl-xL and the cell-proliferation gene c-Myc. These mechanisms may contribute to the development of GAC.
Schematic overview of miR-18a regulatory signalling. We propose the following model: miR-18a targets the STAT3 inhibitor PIAS3, which leads to enhanced activation of genes downstream of STAT3, such as the antiapoptotic genes Survivin and Bcl-xL and the cell-proliferation gene c-Myc, thereby contributing to tumourigenesis.
Change history
19 February 2013
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Acs G, Zhang PJ, McGrath CM, Acs P, McBroom J, Mohyeldin A, Liu S, Lu H, Verma A (2003) Hypoxia-inducible erythropoietin signaling in squamous dysplasia and squamous cell carcinoma of the uterine cervix and its potential role in cervical carcinogenesis and tumour progression. Am J Pathol 162: 1789–1806
Alexander WS (2002) Suppressors of cytokine signalling (SOCS) in the immune system. Nat Rev Immunol 2: 410–416
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116: 281–297
Brock M, Trenkmann M, Gay RE, Gay S, Speich R, Huber LC (2011) microRNA-18a enhances the interleukin-6-mediated production of the acute-phase proteins fibrinogen and haptoglobin in human hepatocytes. J Biol Chem 286: 40142–40150
Brock M, Trenkmann M, Gay RE, Michel BA, Gay S, Fischler M, Ulrich S, Speich R, Huber LC (2009) Interleukin-6 modulates the expression of the bone morphogenic protein receptor type II through a novel STAT3-microRNA cluster 17/92 pathway. Circ Res 104: 1184–1191
Bromberg J (2002) Stat proteins and oncogenesis. J Clin Invest 109: 1139–1142
Bromberg JF, Wrzeszczynska MH, Devgan G, Zhao Y, Pestell RG, Albanese C, Darnell JE (1999) Stat3 as an oncogene. Cell 98: 295–303
Catlett-Falcone R, Landowski TH, Oshiro MM, Turkson J, Levitzki A, Savino R, Ciliberto G, Moscinski L, Fernandez-Luna JL, Nunez G, Dalton WS, Jove R (1999) Constitutive activation of Stat3 signaling confers resistance to apoptosis in human U266 myeloma cells. Immunity 10: 105–115
Chen JP, Lin C, Xu CP, Zhang XY, Fu M, Deng YP, Wei Y, Wu M (2001) Molecular therapy with recombinant antisense c-myc adenovirus for human gastric carcinoma cells in vitro and in vivo. J Gastroenterol Hepatol 16: 22–28
Chung CD, Liao J, Liu B, Rao X, Jay P, Berta P, Shuai K (1997) Specific inhibition of Stat3 signal transduction by PIAS3. Science 278: 1803–1805
Dews M, Homayouni A, Yu D, Murphy D, Sevignani C, Wentzel E, Furth EE, Lee WM, Enders GH, Mendell JT, Thomas-Tikhonenko A (2006) Augmentation of tumour angiogenesis by a Myc-activated microRNA cluster. Nat Genet 38: 1060–1065
Epling-Burnette PK, Liu JH, Catlett-Falcone R, Turkson J, Oshiro M, Kothapalli R, Li Y, Wang JM, Yang-Yen HF, Karras J, Jove R, Loughran TP Jr (2001) Inhibition of STAT3 signaling leads to apoptosis of leukemic large granular lymphocytes and decreased Mcl-1 expression. J Clin Invest 107: 351–362
Filipowicz W, Bhattacharyya SN, Sonenberg N (2008) Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 9: 102–114
Grimson A, Farh KK, Johnston WK, Garrett-Engele P, Lim LP, Bartel DP (2007) MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell 27: 91–105
Guo J, Miao Y, Xiao B, Huan R, Jiang Z, Meng D, Wang Y (2009) Differential expression of microRNA species in human gastric cancer versus non-tumourous tissues. J Gastroenterol Hepatol 24: 652–657
Hayashita Y, Osada H, Tatematsu Y, Yamada H, Yanagisawa K, Tomida S, Yatabe Y, Kawahara K, Sekido Y, Takahashi T (2005) A polycistronic microRNA cluster, miR-17–92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res 65: 9628–9632
Ivanov VN, Bhoumik A, Krasilnikov M, Raz R, Owen-Schaub LB, Levy D, Horvath CM, Ronai Z (2001) Cooperation between STAT3 and c-jun suppresses Fas transcription. Mol Cell 7: 517–528
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D (2011) Global cancer statistics. CA Cancer J Clin 61: 69–90
Junicho A, Matsuda T, Yamamoto T, Kishi H, Korkmaz K, Saatcioglu F, Fuse H, Muraguchi A (2000) Protein inhibitor of activated STAT3 regulates androgen receptor signaling in prostate carcinoma cells. Biochem Biophys Res Commun 278: 9–13
Kondo S, Shinomura Y, Kanayama S, Higashimoto Y, Miyagawa JI, Minami T, Kiyohara T, Zushi S, Kitamura S, Isozaki K, Matsuzawa Y (1996) Over-expression of bcl-xL gene in human gastric adenomas and carcinomas. Int J Cancer 68: 727–730
Kubo M, Hanada T, Yoshimura A (2003) Suppressors of cytokine signaling and immunity. Nat Immunol 4: 1169–1176
Landgraf P, Rusu M, Sheridan R, Sewer A, Iovino N, Aravin A, Pfeffer S, Rice A, Kamphorst AO, Landthaler M, Lin C, Socci ND, Hermida L, Fulci V, Chiaretti S, Foa R, Schliwka J, Fuchs U, Novosel A, Muller RU, Schermer B, Bissels U, Inman J, Phan Q, Chien M, Weir DB, Choksi R, De Vita G, Frezzetti D, Trompeter HI, Hornung V, Teng G, Hartmann G, Palkovits M, Di Lauro R, Wernet P, Macino G, Rogler CE, Nagle JW, Ju J, Papavasiliou FN, Benzing T, Lichter P, Tam W, Brownstein MJ, Bosio A, Borkhardt A, Russo JJ, Sander C, Zavolan M, Tuschl T (2007) A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129: 1401–1414
Matsubara H, Takeuchi T, Nishikawa E, Yanagisawa K, Hayashita Y, Ebi H, Yamada H, Suzuki M, Nagino M, Nimura Y, Osada H, Takahashi T (2007) Apoptosis induction by antisense oligonucleotides against miR-17-5p and miR-20a in lung cancers overexpressing miR-17–92. Oncogene 26: 6099–6105
Mendell JT (2008) miRiad roles for the miR-17–92 cluster in development and disease. Cell 133: 217–222
Pichiorri F, Suh SS, Ladetto M, Kuehl M, Palumbo T, Drandi D, Taccioli C, Zanesi N, Alder H, Hagan JP, Munker R, Volinia S, Boccadoro M, Garzon R, Palumbo A, Aqeilan RI, Croce CM (2008) MicroRNAs regulate critical genes associated with multiple myeloma pathogenesis. Proc Natl Acad Sci USA 105: 12885–12890
Poliseno L, Salmena L, Riccardi L, Fornari A, Song MS, Hobbs RM, Sportoletti P, Varmeh S, Egia A, Fedele G, Rameh L, Loda M, Pandolfi PP (2010) Identification of the miR-106b∼25 microRNA cluster as a proto-oncogenic PTEN-targeting intron that cooperates with its host gene MCM7 in transformation. Sci Signal 3: ra29
Shuai K, Liu B (2005) Regulation of gene-activation pathways by PIAS proteins in the immune system. Nat Rev Immunol 5: 593–605
Tanaka M, Oikawa K, Takanashi M, Kudo M, Ohyashiki J, Ohyashiki K, Kuroda M (2009) Down-regulation of miR-92 in human plasma is a novel marker for acute leukemia patients. PLoS One 4: e5532
Valladares-Ayerbes M, Blanco M, Haz M, Medina V, Iglesias-Diaz P, Lorenzo-Patino MJ, Reboredo M, Santamarina I, Figueroa A, Anton-Aparicio LM, Calvo L (2011) Prognostic impact of disseminated tumour cells and microRNA-17-92 cluster deregulation in gastrointestinal cancer. Int J Oncol 39: 1253–1264
Ventura A, Jacks T (2009) MicroRNAs and cancer: short RNAs go a long way. Cell 136: 586–591
Wang J, Wang Q, Liu H, Hu B, Zhou W, Cheng Y (2010) MicroRNA expression and its implication for the diagnosis and therapeutic strategies of gastric cancer. Cancer Lett 297: 137–143
Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M, Stephens RM, Okamoto A, Yokota J, Tanaka T, Calin GA, Liu CG, Croce CM, Harris CC (2006) Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 9: 189–198
Yao Y, Suo AL, Li ZF, Liu LY, Tian T, Ni L, Zhang WG, Nan KJ, Song TS, Huang C (2009) MicroRNA profiling of human gastric cancer. Mol Med Report 2: 963–970
Yokozaki H (2000) Molecular characteristics of eight gastric cancer cell lines established in Japan. Pathol Int 50: 767–777
Yu CL, Meyer DJ, Campbell GS, Larner AC, Carter-Su C, Schwartz J, Jove R (1995) Enhanced DNA-binding activity of a Stat3-related protein in cells transformed by the Src oncoprotein. Science 269: 81–83
Yu H, Jove R (2004) The STATs of cancer – new molecular targets come of age. Nat Rev Cancer 4: 97–105
Yu J, Leung WK, Ebert MP, Ng EK, Go MY, Wang HB, Chung SC, Malfertheiner P, Sung JJ (2002) Increased expression of survivin in gastric cancer patients and in first degree relatives. Br J Cancer 87: 91–97
Acknowledgements
This work was supported by Grants-in-Aids from the Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT); the Ministry of Health, Labour and Welfare of Japan; and the ‘Private University Strategic Research Based Support Project: Epigenetics research project aimed at general cancer cure using epigenetic targets’ from MEXT.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Wu, W., Takanashi, M., Borjigin, N. et al. MicroRNA-18a modulates STAT3 activity through negative regulation of PIAS3 during gastric adenocarcinogenesis. Br J Cancer 108, 653–661 (2013). https://doi.org/10.1038/bjc.2012.587
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2012.587
Keywords
This article is cited by
-
Molecular targeting of vulnerable RNA sequences in SARS CoV-2: identifying clinical feasibility
Gene Therapy (2022)
-
Circular RNA circRHOBTB3 is downregulated in hepatocellular carcinoma and suppresses cell proliferation by inhibiting miR-18a maturation
Infectious Agents and Cancer (2021)
-
Noncoding RNAs: the shot callers in tumor immune escape
Signal Transduction and Targeted Therapy (2020)
-
Epstein–Barr virus-encoded miR-BART5-5p upregulates PD-L1 through PIAS3/pSTAT3 modulation, worsening clinical outcomes of PD-L1-positive gastric carcinomas
Gastric Cancer (2020)
-
The dual functional role of MicroRNA‐18a (miR‐18a) in cancer development
Clinical and Translational Medicine (2019)