Abstract
During the hair follicle (HF) cycle, HR protein expression is not concordant with the presence of the Hr mRNA transcript, suggesting an elaborate regulation of Hr gene expression. Here we present evidence that the 5′ untranslated region (UTR) of the Hr gene has internal ribosome entry site (IRES) activity and this activity is regulated by the binding of poly (rC) binding protein 2 (PCBP2) to Hr mRNA. Overexpression and knockdown of PCBP2 resulted in a decrease in Hr 5′ UTR IRES activity and an increase in HR protein expression without changing mRNA levels. We also found that this regulation was disrupted in a mutant Hr 5′ UTR that has a mutation responsible for Marie Unna hereditary hypotrichosis (MUHH) in both mice and humans. These findings suggest that Hr mRNA expression is regulated at the post-transcriptional level via IRES-mediated translation control through interaction with PCPB2, but not in MUHH.
Similar content being viewed by others
Introduction
Hairless (HR) is a transcriptional co-factor that regulates downstream gene expression as a co-repressor of nuclear receptors, including thyroid hormone receptors, retinoic acid receptors and vitamin D receptors.1, 2, 3, 4 The Hr gene is expressed mainly in the brain and skin and has been shown to be associated with the Wnt signaling pathway in hair follicle (HF) development.4, 5, 6, 7 In humans, mutations of the HR gene cause hair loss disorders, including alopecia universalis congenita, atrichia papular lesions and Marie Unna hereditary hypotrichosis (MUHH).8, 9, 10 Additionally, many Hr mutant mice, such as Hrhr, Hrrh, Hrm1Enu and HrHp, have been used as mouse models for human hair loss disorders.11, 12, 13 The numerous mutations of Hr that manifest an abnormal hair phenotype in humans and mice suggest that the function of the HR protein is critically important in HF development. It is interesting to note that both the lack and excess of HR result in the hair loss phenotype. Recently, we reported that overexpression of the HR protein caused hair loss disease in humans (MUHH) and mice (HrHp). More specifically, we demonstrated that the HrHp mice had a regulatory mutation in the Hr 5′ untranslated region (UTR) resulting in overexpression of the HR protein, excessive induction of Wnt signaling,14 and abnormal formation of HF structures.15 This evidence indicated that tight regulation of HR expression levels was important in HF development and normal hair cycling.
In general, translation of mRNAs is initiated with the recognition of the 5′-cap structure of mRNAs by eukaryotic translation initiation factor (eIF) 4F (eIF4F) that consists of three subunits including eIF4E (the direct cap-binding protein), eIF4A and eIF4G (a scaffold for the assembly of eIF4E and eIF4A, which links the mRNAs to ribosomes via eIF3). Then, the 43S preinitiation complex consisting of a 40S ribosomal subunit, eIF2, eIF3 and eIF5 attaches to the capped 5′ proximal region of mRNAs and scans downstream for the initiation codon.16, 17 Alternatively, translation of some mRNAs is regulated by another mechanism that occurs in a cap-independent manner. The 5′ UTR of these mRNAs are usually long, have a high GC content and contain several upstream open reading frames (uORFs), which creating a blockade to ribosomal scanning for initiation of translation.18, 19 Although they have an inhibitory structure, these mRNAs also contain specific sequences in their 5′ UTRs, termed internal ribosome entry sites (IRESs), to which ribosomes bind directly to initiate cap-independent translation.20, 21 To regulate their activities, it requires specific partners for their IRES-mediated translation referred to as IRES-trans acting factors (ITAFs). Several RNA-binding proteins, including Unr, La, several hnRNP family members and others have been identified as ITAFs. These ITAFs bind to IRES-containing mRNAs through recognition of specific sequences or secondary structures in order to regulate IRES activity.20, 22
IRES-mediated translation control is used by specific mRNAs to regulate protein expression under particular conditions in which cap-dependent translation is suppressed, such as apoptosis, cell cycling, development and differentiation.23, 24 The temporal expression of the Hr gene has been shown to be discordant with regard to the presence of its mRNA and protein during the HF cycle. Whereas Hr mRNA expression starts at growth phase (anagen) and continues through regressing phase (catagen), HR protein expression starts mainly at the catagen stage and diminishes at the anagen stage.5, 6, 7 This suggests that HR protein expression is regulated by a mechanism working at the post-transcriptional level. Interestingly, the Hr 5′ UTR sequence is 695 bp long, has a high GCcontent (71.2%), and contains four uORFs. Furthermore, the 5′ UTR of the Hr gene has been shown to inhibit the translation of the main coding sequence,10, 14 which raises a fundamental question regarding exactly how the Hr mRNA transcript overcomes the inhibition of translation by this complex and inhibitory 5′ UTR.
In the present study, we demonstrated that the 5′ UTR of Hr had IRES activity and that Hr mRNA was translated under conditions where the cap-dependent translation was blocked. Additionally, we identified poly(rC) binding protein 2 (PCBP2) as an ITAF for the Hr 5′ UTR IRES activity and showed that PCBP2 negatively regulated HR expression. Furthermore, we found that this regulation was suppressed by the MUHH mutant 5′ UTRs in both the mouse and human HR gene. These results suggested that HR expression is tightly regulated at the translational level via IRES activity during the HF cycle, and the disruption of this regulation might cause MUHH disease.
Materials and methods
Plasmid construction
All PCR amplification was performed using the Expend High Fidelity PCR system (Roche, Indianapolis, IN, USA) using specific primers. The Hr mRNA containing the 5′ UTR and coding sequence (CDS) (Hr 5′ UTR-X) was amplified using RT-PCR from wild-type mouse skin, and then cloned into pcDNA 3.1 (Invitrogen, Carlsbad, CA, USA). The full-length wild-type Hr mRNA (FL-mHr) and T403A mutant mRNA (FL-mHr_T403A) including the 5′ UTR, CDS and 3′ UTR were amplified using RT-PCR from wild-type and HrHP mutant skin respectively, and cloned into the pCMV·SPORT6 vector (Invitrogen). We constructed the bicistronic vector (pRF) with Renilla luciferase (Rluc) as the first cistron and firefly luciferase (Fluc) as the second cistron based on the pGL3 basic vector according to a previous report.25 For construction of the pRF vector, the chimeric intron and Rluc were amplified from the pRL-TK vector and then cloned into pGL3 using a KpnI site, yielding pGL3/Chi.intron-Rluc. The SV40 enhancer element was amplified from the pEGFP-N2 vector and then cloned into pGL3/Chi.intron-Rluc using a BamHI/SalI site, yielding pGL3/Chi.intron-Rluc/SV40-enh. The SV40 promoter was amplified from pSV-beta-Galactosidase and then cloned into pGL3/Chi.intron-Rluc/SV40-enh using a PstI site, yielding pGL3/Chi.intron-Rluc/SV40-enh/SV40-pro pRF (Figure 1c). The mouse and human HR 5′ UTR were amplified as previously reported14 for pRF_m695 and pRF_h690, respectively. The IRES sequence fragment of encephalomyocarditis virus (EMCV) was amplified from the pIRES-EGFP vector as a positive control measurement of IRES activity for pRF_EMCV. We also generated constructs containing reverse mouse and human HR 5′ UTR, which had the 5′ UTRs inserted in the reverse orientation into the pRF vector as a control for sequence specificity of IRES (pRF_m695rev and pRF_h690rev, respectively). The pRF_mT403A and pRF_hT-320C mutant constructs were generated using a Site-directed Mutagenesis Kit (Agilent Technologies, Santa Clara, CA, USA) according to the manufacturer’s instructions. For construction of serial deletion constructs, we amplified DNA fragments by PCR using specific primer pairs designed for each deletion. We designed a 5′ proximal nt deletion for pRF_m84-695, a 3′ proximal nt deletion construct for pRF_m1-506, both proximal end deletion constructs for pRF_m84-506, a 3′ nt deletion construct for pRF_m1-120, and a 5′ nt deletion construct for pRF_m402-695. All PCR amplified fragments for analysis of IRES activity were inserted between Rluc and Fluc coding sequences using the BglII site of the pRF vector. The coding sequence of human PCBP2 was amplified using cDNA extracted from HEK293T cells by PCR and the sequence-verified CDS was then cloned into the pcDNA-FLAG vector (Invitrogen). For bacterial expression, the CDS of PCBP2 was cloned into the pGEX4T-1 vector and purified according to the manufacturer’s instructions (GE Healthcare, Pittsburgh, PA, USA). All primers used for plasmid construction and for mutagenesis are available upon request. All mice were maintained as described previously.11 All animal experiments were approved by the Institutional Animal Care and Use Committee (IACUC) of the Catholic University of Korea. All experiments were carried out in accordance with the Guidelines for Animal Experimentation. All experiments using human sample were approved by the Institutional Review Board and Ethics Committee of the Catholic University of Korea.
The 5′ UTR of Hr has IRES activity. (a) Schematic diagrams of the Hr-5′ UTR-X constructs which includes the 5′ UTR and CDS of Hr mRNA. Western blotting was performed using lysates of HEK293T cells transfected with the Hr-5′ UTR-X construct. (b) Rapamycin (Rapa)- or cycloheximide (CHX)-treated HEK293T cells were harvested at indicated time points, followed by the determination of HR protein expression levels by western blotting. Dimethyl sulfoxidewas used as a vehicle control treatment. (c) Schematic diagrams of the pRF bicistronic vectors containing the 5′ UTRs from mouse Hr or human HR with forward (pRF_m695 and pRF_h690) or reverse (pRF_m695rev and pRF_h690rev) orientations, and EMCV IRES elements (pRF_EMCV). The reporter constructs were transfected into HEK293T cells, and the IRES activities were measured and normalized by β-galactosidase activity 48 h after transfection. (d) PAM212 mouse keratinocytes were transfected with pRF_m695, and IRES activities were measured after 48 h. (e) Schematic diagrams of the pRF reporter vectors that contain several deletion fragments of the Hr 5′ UTR. The plasmid pRF_m695 includes the full-length Hr 5′ UTR, and the numbers associated with each of the constructs represent the nucleotide positions of the Hr 5′ UTR sequences. IRES activities were measured 48 h after transfection. All luciferase experiments were performed three times in triplicate, and transfection efficiencies were normalized against β-galactosidase activity. Activities are expressed as the mean±s.e.m. **P<0.01, ***P<0.001, ****P<0.0001.
Cell culture, RNA interference and the luciferase reporter assay
The HEK293T and PAM212 cell lines were maintained in Dulbecco’s modified Eagle’s medium, supplemented with 10% fetal bovine serum at 37 °C with 5% CO2. HEK293T cells stably expressing HR protein were treated with 20 nM of rapamycin or 100 μg/ml of cycloheximide (CHX) and harvested at the indicated times. DMSO was used as a vehicle control. For generation of HR stable cell lines, HEK293T cells were transfected with the pcDNA/Hr-5′ UTR-CDS construct and resistant colonies were selected using G418 (200 μg ml−1, Invitrogen). All transfection experiments were performed using Lipofectamine 2000 (Invitrogen) according to manufacturer’s instructions. Luciferase assays were performed using the Dual-Luciferase Assay Kit (Promega, Madison, WI, USA) at 48 h after transfection. The IRES activities were calculated as the ratio between Flucand Rlucluciferase activity (Fluc/Rluc) normalized for transfection efficiency by β-galactosidase activity. The small interfering RNAs (siRNAs) were designed for the endogenous human poly (rC) binding protein 2 (siPCBP2) knockdown, and we synthesized 2 siRNAs for PCBP2 (siPCBP2) as previously reported.26 The siPCBP2s (20 nM) were introduced to cells at 24 h after transfection of the reporter construct or the Hr full-length construct, and then the IRES activities were measured or western blotting was performed after further incubation for 24 h.
RNA isolation and quantitative RT-PCR
Total RNA was extracted from cells using TRIZOL (Invitrogen) following the manufacturer’s instructions. Single-stranded cDNAs were synthesized by reverse transcription reactions using the SuperScript First-Strand Synthesis System (Invitrogen). Quantitative RT-PCR was carried out in a reaction mixture containing SYBR Premix Ex Taq (Clontech, Mountain View, CA, USA) using a CFX96PCR machine (Bio-Rad, Hercules, CA, USA). The relative levels of mRNA expression were determined by the comparative ΔΔCt method.27 Sequences of gene specific primers for qRT-PCR were as follows: mHr_F, 5′-gagaagagtggggtgtgagc-3′; mHr_R, 5′-ctcggttacctacccaccac-3′; GAPDH_F, 5′-aactttggcattgtggaagg-3′; and GAPDH_R, 5′-acacattgggggtaggaaca-3′.
RNA affinity chromatography, RNA pull-down and UV cross-linking assays
HEK293T cell lysates were used for RNA affinity chromatography using modified protocols.28 For RNA affinity chromatography and the RNA pull-down assay, the Hr 5′ UTR was labeled with biotin (Roche) by in vitro transcription using T7 or SP6 RNA polymerase (Clontech). To identify the Hr 5′ UTR binding proteins, we performed sodium dodecyl sulfate PAGE (SDS-PAGE) using eluted protein from RNA affinity chromatography. The proteins were then stained with Coomassie Blue G-250 (Sigma, St. Louis, MO, USA). The sequences of the protein bands were analyzed using LC/MS/MS analysis (Korea Basic Science Institute, Seoul, Korea). For RNA pull-down experiments, we also used biotin-labeled wild-type or mutant Hr 5′ UTR probes with HEK293T cell lysates prepared from the cells transfected with FLAG-PCBP2 or purified GST-PCBP2 protein using a modified protocol.29, 30 After labeling, the Hr 5′ UTR probe (3 μg) was incubated with cell lysates (30 μg) or purified GST-PCBP2 protein (100 ng) in incubation buffer (10 mM Tris-Cl, pH 7.4, 150 mM KCl, 1.5 mM MgCl2, 0.5 mM dithiothreitol (DTT), 0.5 mM phenylmethylsulfonyl fluoride, 0.05% (v/v) Nonidet P-40, 120) with 120 μg of yeast tRNA (Sigma) and 10 U of RNase Inhibitor (Clontech) at 4 °C overnight with continuous rotation. Then, the RNA–protein complex was incubated with Streptavidin-Agarose beads (Thermo Scientific, Waltham, MA, USA) for 1 h at 4 ºC, and beads were washed 4 × 5 min using incubation buffer. After washing, proteins were eluted using 1 × sample buffer and were separated by 10% SDS-PAGE. Western blotting analysis was performed as described above. The UV cross-linking analysis was performed as described previously.28 The [32P]-labeled Hr 5′ UTR probe and unlabeled Hr 5′ UTR probe for the competition assay were incubated with GST or purified GST-PCBP2 protein in reaction buffer (0.5 mM DTT, 5 mM HEPES (pH 7.6), 75 mM KCl, 2 mM MgCl2, 0.1 mM EDTA, 4% glycerol, 20 U of RNasin, 3 μg of yeast tRNA) at 30 °C for 20 min. RNA–protein complexes were irradiated on ice with 254-nm UV light at 400 000 μJ cm−2 for 15 min. After irradiation, un-bound RNAs were digested using RNase cocktail (Thermo Scientific) at 37 °C for 20 min. The RNA–protein complexes were separated by 10% SDS-PAGE and visualized by autoradiography.
Western blotting and immunostaining
Western blotting and immunostaining analyses were performed as previously described.14 Proteins were extracted using RIPA buffer (150 mM NaCl, 1% NP-40, 0.5% NaN3, 0.1% SDS, 50 mM Tris-HCl, pH 8.0). Protein samples were eluted in 1 × sample buffer and separated using 8 or 10% SDS-PAGE. The signal was visualized using an enhanced chemiluminescence detection kit (GE Healthcare). For western blotting, the following antibodies were used anti-HR rabbit polyclonal,14 anti-PCBP2 (Aviva, San Diego, CA, USA), anti-FLAG (Abcam, Cambridge, MA, USA), anti-phospho-S6 ribosomal protein (Cell Signaling, Danvers, MA, USA), anti-GFP, anti-GAPDH, and anti-actin (Santa Cruz, Dallas, TX, USA). The relative expression levels of proteins were determined based on densitometry analysis. For histological analysis, we used mouse skin of the BALB/c strain, and mice were maintained as described previously.14 The mouse dorsal skin samples were prepared at postnatal day 7 (P7), P14, P17, P21, P25 and P28. The mouse skin samples were embedded in a paraffin block. The hematoxylin and eosin staining and immunostaining were performed as described previously.11, 14 For immunostaining, the sections were incubated overnight at 4 °C with anti-PCBP2 antibody (1:200, Aviva). After washing, the sections were incubated for 1 h with Alexa Flour 594 goat anti-rabbit IgG antibody (Invitrogen) and counterstained with Hoechst dye. The images were taken on an Olympus AX-70 microscope.
Results
The 5′ UTR of the Hr gene inhibits the expression of the HR protein and has IRES activity
Using a heterologous reporter gene, the effect of the uORFs in the 5′ UTR of the Hr gene on the expression of the main ORF was explored and uncovered an inhibitory effect of the second uORF on the translation of the main ORF of the GFP reportergene.10 Because the effect of the uORFs on the synthesis of the HR protein has not been studied, we investigated the function of the Hr 5′ UTR on HR protein expression using several mouse Hr mutant constructs. Consistent with the results from previous reports,10, 14 removal of uORF 1 and uORF 2 showed decreased and increased HR expression, respectively. Interestingly, disruption of uORF 3 or uORF 4 resulted in increased HR protein expression in contrast to a previous report,10 in which the heterologous luciferase reporter gene was used (Supplementary Figure 1a and b), indicating that the uORFs 2, 3 and 4 of the Hr gene repress its protein expression. These results suggested that HR expression was suppressed by its 5′ UTR cis-element.
To elucidate a mechanism by which Hr mRNA overcomes the repression of its activity via its 5′ UTR, we hypothesized that the Hr 5′ UTR had IRES activity that allowed for the translation of HR to be initiated directly and in a cap-independent manner without inhibition by uORFs. To test this hypothesis, we investigated whether HR protein translation could be initiated when cap-dependent translation was blocked. First, we generated a stable cell line expressing the Hr mRNA containing 5′ UTR and coding sequence (CDS) of Hr (Figure 1a). Regulation of the translational initiation of Hr was explored using this stable cell line and differential inhibitors of translation. We treated these cells with rapamycin, an inhibitor of the mTOR serine/threonine protein kinase, which causes inhibition of cap-dependent translation initiation.31 Additionally, we treated cells with the general translation elongation inhibitor cycloheximide. Compared to the vehicle control, HR protein expression was drastically reduced by treatment with cycloheximide. On the other hand, HR protein expression was maintained at normal levels in the presence of rapamycin, while that of the phosphorylated S6 ribosomal protein (p-RPS6), a positive control for rapamycin activity, was clearly affected (Figure 1b). This result suggested that HR protein synthesis could be initiated in a cap-independent manner, as HR protein expression continued under a cap-dependent translation blocking condition. To examine the IRES activity of the Hr 5′ UTR, we generated bi-cistronic reporter constructs with 5′ UTR sequence fragments from the Renilla luciferase (Rluc) and firefly luciferase (Fluc) genes as previously reported.25 The IRES activities of the 5′ UTRs of the mouse (pRF_m695) and human (pRF_h690) Hr genes were 8.52 (±0.50)- and 5.55 (±0.28)-fold higher than that of the control pRF empty vector, respectively, which is comparable to that of the positive control, pRF_EMCV (6.85±0.35-fold). Reverse forms of both 5′ UTRs did not display any recognizable activities (m695_rev, 1.29±0.003-fold; h690_rev, 1.21±0.25-fold), indicating that the IRES activity of the Hr 5′ UTR was sequence-specific (Figure 1c). To rule out the possibility that the Hr 5′ UTR could contain a cryptic promoter activity or splicing site, we checked expression levels of each reporter gene by quantitative real-time PCR. There was no significant difference of the Fluc relative to Rluc expression between pRF and pRF_m695 (Supplementary Figure 2). To test cell-type specificity, we assessed the IRES activity of the pRF_m695 construct in PAM212 mouse keratinocytes, which normally express Hr mRNA. The Fluc activity was increased in the PAM212 cells transfected with the pRF_m695 construct (10.54±0.85-fold) (Figure 1d) and was also increased in the glioblastoma cell line U251 (Supplementary Figure 3), suggesting that the IRES activity of the 5′ UTR of the Hr gene was not cell type-dependent. To determine the cis-acting region of the Hr 5′ UTR that conferred the IRES activity, we generated several deletion constructs of pRF_m695 and performed a luciferase assay. Interestingly, all deletion constructs showed decreased IRES activities compared to that of the full-length 5′ UTR of Hr (Figure 1e). Thus, we did not detect any specific cis-acting region responsible for the IRES activity in the Hr 5′ UTR. These data suggested that the 5′ UTR of the Hr gene had IRES activity and that its activity was not cell type-specific.
PCBP2 binds to the Hr 5′ UTR, and PCBP2 expression is inversely correlated with HR expression during the HF cycle
To identify the partner ITAF involved in the IRES function of the Hr 5′ UTR, we performed RNA affinity chromatography using a biotin-labeled Hr 5′ UTR probe (m695-Biotin) (Supplementary Figure 4). The bands specifically bound by the probe were further analyzed using liquid chromatography-tandem mass spectrometry (LC/MS/MS) analysis (Supplementary Table 1). Based on the results of peptide sequencing (Supplementary Figure 4b and c), we focused on a protein of approximately 40 kDa (Supplementary Figure 4a, arrow) identified as PCBP2.
To confirm the binding of PCBP2 to the Hr 5′ UTR, we carried out an RNA pull-down assay using a m695-biotin probe. Western blot analysis of pulled-down proteins from the FLAG-PCBP2 overexpressing cell lysates or the purified GST-PCBP2 protein detected binding of PCBP2 to the Hr 5′ UTR (Figure 2a and b). The specificity of this binding was further confirmed by competition assays using ultraviolet (UV) cross-linking experiment. The radioactive signal was increased with increasing amounts of PCBP2 protein, and the signal was significantly decreased by the addition of an unlabeled probe (Figure 2c). These results indicated that PCBP2 bound the Hr mRNA 5′ UTR specifically and raised the possibility that PCBP2 was an ITAF associated with the IRES function of the Hr 5′ UTR.
Poly(rC) binding protein 2 interactions with the 5′ UTR of Hr mRNA and expression patterns during the HF cycle. (a, b) RNA pull-down experiments were performed using FLAG-PCBP2 overexpressing HEK293T cell lysates or purified GST-PCBP2 protein with or without a biotinylated Hr 5′ UTR probe (Biotin-m695). PCBP2 protein was detected by western blotting. (c) A competition analysis was performed in a UV cross-linking experiment. A radiolabeled Hr 5′ UTR RNA probe was incubated with different amounts of purified PCBP2 protein (+; 100 ng/rxn, ++; 200 ng/rxn, +++; 300 ng/rxn). The signal shows a concentration-dependent increase of the interaction, which was decreased specifically by the addition of unlabeled Hr 5′ UTR RNA. GST protein was used as a negative control, and signals were visualized by autoradiography. (d) Mouse skin sections from each time point were subjected to hematoxylin and eosin staining or immunostaining using the anti-PCBP2 antibody. Hoechst dye was used for counterstaining. Scale bar, 200 μm. (e, f) Western blotting was performed using mouse skin tissue from each time point. The quantitative densitometry of PCBP2 expression was presented as the mean±s.e.m. of three mice per time point.
As previously mentioned, the HR protein is expressed mainly during the catagen and telogen phases of HF cycling. We tested whether there was a correlation between the expression of HR and PCBP2 during the HF cycle. The PCBP2 protein was expressed mainly in the matrix and outer root sheath of the HF (Figure 2d). Interestingly, the PCBP2 protein was highly expressed during the anagen phase, such as at P7, P25 and P28. Conversely, PCBP2 expression was decreased at early-(P14), as well as mid-(P17) catagen. Additionally, it was difficult to detect the PCBP2 protein during the telogen (P21) phase. We confirmed this expression pattern of PCBP2 protein in HF using western blotting (Figure 2e and f). These results suggested that there was an inverse relationship between the expression patterns of HR and PCBP2 during HF cycling.
PCBP2 acts as a negative regulator of the IRES activity of the Hr 5′ UTR and IRES-mediated translation of the HR protein
We next investigated whether PCBP2 could affect the IRES activity of the Hr 5′ UTR. Impressively, the IRES activity of the Hr 5′ UTR (pRF_m695) was markedly decreased by PCPB2 overexpression (Figure 3a). Furthermore, this negative regulation of PCBP2 on the IRES activity of the Hr 5′ UTR showed a dose–dependent response (Figure 3b). As expected, the IRES activity of the Hr 5′ UTR was decreased by PCBP2 overexpression in PAM212 keratinocytes (Figure 3c). The negative regulation of PCBP2 on the IRES activity of the 5′ UTR of the human HR (pRF_h690) was similar to that on the 5′ UTR of the mouse Hr (Figure 3d). Next, we tested whether this negative effect of PCBP2 on the Hr 5′ UTR IRES activity could be circumvented by the knockdown of PCBP2 expression. Decreased expression of PCBP2 was achieved by using specific siRNAs against PCBP2 (siPCBP2) (Figure 3e). As expected, knockdown of PCBP2 resulted in the increased IRES activity of the Hr 5′ UTR (Figure 3f).
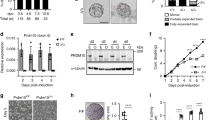
PCBP2 negatively regulates the IRES activity of the Hr 5′ UTR and HR protein expression. (a) The IRES activities were measured in HEK293T cells co-transfected with pRF_m695 and pFLAG-PCBP2 at 48 h after transfection. The expression of the FLAG-PCBP2 protein was confirmed by western blotting. (b) The IRES activities of pRF_m695 were measured at 48 h after transfection with various concentrations of FLAG-PCBP2. (c) PAM212 mouse keratinocytes were transfected with pRF_m695 and FLAG-PCBP2, and then IRES activities were measured. (d) The IRES activity of human HR 5′ UTR (pRF_h690) was measured in HEK293T cells co-transfected with the pFLAG-PCBP2 expression construct. (e) SiPCBP2 or a scrambled control were transfected into HEK293T cells. At 48 h post transfection, protein extracts were subjected to western blot analysis and expression level was determined using quantitative densitometry of the western blots from three independent experiments. (f) HEK293T cells were transfected with siPCBP2s or a scrambled control and incubated for 24 h. Then, the pRF_m695 reporter construct was transfected and further incubated. Twenty-four hours later, the IRES activities were measured, and cell lysates were subjected to western blotting. (g) HEK293T cells were co-transfected with the pCMV-FL-mHr and pFLAG-PCBP2 constructs. Forty-eight hours later, cells were harvested and HR expression determined by western blot analysis. Actin and GAPDH were used as loading controls, and GFP was used for normalization of the transfection efficiency. Additionally, Hr mRNA expression levels were determined by quantitative real time PCR, and the relative expression level was determined against Gapdh mRNA expression. (h) Increasing amounts of the pFLAG–PCBP2 construct were transfected into the HR stable cell line. Cell lysates were subjected to western blot analysis at 48 h post transfection. RT-PCR revealed no change in the Hr mRNA expression level. (i) HR-expressing stable cells were transfected with siPCBP2s, and cell lysates were subjected to western blotting after 48 h of incubation. All experiments for IRES activity were performed three times in triplicate, and transfection efficiencies were normalized by β-galactosidase activity. Activities are expressed as the mean±SEM. *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001.
The negative effect of PCBP2 on the IRES activity of the Hr 5′ UTR was confirmed by analyzing HR protein expression. The overexpression of PCBP2 resulted in a drastic reduction in the HR protein level, while the expression of Hr mRNA was not reduced in either transiently transfected cells (Figure 3g) or the HR stable cell line (Figure 3h). We further confirmed that PCBP2 overexpression did not affect the level of expression of the endogenous Hr mRNA, as shown by quantitative real-time PCR using PAM212 keratinocytes (Supplementary Figure 5). Similar to the results of the reporter system experiments, HR protein expression was increased by treatment with siPCBP2 compared to that of the scrambled control (Figure 3i). These results strongly suggested that PCBP2 suppressed HR protein expression without changing Hr mRNA expression by negatively regulating the IRES activity of the Hr 5′ UTR as an ITAF.
Mutation of Hr 5′ UTR suppressed the regulation of Hr 5′ UTR by PCBP2 in MUHH
Next, we investigated whether this IRES-mediated translational control for HR protein could affect mutant Hr 5′ UTR, which has the mutation corresponding to MUHH. First, we assessed the expression level of PCBP2 mRNA using skin tissue of wild-type and HrHp mutant mice at the anagen phase which showed highly expressed PCBP2 as seen in (Figure 2d–f). The PCBP2 mRNA expression levels were similar between the wild-type and HrHp mutant skin (Figure 4a). Additionally, we compared IRES activities between wild-type (pRF_m695) and mutant form of mouse Hr 5′ UTR (pRF_mT403A) that has a T403A substitution mutation as previously reported.14 The IRES activity of mutant Hr 5′ UTR was similar to, even slightly higher than that of the wild-type 5′ UTR (Figure 4b). We next investigated whether PCBP2 could affect the IRES activity of the mutant Hr 5′ UTR. Surprisingly, the IRES activity of the mutant form, pRF_mT403A, was not affected by PCBP2 overexpression (Figure 4c). We also confirmed this effect on the HR protein expression itself using a full-length mutant Hr construct that had a T403A mutation in the 5′ UTR (FL-mHr_T403A). As expected, the overexpression of PCBP2 showed no effect on the HR protein level of FL-mHr_T403A (Figure 4d). We next investigated whether PCBP2 binds to the mutant 5′ UTR (mT403A) using an RNA pull-down assay as above. We found that the PCBP2 weakly bound to biotin-labeled mT403A, while it bound strongly to wild type (Biotin-m695) (Figure 4e). We confirmed this binding was dose-dependent in both wild-type and mutant forms (Figure 4f). Furthermore, we performed an IRES activity assay using the human HR 5′ UTR mutant that has the mutation T-320C of MUHH patients (pRF_hT-320C) as reported previously.14 The IRES activity of the MUHH mutant form was also not affected by PCBP2 overexpression (Figure 4g). These results strongly suggested that binding of PCBP2 to the Hr 5′ UTR was suppressed in mutant forms that had mutations corresponding to MUHH in both mice and humans. We suggest that this suppression of PCBP2 binding efficiency may be the cause of maintained IRES activities in Hr 5′ UTR mutants.
MUHH mutant forms of the 5′ UTR were not affected by PCBP2 negative regulation in both mice and humans. (a) PCBP2 mRNA expression levels were determined by quantitative real time PCR using wild-type and HrHp mutant P28 mouse skin tissues, and the relative expression level was determined against Gapdh mRNA expression. (b) The IRES activities of pRF_m695 (wild type) and pRF_mT403A (HrHp mutant type) were measured 48 h after transfection. (c) The pRF_mT403A and pFLAG-PCBP2 constructs were co-transfected into HEK293T cells. The IRES activities were measured 48 h after transfection. The expression of the FLAG-PCBP2 protein was confirmed by western blotting. (d) HEK293T cells were co-transfected with the pCMV-FL-mHr_T403A and the pFLAG-PCBP2 constructs. Forty-eight hours later, cells were harvested and HR expression determined by western blot analysis. GAPDH was used as a loading control, and GFP was used for normalization of the transfection efficiency. (e, f) RNA pull-down experiments were performed using FLAG-PCBP2 overexpressing HEK293T cell lysates with a wild-type (Biotin-m695) or mutant (Biotin-mT403A) biotinylated Hr 5′ UTR probe. The PCBP2 protein was detected by western blotting. (g) The pRF_hT-320C and pFLAG-PCBP2 constructs were co-transfected into HEK293T cells. The IRES activities were measured 48 h after transfection. The expression of the FLAG-PCBP2 protein was confirmed by western blotting. All experiments for IRES activity were performed three times in triplicate, and transfection efficiencies were normalized by β-galactosidase activity. Activities are expressed as the mean±s.e.m. *P<0.05.
Discussion
The sequences and structures of the Hr 5′ UTR suggested that it may inhibit efficient translation initiation by the conventional cap-dependent ribosome scanning mechanism. We recently reported that the Hr 5′ UTR mutation T403A resulted in the MUHH phenotype and overexpression of the HR protein.14 Here we showed that the uORFs of Hr gene (except for uORF 1) function as inhibitory translational control elements. These features suggested that the Hr 5′ UTR could be a functional cis-element for the expression of the HR protein. Although the results of one study indicated that the 5′ UTR of the Drosophila Hr had an IRES function, its sequence and structure were entirely different from those of the mammalian Hr genes.32 As we reported previously, the similarity of the 5′ UTR of mammalian Hr genes, such as human, mouse, and rat, is more than 80%,14 and there have been no reports pertaining to the function of the mammalian Hr 5′ UTR. In the present study, we demonstrated that the Hr 5′ UTR has IRES activity and that this activity seemed to be dependent on the secondary structure rather than specific sequences. Additionally, we confirmed that this IRES activity was functional in several cell lines including keratinocytes, which is a cell type known to express the HR protein.
PCBP2 is an RNA-binding protein that functions as an ITAF for the IRESs of several viruses, such as hepatitis C, hepatitis A and picornavirus, as well as of Myc cellular mRNA.33, 34, 35, 36 In this report, we showed that the PCBP2 protein bound to the Hr 5′ UTR and modulated its IRES activity. Furthermore, we found that the PCBP2 protein suppressed the expression of the HR protein and Hr 5′ UTR IRES activity without changing its mRNA expression. We suggest that the Hr mRNA is a novel binding target of the PCBP2 protein through its 5′ UTR and that PCBP2 acts as a negative regulator of Hr gene translation via post-transcriptional modulation as an ITAF.
As previously mentioned, Hr mRNA is expressed during all phases of the HF cycle. However, the HR protein is mainly expressed during the late catagen through telogen phases. In the anagen stage, which is the growth phase of the HF cycle, the HR protein is weakly expressed, although significant amounts of the mRNA are expressed.5, 6 The catagen stage is a regressing phase in which cells consisting of the HF undergo cell death, and the telogen stage is a resting phase in which cell metabolism is quiescent. During cell death, protein expression is blocked by decreased cap-dependent translation initiation. However, mRNAs can be translated into proteins by cap-independent mechanisms, such as IRES-mediated translational initiation.23 Based on our results, we suggest that HR protein expression may be maintained during the cell death and resting phases of the HF cycle by an IRES mechanism mediated via its 5′ UTR, even though the overall cap-dependent initiation of translation is blocked. Furthermore, we suggest that this IRES-mediated translational control of the HR protein is modulated by PCBP2. There have been no reports regarding the PCBP2 expression pattern in the skin or HF. Here we showed the expression pattern of the PCBP2 protein during the HF cycle and found that the PCBP2 protein level differed significantly depending on the phases of the HF cycle. Specifically, PCBP2 was highly expressed during the growth phase, which is an active stage of the cell cycle or proliferation. This result may suggest a mechanism for the discrepant expression of the Hr mRNA and the HR protein during the HF cycle. HR protein expression may be decreased during the anagen phase by the highly expressed PCBP2 protein. Conversely, increased HR protein expression during the catagen and telogen phases may be attributable to decreased expression of the PCBP2 negative regulator protein, allowing for the activation of IRES-mediated translation.
Interestingly, we showed that the regulation of Hr 5′ UTR IRES activity and HR protein expression by PCBP2 was suppressed in the MUHH mutant 5′ UTRs in both mouse and human cells. Additionally, we presented that the binding of PCBP2 to the mutant Hr 5′ UTR was decreased compared to wild type. We compared secondary structures of wild-type and T403A mutant Hr 5′ UTRs predicted by the Vienna RNA folding server.37 Interestingly, the WT and the mutant Hr 5′ UTR had very different secondary structures (Supplementary Figure 6), which could explain their different binding efficiencies with the PCBP2 protein. However, further study should be conducted to understand how these different secondary structures of Hr 5′ UTRs affect their interaction with PCBP2. These results suggest that divergent HR protein and mRNA expression is regulated by an IRES mechanism of its 5′ UTR cis-element through PCBP2 during HF cycling in wild type, but this regulation is disrupted in MUHH. Thus, we suggest that the failure of this fine translational control may cause abnormal overexpression of the HR protein during HF cycling in MUHH (Figure 5).
Model for the regulation of HR expression via IRES-mediated translational control by PCBP2 in wild type and MUHH during HF cycling. In wild type, PCBP2 inhibits the IRES activity of Hr 5′ UTR in the anagen phase, then this inhibition is lifted at the catagen phase as PCBP2 expression decreases. However, in the MUHH mutant, the IRES activity of the Hr 5′ UTR is not inhibited by PCBP2 during the anagen phase, even though the PCBP2 expression is high because of the weak interaction between PCBP2 and Hr mRNA. Consequently, it results in abnormal overexpression of the HR protein during the whole HF cycle.
Here we suggest a new mechanism that regulates HR protein expression and MUHH, a hair loss disease caused by changes in HR protein expression. Our results may be helpful in the further elucidation of the mechanisms of hair loss disease.
References
Hsieh JC, Sisk JM, Jurutka PW, Haussler CA, Slater SA, Haussler MR et al. Physical and functional interaction between the vitamin D receptor and hairless corepressor, two proteins required for hair cycling. J Biol Chem 2003; 278: 38665–38674.
Moraitis AN, Giguere V, Thompson CC . Novel mechanism of nuclear receptor corepressor interaction dictated by activation function 2 helix determinants. Mol Cell Biol 2002; 22: 6831–6841.
Potter GB, Beaudoin GM 3rd, DeRenzo CL, Zarach JM, Chen SH, Thompson CC . The hairless gene mutated in congenital hair loss disorders encodes a novel nuclear receptor corepressor. Genes Dev 2001; 15: 2687–2701.
Thompson CC . Thyroid hormone-responsive genes in developing cerebellum include a novel synaptotagmin and a hairless homolog. J Neurosci 1996; 16: 7832–7840.
Beaudoin GM 3rd, Sisk JM, Coulombe PA, Thompson CC . Hairless triggers reactivation of hair growth by promoting Wnt signaling. Proc Natl Acad Sci USA 2005; 102: 14653–14658.
Panteleyev AA, Paus R, Christiano AM . Patterns of hairless (hr) gene expression in mouse hair follicle morphogenesis and cycling. Am J Pathol 2000; 157: 1071–1079.
Thompson CC, Sisk JM, Beaudoin GM 3rd . Hairless and Wnt signaling: allies in epithelial stem cell differentiation. Cell Cycle 2006; 5: 1913–1917.
Ahmad W, Faiyaz ul Haque M, Brancolini V, Tsou HC, ul Haque S, Lam H et al. Alopecia universalis associated with a mutation in the human hairless gene. Science 1998; 279: 720–724.
Ahmad W, Zlotogorski A, Panteleyev AA, Lam H, Ahmad M, Faiyaz ul Haque M et al. Genomic organization of the human hairless gene (HR) and identification of a mutation underlying congenital atrichia in an Arab Palestinian family. Genomics 1999; 56: 141–148.
Wen Y, Liu Y, Xu Y, Zhao Y, Hua R, Wang K et al. Loss-of-function mutations of an inhibitory upstream ORF in the human hairless transcript cause Marie Unna hereditary hypotrichosis. Nat Genet 2009; 41: 228–233.
Baek IC, Kim JK, Cho KH, Cha DS, Cho JW, Park JK et al. A novel mutation in Hr causes abnormal hair follicle morphogenesis in hairpoor mouse, an animal model for Marie Unna Hereditary Hypotrichosis. Mamm Genome 2009; 20: 350–358.
Nam Y, Kim JK, Cha DS, Cho JW, Cho KH, Yoon S et al. A novel missense mutation in the mouse hairless gene causes irreversible hair loss: genetic and molecular analyses of Hr m1Enu. Genomics 2006; 87: 520–526.
Sundberg JP . Handbook of Mouse Mutations with Skin and Hair Abnormalities Animal Models and Biomedical Tools. CRC Press: Boca Raton, FL, USA, 1994.
Kim JK, Kim E, Baek IC, Kim BK, Cho AR, Kim TY et al. Overexpression of Hr links excessive induction of Wnt signaling to Marie Unna hereditary hypotrichosis. Hum Mol Genet 2010; 19: 445–453.
Kim BK, Lee HY, Choi JH, Kim JK, Yoon JB, Yoon SK . Hairless plays a role in formation of inner root sheath via regulation of Dlx3 gene. J Biol Chem 2012; 287: 16681–16688.
Gingras AC, Raught B, Sonenberg N . eIF4 initiation factors: effectors of mRNA recruitment to ribosomes and regulators of translation. Annu Rev Biochem 1999; 68: 913–963.
Sonenberg N, Hinnebusch AG . Regulation of translation initiation in eukaryotes: mechanisms and biological targets. Cell 2009; 136: 731–745.
Morris DR, Geballe AP . Upstream open reading frames as regulators of mRNA translation. Mol Cell Biol 2000; 20: 8635–8642.
Sachs MS, Geballe AP . Downstream control of upstream open reading frames. Genes Dev 2006; 20: 915–921.
Gebauer F, Hentze MW . Molecular mechanisms of translational control. Nat Rev Mol Cell Biol 2004; 5: 827–835.
Hellen CU, Sarnow P . Internal ribosome entry sites in eukaryotic mRNA molecules. Genes Dev 2001; 15: 1593–1612.
Stoneley M, Willis AE . Cellular internal ribosome entry segments: structures, trans-acting factors and regulation of gene expression. Oncogene 2004; 23: 3200–3207.
Holcik M, Sonenberg N . Translational control in stress and apoptosis. Nat Rev Mol Cell Biol 2005; 6: 318–327.
Spriggs KA, Bushell M, Mitchell SA, Willis AE . Internal ribosome entry segment-mediated translation during apoptosis: the role of IRES-trans-acting factors. Cell Death Differ 2005; 12: 585–591.
Stoneley M, Paulin FE, Le Quesne JP, Chappell SA, Willis AE . C-Myc 5' untranslated region contains an internal ribosome entry segment. Oncogene 1998; 16: 423–428.
Han W, Xin Z, Zhao Z, Bao W, Lin X, Yin B et al. RNA-binding protein PCBP2 modulates glioma growth by regulating FHL3. J Clin Invest 2013; 123: 2103–2118.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001; 25: 402–408.
Holcik M, Korneluk RG . Functional characterization of the X-linked inhibitor of apoptosis (XIAP) internal ribosome entry site element: role of La autoantigen in XIAP translation. Mol Cell Biol 2000; 20: 4648–4657.
Kim JH, Paek KY, Choi K, Kim TD, Hahm B, Kim KT et al. Heterogeneous nuclear ribonucleoprotein C modulates translation of c-myc mRNA in a cell cycle phase-dependent manner. Mol Cell Biol 2003; 23: 708–720.
Lewis SM, Veyrier A, Hosszu Ungureanu N, Bonnal S, Vagner S, Holcik M . Subcellular relocalization of a trans-acting factor regulates XIAP IRES-dependent translation. Mol Biol Cell 2007; 18: 1302–1311.
Shah OJ, Anthony JC, Kimball SR, Jefferson LS . 4E-BP1 and S6K1: translational integration sites for nutritional and hormonal information in muscle. Am J Physiol Endocrinol Metab 2000; 279: E715–E729.
Maier D, Nagel AC, Preiss A . Two isoforms of the Notch antagonist hairless are produced by differential translation initiation. Proc Natl Acad Sci USA 2002; 99: 15480–15485.
Ali N, Siddiqui A . Interaction of polypyrimidine tract-binding protein with the 5' noncoding region of the hepatitis C virus RNA genome and its functional requirement in internal initiation of translation. J Virol 1995; 69: 6367–6375.
Evans JR, Mitchell SA, Spriggs KA, Ostrowski J, Bomsztyk K, Ostarek D et al. Members of the poly(rC) binding protein family stimulate the activity of the c-myc internal ribosome entry segment in vitro and in vivo. Oncogene 2003; 22: 8012–8020.
Graff J, Cha J, Blyn LB, Ehrenfeld E . Interaction of poly(rC) binding protein 2 with the 5' noncoding region of hepatitis A virus RNA and its effects on translation. J Virol 1998; 72: 9668–9675.
Walter BL, Nguyen JH, Ehrenfeld E, Semler BL . Differential utilization of poly(rC) binding protein 2 in translation directed by picornavirus IRES elements. Rna 1999; 5: 1570–1585.
Gruber AR, Lorenz R, Bernhart SH, Neubock R, Hofacker IL . The Vienna RNA websuite. Nucleic Acids Res 2008; 36: W70–W74.
Acknowledgements
This work was supported by Basic Science Research Programs through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2011-355-C00085 and 2012R1A2A2A01047225) and the Ministry of Health and Welfare (grant number: HI17C0616).
Author contributions: This study was designed by J-KK, IK and SKY. The manuscript was written by J-KK and SKY. All experiments were planned and executed by J-KK, IK and SKY. Histological analysis was performed by KC and H-YL. Recombinant protein expression and purification were performed by EK and JKP. Plasmid constructions were performed by J-HC, H-YL and JKP.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information accompanies the paper on Experimental & Molecular Medicine website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Kim, JK., Kim, I., Choi, K. et al. Poly(rC) binding protein 2 acts as a negative regulator of IRES-mediated translation of Hr mRNA. Exp Mol Med 50, e441 (2018). https://doi.org/10.1038/emm.2017.262
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/emm.2017.262