Abstract
Background:
We previously conducted gene expression microarray analyses to identify novel indicators for colorectal cancer (CRC) metastasis and prognosis from which we identified PVT-1 as a candidate gene. PVT-1, which encodes a long noncoding RNA, mapped to chromosome 8q24 whose copy-number amplification is one of the most frequent events in a wide variety of malignant diseases. However, PVT-1 molecular mechanism of action remains unclear.
Methods:
We conducted cell proliferation and invasion assays using colorectal cancer cell lines transfected with PVT-1siRNA or negative control siRNA. Gene expression microarray analyses on these cell lines were also carried out to investigate the molecular function of PVT-1. Further, we investigated the impact of PVT-1 expression on the prognosis of 164 colorectal cancer patients by qRT–PCR.
Results:
CRC cells transfected with PVT-1 siRNA exhibited significant loss of their proliferation and invasion capabilities. In these cells, the TGF-β signalling pathway and apoptotic signals were significantly activated. In addition, univariate and multivariate analysis revealed that PVT-1 expression level was an independent risk factor for overall survival of colorectal cancer patients.
Conclusion:
PVT-1, which maps to 8q24, generates antiapoptotic activity in CRC, and abnormal expression of PVT-1 was a prognostic indicator for CRC patients.
Similar content being viewed by others
Main
To identify novel indicators for colorectal cancer (CRC) metastasis and prognosis, we previously conducted gene expression array analysis using clinical samples resulting in the identification of several genes, which are involved in cancer progression and metastasis (Takahashi et al, 2013). The candidate genes include the PVT-1 oncogene (PVT-1), which encodes a long noncoding RNA (lncRNA) and maps to chromosome 8q24 (8q24) (Carninci et al, 2005; Amaral et al, 2008; Guttman et al, 2009). Amplification of 8q24 is one of the most frequent events in a wide variety of malignant diseases including CRC, and has been associated with reduced survival duration in several studies (Popescu and Zimonjic, 2002; Lancaster et al, 2004; Guan et al, 2007). In addition to PVT-1, the well-established oncogene MYC has also been mapped to 8q24, and these two genes are co-amplified in CRC cell lines (Shtivelman and Bishop, 1989).
lncRNAs were previously believed to represent random transcriptional noise. However, their expression levels have been observed to vary spatially, temporally and in response to various stimuli (Mercer et al, 2009; Ponting et al, 2009). Moreover, several lncRNAs exhibit very precise expression patterns in different tissues (Amaral and Mattick, 2008; Mercer et al, 2008). In spite of emerging evidence, our understanding of the functions of these lncRNAs is limited. PVT-1 produces a wide variety of spliced noncoding RNAs as well as a cluster of six annotated microRNAs: miR-1204, miR-1205, miR-1206, miR-1207-5p, miR-1207-3p and miR-1208 (Barsotti et al, 2012). However, PVT-1 exerts its influence as an lncRNA (Guan et al, 2007; Alvarez and DiStefano, 2011; You et al, 2011). The molecular mechanism of PVT-1 transcripts in gene regulation remains unclear. In the current study, we investigated the clinical significance of PVT-1 expression in CRC.
Materials and methods
Patients and sample collection
We used a total of 312 CRC samples, of which 148 (set 1) were used as pure cancer tissues separated by laser microdissection (all cases were used for gene expression array and 130 cases were used for array-CGH), and 164 (set 2) were used in bulk for quantitative reverse transcription–PCR. All samples were obtained during surgery from patients who underwent resection of the primary tumour at Kyushu University Hospital at Beppu and affiliated hospitals between 1992 and 2007. Written informed consent was obtained from all patients. All patients had a clear histological diagnosis of CRC and were closely followed up every 3 months. The follow-up period in set 1 ranged from 0.1 months to 3.2 years with a mean of 2.1 years; follow-up in set 2 ranged from 0.1 to 12.3 years, with a mean of 3.8 years. Resected cancer tissues were immediately cut and stored in RNAlater (Ambion, Palo Alto, CA, USA) or embedded in Tissue-Tek OCT (optimum cutting temperature) medium (Sakura, Tokyo, Japan), frozen in liquid nitrogen and kept at −80 °C until DNA and RNA extraction. For DNA extraction, the QIAamp DNA Micro Kit (Qiagen, Hilden, Germany) was used following the manufacturer’s protocol. For RNA extraction, frozen tissue specimens were homogenised in guanidinium thiocyanate, and total RNA was obtained by ultracentrifugation through a caesium chloride cushion. cDNA for reverse transcription–PCR was synthesised from 8.0 μg of total RNA with M-MLV Reverse Transcriptase (Invitrogen, Carlsbad, CA, USA). Clinicopathological factors and clinical stage were classified using the TNM system of classification. All sample data, including age, gender, histology, tumour depth, lymph node metastasis, lymphatic invasion, vascular invasion, liver metastasis and postoperative liver recurrence, were obtained from the clinical and pathological records.
Laser microdissection
Tissue samples were microdissected using an LMD6000 laser microdissection system (Leica Laser Microdissection System; Leica Microsystems, Wetzlar, Germany). For LMD, 5 μm-thick frozen sections were fixed in 70% ethanol for 30 s, stained with hematoxylin and eosin and de-hydrated as follows: 5 s each in 70%, 95% and 100% ethanol. Sections were air dried and then microdissected with the LMD system. Target cells were excised, with at least 100 cells per section, and bound to the transfer film. Total RNA was then extracted.
Quantitative real-time reverse transcription–PCR
For quantitative real-time reverse transcription (qRT)–PCR, PVT-1 (NR_003367.1) primer sequences were 5′-TGAGAACTGTCCTTACGTGACC-3′ and 5′-AGAGCACCAAGACTGGCTCT-3′, and MYC (NM_002467.4) primer sequences were 5′-CACCAGCAGCGACTCTGA-3′ and 5′-GATCCAGACTCTGACCTTTTGC-3′. To normalise for RNA concentration, glyceraldehyde-3-phosphate dehydrogenase (GAPDH) served as an internal control. The sequences of the GAPDH primers were sense, 5′-TTGGTATCGTGGAAGGACTCA-3′ and antisense, 5′-TGTCATCATATTTGGCAGGTT-3′. The amplification protocol included an initial denaturation step at 95 °C for 10 min, followed by 45 cycles of 95 °C for 10 s and 60 °C for 30 s. qRT–PCR was performed in a LightCycler 480 instrument (Roche Applied Science, Basel, Switzerland) using the LightCycler 480 Probes Master kit (Roche Applied Science). All concentrations were calculated relative to the concentration of cDNA using Human Universal Reference Total RNA (Clontech, Palo Alto, CA, USA). The concentration of PVT-1 was then divided by the concentration of the endogenous reference (GAPDH) to obtain normalised expression values. For qRT–PCR, for miR-1204, miR-1205, miR-1206, miR-1207-3p, miR-1207-5p and miR-1208, cDNA was synthesised from total RNA using TaqMan MicroRNA specific primers for each microRNAs (Applied Biosystems, Foster City, CA, USA) and a TaqMan MicroRNA Reverse Transcription kit (Applied Biosystems). Expression of target miRNAs was normalised to the expression of a small nuclear RNA, RNU6B (Applied Biosystems).
Cell lines
The human colorectal cancer cell lines RKO and HCT116 were obtained from the Japanese Cancer Research Bank (Tokyo, Japan) and maintained in Dulbecco’s modified Eagle’s medium containing 10% fetal bovine serum, 100 units ml−1 penicillin and 100 μg ml−1 streptomycin sulphate. All cells were cultured in a humidified 5% CO2 incubator at 37 °C.
PVT-1 siRNA transfection
PVT-1-specific siRNAs (PVT-1siRNA-1: sense GCUUGGAGGCUGAGGAGUUTT and antisense AACUCCUCAGCCUCCAAGCTT and PVT-1 siRNA-2: sense CCCAACAGGAGGACAGCUUTT and antisense AAGCUGUCCUCCUGUUGGGTT) and negative control siRNA were purchased from Cosmo Bio (Tokyo, Japan). siRNA oligonucleotide (30 nmol l−1) in Opti-MEM (Invitrogen) were transfected into RKO and HCT116 cells using Lipofectamine RNAiMAX (Invitrogen) following the manufacturer’s protocol. Cells in logarithmic growth phase were diluted without antibiotics and were seeded at 2 × 105 or 5 × 104 cells per well in a final volume of 2 ml or 500 μl, respectively, in 6- or 24-well flat-bottom microtiter plates, respectively. The cells were incubated in a humidified atmosphere (37 °C and 5% CO2) for 24 h before use in assays.
Cell proliferation assay and cell invasion assay
To conduct the cell proliferation assay, 10 μl of MTT-labelling reagent (at a final concentration of 0.5 mg ml−1) was added to each well, and the plate was incubated for 4 h in a humidified atmosphere. Solubilisation solution (100 μl) was added to each well, and the plate was incubated overnight in a humidified atmosphere. After confirming that the purple formazan crystals were completely solubilised, the absorbance of each well was measured using a Model 550 series microplate reader (Bio-Rad Laboratories, Hercules, CA, USA) at a wavelength of 570 nm corrected to 655 nm. The assay was performed using six replicates.
Invasion assays were done using the BD Biocoat Matrigel Invasion Chamber (pore size: 8 μm, 24-well; BD Biosciences, San Jose, CA, USA) following the manufacturer's protocol. A total of 5 × 104 cells were plated in the upper chamber in serum-free medium. The bottom chamber contained medium with 10% FBS. After 72 h, the bottom of the chamber insert was stained with Calcein AM (Invitrogen). The cells that had invaded through the membrane to the lower surface were evaluated in a fluorescence plate reader at excitation/emission wavelengths of 485/530 nm. Each assay was conducted in at least three replicates.
Western blotting
Total protein was extracted from each cell line. Aliquots of total protein (40 μg) were electrophoresed in 10% concentrated poly-acrylamide gel and then electrophoresed and then electroblotted as previously described (Ieta et al, 2007). The primary mouse monoclonal antibodies against caspase 3 (Cell Signaling Technology, Danvers, MA, USA), SMAD4 (Santa Cruz Biotechnbology, Dallas, TX, USA) were used at a dilution of 1 : 1000 and 1 : 200, respectively. Primary antibodies were detected using horseradish peroxidase (HRP)-conjugated secondary antibodies (GE Healthcare Japan, Tokyo, Japan) and the ImageQuant LAS 4000mini system (GE Healthcare Japan). Caspase 3 and SMAD4 proteins were normalised to the level of β-tubulin protein (Cell Signaling Technology) diluted 1 : 1000.
Array-CGH and copy-number analysis
A total of 130 tumour samples were prepared for array-CGH analysis. For genome profiling, the labelling and hybridisation of genomic DNA onto the Agilent Human Genome Microarray Kit 244 K (Agilent Technologies, Santa Clara, CA, USA) were performed according to the manufacturer’s instructions. The raw signal intensities were measured and transformed into log ratios to reference DNA with ‘Feature Extraction’ software (version 9.1) from Agilent Technologies. Arrays were analysed using the Agilent DNA microarray scanner. The log ratio was thereafter used as the signal intensity of each probe. The raw copy-number data for each sample provided by array CGH was analysed using the GISTIC algorithm (Beroukhim et al, 2007; Mermel et al, 2011).
Gene expression array
The commercially available whole-Human Genome Microarray Kit and SurePrint G3 Human GE Microarray Kit (Agilent Technologies) were used to investigate globally altered genes. In brief, cyanine-labelled cRNA was prepared using the T7 linear amplification technique described in the Agilent Low RNA Input Fluorescent Linear Amplification Kit Manual (Agilent Technologies). Hundred nanograms of total RNA was extracted using the LMD technique and reverse transcribed to generate double-stranded cDNA using an oligo dT T7 promoter primer. Subsequently, cRNA was synthesised using T7 RNA polymerase, which simultaneously incorporated Cy3-labelled CTP. The quality of the cRNA was verified with the Agilent Bioanalyzer 2100. One microgram aliquots of Cy3-labelled cRNA were combined and fragmented in a hybridisation cocktail (Agilent Technologies). Labelled cRNA was then fragmented and hybridised to an oligonucleotide microarray. Fluorescence intensities were determined with an Agilent DNA microarray scanner. The gene expression profiles obtained from microarray data were quantile normalised. The batch effect in microarray experiments was also adjusted by an empirical Bayesian approach (Johnson et al, 2007).
Gene set enrichment analysis
For GSEA, PVT-1 expression was treated as a numeric variable. We applied a continuous-type CLS file of the PVT-1 profile to phenotype labels in GSEA. The metric for ranking genes in GSEA was set as ‘pearson,’ and the other parameters were set to their default values (Subramanian et al, 2005).
Statistical analysis
For continuous variables, data were expressed as means±s.d. The relationship between PVT-1 expression and clinicopathological factors was analysed using a χ2-test and Student’s t-test. Findings were considered significant when the P-value was <0.05. Correlation coefficients were analysed using the Pearson product-moment correlation coefficient. All tests were performed using the JMP software (SAS Institute Inc., Cary, NC, USA).
Results
8q24 copy-number amplification and PVT-1 expression
First, we applied array-CGH and gene expression array analyses on 130 CRC tumours to investigate the correlation between chromosome 8q24 copy-number and PVT-1 expression, and significant correlation was observed (Figure 1A; R2=0.181, P<0.0001). This result was validated with the Cancer Cell Line Encyclopedia (CCLE, http://www.broadinstitute.org/ccle/home), which is a recently compiled public resource that contains gene expression and chromosomal copy-number data from nearly 1000 cancer cell lines(Barretina et al, 2012). A significant positive correlation between 8q24 copy-number and PVT-1 expression was also observed in 50 CRC cell lines (Figure 1B; R2=0.420, P<0.0001). These findings showed that 8q24 copy-number gain promoted PVT-1 expression and suggested that aberrant expression of PVT-1, which we previously identified to be a candidate gene for CRC metastasis and progression (Takahashi et al, 2013), was of significance accompanied by genomic alteration in CRC. As MYC is a well-established oncogene, which also maps to 8q24, we investigated the correlation between 8q24 copy-numbers and MYC expression. Copy-numbers of 8q24 and MYC expression were also positively correlated in CRC tissues and CRC cell lines. However, the correlation between MYC and 8q24 copy-numbers was weaker than that between PVT-1 and 8q24 copy-numbers (Figures 1C and D).
Positive correlation between chromosome 8q24 copy-number and PVT-1 or MYC expression. (A) Chromosome 8q24 copy-number (horizontal axis) and PVT-1 expression (vertical axis) analysed by array-CGH and gene expression array in 130 colorectal cancer cases. Dot plots indicate each case. (B) Chromosome 8q24 copy-number (horizontal axis) and PVT-1 expression (vertical axis) in 50 colorectal cancer cell lines (the Cancer Cell Line Encyclopedia database). Dot plots indicate each cell line. (C) Chromosome 8q24 copy-number (horizontal axis) and MYC expression (vertical axis) analysed by array-CGH and gene expression array in 130 colorectal cancer cases. Dot plots indicate each case. (D) Chromosome 8q24 copy-number (horizontal axis) and MYC expression (vertical axis) in 50 colorectal cancer cell lines (the CCLE database). Dot plots indicate each cell line.
Knockdown of PVT-1 promotes apoptosis in colorectal cancer cell lines
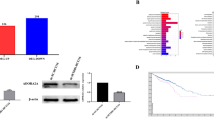
Next, we conducted PVT-1 knockdown assays using siRNA to investigate the biological function of PVT-1 transcripts in CRC cells. Using qRT–PCR, we confirmed that PVT-1 expression in RKO and HCT116 cells transfected with PVT-1-specific siRNA was significantly lower than that in cells transfected with the negative control siRNA (Figure 2A). Cell proliferation assays were carried out in RKO and HCT116 colorectal cancer cells transfected with PVT-1 siRNA or negative control siRNA. Both RKO and HCT116 cell lines have 8q24 copy-number amplification and PVT-1 upregulation according to the CCLE database. PVT-1 knockdown significantly inhibited cancer cell proliferation in both cell lines (Figure 2B). This finding suggested that aberrant expression of PVT-1 promoted the cell proliferation capability of colorectal cancer cells. Next, we conducted cancer cell invasion assays to verify the effect of PVT-1 on metastasis, since we had originally identified PVT-1 as a metastasis regulating gene. Then we observed that PVT-1 siRNA transfected cells had reduced invasive abilities compared with negative control cells (Figure 2C).
Knockdown of PVT-1 promoted apoptosis in colorectal cancer cells. (A) qRT–PCR analysis showed that PVT-1 expression levels of both cell lines transfected with PVT-1 siRNA were significantly lower than negative control (nc) cells. (B) Cell proliferation ratio of RKO cells (left panel) and HCT116 cells (right panel) with or without PVT-1 knockdown. (C) Invasive properties of RKO cells (left panel) and HCT116 cells (right panel) with or without PVT-1 knockdown. (D) A total of 1187 genes with significant alteration of expression levels (649 upregulated and 538 downregulated) by knockdown of PVT-1(P<0.05). Representative genes are shown in the right side of the heat map. (E) Western blot (left, RKO; right, HCT116) analyses of cells transfected with PVT-1 siRNA, and negative control cells show the activation of SMAD4 and cleavage of caspase3. Histograms represent the means±s.d. of three independent experiments. nc, cells transfected with negative control siRNA. si-1, cells transfected with PVT-1 siRNA-1. si-2, cells transfected with PVT-1 siRNA-2. *, statistically significant (P<0.05).
To uncover the underlying mechanisms of anticancer activity of PVT-1 knockdown, we performed gene expression microarray assays on cells transfected with PVT-1 siRNA and negative control siRNA. A total of 649 genes including several apoptosis-relating genes were upregulated and 538 genes including some oncogenes were downregulated by PVT-1 knockdown (Figure 2D, Supplementary Tables S1 and S2). Functional annotation analyses were performed using David Bioinformatics Resources 6.7, which is a web-based bioinformatics application (http://david.abcc.ncifcrf.gov/) (Imamichi et al, 2012), and we found that genes in the pathways for transforming growth factor beta (TGF-β) signalling and apoptosis were activated in cells with PVT-1 knockdown (Supplementary Table S3). Actually, western blot analyses showed that the protein expression of SMAD4, which is in the TGF-β signalling pathway, was induced, and the cleavage of caspase 3 was observed in PVT-1 knockdown cells (Figure 2E). These data showed that apoptosis was induced to CRC cells transfected with PVT-1 siRNA via activation of TGF-β signalling. In order to confirm whether apoptosis was induced by knockdown of PVT-1 lncRNA or by knockdown of microRNAs encoded by PVT-1, which seemed to be downregulated following knockdown of PVT-1 transcripts, we conducted qRT–PCR assays for these six microRNAs in CRC cells transfected with PVT-1 siRNA and negative control siRNA. Five of six microRNAs were detected by qRT–PCR, and no microRNAs were downregulated by PVT-1 knockdown in CRC cells (Supplementary Figure S1).
PVT-1 expression is a prognostic indicator for colorectal cancer patients
PVT-1 expression levels in 164 tumour tissues and corresponding normal tissues were examined by qRT–PCR to investigate the clinical significance of PVT-1 in CRC. PVT-1 expression levels in cancerous tissues were significantly higher than those in non-cancerous tissues (P<0.0001; Figure 3A). We divided the 164 patients with CRC into a high PVT-1 expression group (n=131) and a low PVT-1 expression group (n=33), classified as having expression levels higher or lower than the 20 percentile value, respectively, which is an approximation of the highest value among normal tissues. Clinicopathological factors were then analysed in the high and low PVT-1 expression groups (Table 1). The high PVT-1 expression group showed greater lymph node metastasis and venous invasion compared with the low PVT-1 expression group. With regard to overall survival, patients with high PVT-1 expression had a significantly poorer prognosis than those with low PVT-1 expression (P=0.0101; Figure 3B). Univariate and multivariate analysis showed that PVT-1 expression level was an independent prognostic indicator of overall survival in patients with CRC (relative risk: 2.532, P=0.016; Table 2). The clinical significance of MYC expression was also investigated in the same 164 colorectal cancer cases, as MYC is another important oncogene that maps to 8q24. MYC expression levels in cancerous tissues were significantly higher than those in non-cancerous tissues (P<0.0001; Figure 3C). In addition, we analysed the impact of MYC expression on colorectal cancer patients. We divided the 164 patients with colorectal cancer into a high MYC expression group (n=98) and a low PVT-1 expression group (n=66), classified as having expression levels higher or lower than the 40 percentile value, respectively, which is an approximation of the highest MYC expression value among normal tissues. However, the significance of MYC as a prognosticator for overall survival was not demonstrated in this case set (Figure 3D).
Abundant PVT-1 expression is involved in poor prognosis in colorectal cancer. (A) qRT–PCR analyses of PVT-1expression levels on tumour tissues and paired normal tissues from 164 colorectal cancer samples. (B) Kaplan–Meier overall survival curves for 164 patients with CRC classified according to PVT-1 expression level. (C) qRT–PCR analyses of MYC expression levels on tumour tissues and paired normal tissues from 164 colorectal cancer samples. (D) Kaplan–Meier overall survival curves for 164 patients with CRC classified according to MYC expression level.
Discussion
Several published reports have revealed that PVT-1 is involved in cancer pathophysiology. In the current study, we focused on the clinical significance of aberrant PVT-1 expression caused by copy-number amplification of chromosome 8q24. The MYC oncogene is also a notable oncogene mapping to 8q24 (Popescu and Zimonjic, 2002; Lancaster et al, 2004; Guan et al, 2007). However, the prognostic value of MYC expression was not verified in our case set. Although these data do not contradict the previously demonstrated significance of the MYC oncogene, the meaning of PVT-1 in CRC pathophysiology was strongly suggested.
In our gene expression microarray analysis of CRC cell lines (Figure 2D), genes relating to the TGF-β signalling pathway were upregulated by PVT-1 knockdown. SMAD4, a well-known tumour suppressor gene, is a major mediator of intracellular TGF-β signalling (Heldin et al, 1997; Massague, 1998) and inhibits tumour growth by inducing apoptosis in cancer cells (Brodin et al, 1999; Dai et al, 1999; Jang et al, 2002; Li et al, 2005). In the current study, SMAD4 and apoptosis-relating genes were upregulated by PVT-1 knockdown, suggesting that PVT-1 knockdown promoted apoptosis via TGF-β signalling activation in CRC cells. On the other hand, ROCK1, which is another mediator of TGF-β signalling and promotes cancer invasiveness, was also activated in PVT-1 knockdown cells. However, PVT-1 knockdown cells lose their invasive capability (Figure 2B). We consider that this is because apoptosis has more impact on the phenotypes of CRC cells than ROCK1 activation. Guan et al, have reported that PVT-1 inhibits apoptosis independently of MYC in breast and ovarian cancer (Guan et al, 2007), and our results support their findings. However, the functions of microRNAs encoded by PVT-1 must be taken into account. miR-1207-5p was an only microRNA deregulated by PVT-1 knockdown, and the expression level of miR-1207-5p was upregulated following PVT-1 knockdown (Supplementary Figure S1). miR-1207-5p was predicted to repress CASP9, using the microRNA target prediction online tool, TargetScan (http://www.targetscan.org/), which is involved in apoptosis signal and is one of the representative genes deregulated by PVT-1 knockdown (shown in Figure 3D). Namely, miR-1207-5p was thought to inhibit apoptosis. Therefore, we consider that these microRNAs are regulated by other transcriptional mechanism and that these microRNAs are not involved in apoptosis induced by PVT-1 knockdown.
Although only a small number of functional lncRNAs have been well characterised to date, emerging evidence has suggested that lncRNAs have roles as drivers of tumour suppressive or oncogenic functions in a wide variety of cancer types. In addition, many researchers have suggested mechanisms of action underlying the regulation of gene expression by lncRNAs (Wang and Chang, 2011). For example, lncRNAs can regulate chromosome structure in cis (Jonkers et al, 2008) or in trans (Rinn et al, 2007; Kogo et al, 2011). Other lncRNAs modulate the activity of protein-binding partners (Espinoza et al, 2004; Mariner et al, 2008). However, the molecular function of PVT-1 is still controversial. However, we speculate that PVT-1 regulates chromatin modification by targeting the SWItch/Sucrose NonFermentable (SWI/SNF) complex, which has tumour suppressive function (Roberts and Orkin, 2004; Tolstorukov et al, 2013). GSEA on clinical CRC samples suggested that PVT-1 modulated the expression levels of target genes of SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 (SMARCA2) that constitute an ATPase subunit of SWI/SNF complex (Supplementary Figures S2A and B) and that PVT-1 was associated with the apoptosis pathway (Supplementary Figures S2C). In addition, Shen et al. have reported the positive correlation between the expression of SMARCA2, SMAD4 and other genes on the TGF-β pathway (Shen et al, 2008). Our in vitro study also supported these findings (Supplementary Figure S3). However, a limitation of this current study is that we did not demonstrate a direct molecular function of PVT-1 in apoptosis induction clearly, and further investigation is needed.
In summary, we demonstrated that PVT-1, which encodes an lncRNA and maps to 8q24, generates antiapoptotic activity in colorectal cancer cells and that abnormal expression of PVT-1 was a prognostic indicator for colorectal cancer patients. Further study is required to understand the underlying mechanism of apoptosis inhibition by PVT-1.
Change history
07 January 2014
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Alvarez ML, DiStefano JK (2011) Functional characterization of the plasmacytoma variant translocation 1 gene (PVT1) in diabetic nephropathy. PLoS One 6 (4): e18671.
Amaral PP, Dinger ME, Mercer TR, Mattick JS (2008) The eukaryotic genome as an RNA machine. Science 319 (5871): 1787–1789.
Amaral PP, Mattick JS (2008) Noncoding RNA in development. Mamm Genome 19 (7–8): 454–492.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, Wilson CJ, Lehar J, Kryukov GV, Sonkin D, Reddy A, Liu M, Murray L, Berger MF, Monahan JE, Morais P, Meltzer J, Korejwa A, Jane-Valbuena J, Mapa FA, Thibault J, Bric-Furlong E, Raman P, Shipway A, Engels IH, Cheng J, Yu GK, Yu J, Aspesi P Jr, de Silva M, Jagtap K, Jones MD, Wang L, Hatton C, Palescandolo E, Gupta S, Mahan S, Sougnez C, Onofrio RC, Liefeld T, Mac Conaill L, Winckler W, Reich M, Li N, Mesirov JP, Gabriel SB, Getz G, Ardlie K, Chan V, Myer VE, Weber BL, Porter J, Warmuth M, Finan P, Harris JL, Meyerson M, Golub TR, Morrissey MP, Sellers WR, Schlegel R, Garraway LA (2012) The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483 (7391): 603–607.
Barsotti AM, Beckerman R, Laptenko O, Huppi K, Caplen NJ, Prives C (2012) p53-Dependent induction of PVT1 and miR-1204. J Biol Chem 287 (4): 2509–2519.
Beroukhim R, Getz G, Nghiemphu L, Barretina J, Hsueh T, Linhart D, Vivanco I, Lee JC, Huang JH, Alexander S, Du J, Kau T, Thomas RK, Shah K, Soto H, Perner S, Prensner J, Debiasi RM, Demichelis F, Hatton C, Rubin MA, Garraway LA, Nelson SF, Liau L, Mischel PS, Cloughesy TF, Meyerson M, Golub TA, Lander ES, Mellinghoff IK, Sellers WR (2007) Assessing the significance of chromosomal aberrations in cancer: methodology and application to glioma. Proc Natl Acad Sci USA 104 (50): 20007–20012.
Brodin G, ten Dijke P, Funa K, Heldin CH, Landstrom M (1999) Increased smad expression and activation are associated with apoptosis in normal and malignant prostate after castration. Cancer Res 59 (11): 2731–2738.
Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, Kodzius R, Shimokawa K, Bajic VB, Brenner SE, Batalov S, Forrest AR, Zavolan M, Davis MJ, Wilming LG, Aidinis V, Allen JE, Ambesi-Impiombato A, Apweiler R, Aturaliya RN, Bailey TL, Bansal M, Baxter L, Beisel KW, Bersano T, Bono H, Chalk AM, Chiu KP, Choudhary V, Christoffels A, Clutterbuck DR, Crowe ML, Dalla E, Dalrymple BP, de Bono B, Della Gatta G, di Bernardo D, Down T, Engstrom P, Fagiolini M, Faulkner G, Fletcher CF, Fukushima T, Furuno M, Futaki S, Gariboldi M, Georgii-Hemming P, Gingeras TR, Gojobori T, Green RE, Gustincich S, Harbers M, Hayashi Y, Hensch TK, Hirokawa N, Hill D, Huminiecki L, Iacono M, Ikeo K, Iwama A, Ishikawa T, Jakt M, Kanapin A, Katoh M, Kawasawa Y, Kelso J, Kitamura H, Kitano H, Kollias G, Krishnan SP, Kruger A, Kummerfeld SK, Kurochkin IV, Lareau LF, Lazarevic D, Lipovich L, Liu J, Liuni S, McWilliam S, Madan Babu M, Madera M, Marchionni L, Matsuda H, Matsuzawa S, Miki H, Mignone F, Miyake S, Morris K, Mottagui-Tabar S, Mulder N, Nakano N, Nakauchi H, Ng P, Nilsson R, Nishiguchi S, Nishikawa S, Nori F, Ohara O, Okazaki Y, Orlando V, Pang KC, Pavan WJ, Pavesi G, Pesole G, Petrovsky N, Piazza S, Reed J, Reid JF, Ring BZ, Ringwald M, Rost B, Ruan Y, Salzberg SL, Sandelin A, Schneider C, Schonbach C, Sekiguchi K, Semple CA, Seno S, Sessa L, Sheng Y, Shibata Y, Shimada H, Shimada K, Silva D, Sinclair B, Sperling S, Stupka E, Sugiura K, Sultana R, Takenaka Y, Taki K, Tammoja K, Tan SL, Tang S, Taylor MS, Tegner J, Teichmann SA, Ueda HR, van Nimwegen E, Verardo R, Wei CL, Yagi K, Yamanishi H, Zabarovsky E, Zhu S, Zimmer A, Hide W, Bult C, Grimmond SM, Teasdale RD, Liu ET, Brusic V, Quackenbush J, Wahlestedt C, Mattick JS, Hume DA, Kai C, Sasaki D, Tomaru Y, Fukuda S, Kanamori-Katayama M, Suzuki M, Aoki J, Arakawa T, Iida J, Imamura K, Itoh M, Kato T, Kawaji H, Kawagashira N, Kawashima T, Kojima M, Kondo S, Konno H, Nakano K, Ninomiya N, Nishio T, Okada M, Plessy C, Shibata K, Shiraki T, Suzuki S, Tagami M, Waki K, Watahiki A, Okamura-Oho Y, Suzuki H, Kawai J, Hayashizaki Y, Consortium F Group RGER, Genome Science G (2005) The transcriptional landscape of the mammalian genome. Science 309 (5740): 1559–1563.
Dai JL, Bansal RK, Kern SE (1999) G1 cell cycle arrest and apoptosis induction by nuclear Smad4/Dpc4: phenotypes reversed by a tumorigenic mutation. Proc Natl Acad Sci USA 96 (4): 1427–1432.
Espinoza CA, Allen TA, Hieb AR, Kugel JF, Goodrich JA (2004) B2 RNA binds directly to RNA polymerase II to repress transcript synthesis. Nat Struct Mol Biol 11 (9): 822–829.
Guan Y, Kuo WL, Stilwell JL, Takano H, Lapuk AV, Fridlyand J, Mao JH, Yu M, Miller MA, Santos JL, Kalloger SE, Carlson JW, Ginzinger DG, Celniker SE, Mills GB, Huntsman DG, Gray JW (2007) Amplification of PVT1 contributes to the pathophysiology of ovarian and breast cancer. Clin Cancer Res 13 (19): 5745–5755.
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, Cabili MN, Jaenisch R, Mikkelsen TS, Jacks T, Hacohen N, Bernstein BE, Kellis M, Regev A, Rinn JL, Lander ES (2009) Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458 (7235): 223–227.
Heldin CH, Miyazono K, ten Dijke P (1997) TGF-beta signalling from cell membrane to nucleus through SMAD proteins. Nature 390 (6659): 465–471.
Ieta K, Ojima E, Tanaka F, Nakamura Y, Haraguchi N, Mimori K, Inoue H, Kuwano H, Mori M (2007) Identification of overexpressed genes in hepatocellular carcinoma, with special reference to ubiquitin-conjugating enzyme E2C gene expression. Int J Cancer 121 (1): 33–38.
Imamichi T, Yang J, Huang da W, Sherman B, Lempicki RA (2012) Interleukin-27 induces interferon-inducible genes: analysis of gene expression profiles using Affymetrix microarray and DAVID. Methods Mol Biol 820: 25–53.
Jang CW, Chen CH, Chen CC, Chen JY, Su YH, Chen RH (2002) TGF-beta induces apoptosis through Smad-mediated expression of DAP-kinase. Nat Cell Biol 4 (1): 51–58.
Johnson WE, Li C, Rabinovic A (2007) Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8 (1): 118–127.
Jonkers I, Monkhorst K, Rentmeester E, Grootegoed JA, Grosveld F, Gribnau J (2008) Xist RNA is confined to the nuclear territory of the silenced X chromosome throughout the cell cycle. Mol Cell Biol 28 (18): 5583–5594.
Kogo R, Shimamura T, Mimori K, Kawahara K, Imoto S, Sudo T, Tanaka F, Shibata K, Suzuki A, Komune S, Miyano S, Mori M (2011) Long noncoding RNA HOTAIR regulates polycomb-dependent chromatin modification and is associated with poor prognosis in colorectal cancers. Cancer Res 71 (20): 6320–6326.
Lancaster JM, Dressman HK, Whitaker RS, Havrilesky L, Gray J, Marks JR, Nevins JR, Berchuck A (2004) Gene expression patterns that characterize advanced stage serous ovarian cancers. J Soc Gynecol Invest 11 (1): 51–59.
Li Q, Wu L, Oelschlager DK, Wan M, Stockard CR, Grizzle WE, Wang N, Chen H, Sun Y, Cao X (2005) Smad4 inhibits tumor growth by inducing apoptosis in estrogen receptor-alpha-positive breast cancer cells. J Biol Chem 280 (29): 27022–27028.
Mariner PD, Walters RD, Espinoza CA, Drullinger LF, Wagner SD, Kugel JF, Goodrich JA (2008) Human Alu RNA is a modular transacting repressor of mRNA transcription during heat shock. Mol Cell 29 (4): 499–509.
Massague J (1998) TGF-beta signal transduction. Annu Rev Biochem 67: 753–791.
Mercer TR, Dinger ME, Mattick JS (2009) Long non-coding RNAs: insights into functions. Nat Rev Genet 10 (3): 155–159.
Mercer TR, Dinger ME, Sunkin SM, Mehler MF, Mattick JS (2008) Specific expression of long noncoding RNAs in the mouse brain. Proc Natl Acad Sci USA 105 (2): 716–721.
Mermel CH, Schumacher SE, Hill B, Meyerson ML, Beroukhim R, Getz G (2011) GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol 12 (4): R41.
Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136 (4): 629–641.
Popescu NC, Zimonjic DB (2002) Chromosome-mediated alterations of the MYC gene in human cancer. J Cell Mol Med 6 (2): 151–159.
Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, Chang HY (2007) Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129 (7): 1311–1323.
Roberts CW, Orkin SH (2004) The SWI/SNF complex--chromatin and cancer. Nat Rev Cancer 4 (2): 133–142.
Shen H, Powers N, Saini N, Comstock CE, Sharma A, Weaver K, Revelo MP, Gerald W, Williams E, Jessen WJ, Aronow BJ, Rosson G, Weissman B, Muchardt C, Yaniv M, Knudsen KE (2008) The SWI/SNF ATPase Brm is a gatekeeper of proliferative control in prostate cancer. Cancer Res 68 (24): 10154–10162.
Shtivelman E, Bishop JM (1989) The PVT gene frequently amplifies with MYC in tumor cells. Mol Cell Bio 9 (3): 1148–1154.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102 (43): 15545–15550.
Takahashi Y, Sawada G, Sato T, Kurashige J, Mima K, Matsumura T, Uchi R, Ueo H, Ishibashi M, Takano Y, Akiyoshi S, Eguchi H, Sudo T, Sugimachi K, Tanaka J, Kudo S, Doki Y, Mori M, Mimori K (2013) Microarray analysis reveals that high mobility group A1 is involved in colorectal cancer metastasis. Oncol Rep 30 (3): 1488–1496.
Tolstorukov MY, Sansam CG, Lu P, Koellhoffer EC, Helming KC, Alver BH, Tillman EJ, Evans JA, Wilson BG, Park PJ, Roberts CW (2013) Swi/Snf chromatin remodeling/tumor suppressor complex establishes nucleosome occupancy at target promoters. Proc Natl Acad Sci USA 110 (25): 10165–10170.
Wang KC, Chang HY (2011) Molecular mechanisms of long noncoding RNAs. Mol Cell 43 (6): 904–914.
You L, Chang D, Du HZ, Zhao YP (2011) Genome-wide screen identifies PVT1 as a regulator of Gemcitabine sensitivity in human pancreatic cancer cells. Biochem Biophys Res Commun 407 (1): 1–6.
Acknowledgements
The present study was supported in part by the following grants and foundation; CREST, Japan Science and Technology Agency (JST), the Funding Program for Next Generation World-Leading Researchers (LS094), and Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Scientific Research, grant number 25861199. We thank T Shimooka, K Oda, M Kasagi, S Kono, T Kawano and M Aoyagi for their technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Takahashi, Y., Sawada, G., Kurashige, J. et al. Amplification of PVT-1 is involved in poor prognosis via apoptosis inhibition in colorectal cancers. Br J Cancer 110, 164–171 (2014). https://doi.org/10.1038/bjc.2013.698
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2013.698
Keywords
This article is cited by
-
Upregulation of the c-MYC oncogene and adjacent long noncoding RNAs PVT1 and CCAT1 in esophageal squamous cell carcinoma
BMC Cancer (2023)
-
Identification of LncPVT1 and CircPVT1 as prognostic biomarkers in human colorectal polyps
Scientific Reports (2023)
-
CircPVT1: a pivotal circular node intersecting Long Non-Coding-PVT1 and c-MYC oncogenic signals
Molecular Cancer (2022)
-
lncRNA PVT1: a novel oncogene in multiple cancers
Cellular & Molecular Biology Letters (2022)
-
Unravelling similarities and differences in the role of circular and linear PVT1 in cancer and human disease
British Journal of Cancer (2022)