Abstract
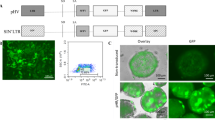
Phage φC31 integrase is a recombinase that can be expressed in mammalian cells to integrate plasmids carrying an attB sequence into the genome at specific pseudo attP locations. We show by immunofluoresence that wild-type φC31 integrase is cytoplasmic and that addition of the SV40 nuclear localization signal (NLS) localized φC31 integrase to the nucleus. Unexpectedly, the NLS depressed integration efficiency in HeLa cells and provided no benefit when used to integrate the human Factor IX (hFIX) gene into mouse liver. As breakdown of the nuclear membrane during mitosis could allow cytoplasmic integrase access to the chromosomes, we analyzed whether cell division was required for integration into liver cells in vivo. Hepatocytes were labeled with iododeoxyuridine to mark cells that underwent DNA replication during the week after hydrodynamic injection. Hydrodynamic delivery led to DNA replication in one-third of hepatocytes. Approximately three out of four cells having φC31 integrase-mediated stable hFIX expression did not undergo replication, indicating that cell division was not required for integrase function in liver. Therefore, although the bulk of φC31 integrase protein seems to be cytoplasmic in mammalian cells, integration can still occur in the nucleus, even without cell division.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Calos MP . The phiC31 integrase system for gene therapy. Curr Gene Ther 2006; 6: 633–645.
Chalberg TW, Genise HL, Vollrath D, Calos MP . phiC31 integrase confers genomic integration and long-term transgene expression in rat retina. Invest Ophthalmol Vis Sci 2005; 46: 2140–2146.
Ortiz-Urda S, Thyagarajan B, Keene DR, Lin Q, Calos MP, Khavari PA . PhiC31 integrase-mediated nonviral genetic correction of junctional epidermolysis bullosa. Hum Gene Ther 2003; 14: 923–928.
Olivares EC, Hollis RP, Chalberg TW, Meuse L, Kay MA, Calos MP . Site-specific genomic integration produces therapeutic Factor IX levels in mice. Nat Biotechnol 2002; 20: 1124–1128.
Bertoni C, Jarrahian S, Wheeler TM, Li Y, Oliveres EC, Calos MP et al. Enhancement of plasmid-mediated gene therapy for muscular dystrophy by directed plasmid integration. Proc Natl Acad Sci USA 2006; 103: 419–424.
Belteki G, Gertsenstein M, Ow DW, Nagy A . Site-specific cassette exchange and germline transmission with mouse ES cells expressing phiC31 integrase. Nat Biotechnol 2003; 21: 321–324.
Allen BG, Weeks DL . Transgenic Xenopus laevis embryos can be generated using phiC31 integrase. Nat Methods 2005; 2: 975–979.
Khan MS, Khalid AM, Malik KA . Phage phiC31 integrase: a new tool in plastid genome engineering. Trends Plant Sci 2005; 10: 1–3.
Bateman JR, Lee AM, Wu CT . Site-specific transformation of Drosophila via phiC31 integrase-mediated cassette exchange. Genetics 2006; 173: 769–777.
Raymond CS, Soriano P . High-efficiency FLP and PhiC31 site-specific recombination in mammalian cells. PLoS One 2007; 2: e162.
Kuhstoss S, Rao RN . Analysis of the integration function of the streptomycete bacteriophage phi C31. J Mol Biol 1991; 222: 897–908.
Rausch H, Lehmann M . Structural analysis of the actinophage phi C31 attachment site. Nucleic Acids Res 1991; 19: 5187–5189.
Groth AC, Olivares EC, Thyagarajan B, Calos MP . A phage integrase directs efficient site-specific integration in human cells. Proc Natl Acad Sci USA 2000; 97: 5995–6000.
Thorpe HM, Smith MC . In vitro site-specific integration of bacteriophage DNA catalyzed by a recombinase of the resolvase/invertase family. Proc Natl Acad Sci USA 1998; 95: 5505–5510.
Thyagarajan B, Oliveres EC, Hollis RP, Ginsburg DS, Calos MP . Site-specific genomic integration in mammalian cells mediated by phage phiC31 integrase. Mol Cell Biol 2001; 21: 3926–3934.
Andreas S, Schwenk F, Küter-Luks B, Faust N, Kühn R . Enhanced efficiency through nuclear localization signal fusion on phage PhiC31-integrase: activity comparison with Cre and FLPe recombinase in mammalian cells. Nucleic Acids Res 2002; 30: 2299–2306.
Chen JZ, Ji CN, Xu GL, Pang RY, Yao JH, Zhu HZ et al. DAXX interacts with phage PhiC31 integrase and inhibits recombination. Nucleic Acids Res 2006; 34: 6298–6304.
Liu F, Song Y, Liu D . Hydrodynamics-based transfection in animals by systemic administration of plasmid DNA. Gene Therapy 1999; 6: 1258–1266.
Zhang G, Budker V, Wolff JA . High levels of foreign gene expression in hepatocytes after tail vein injections of naked plasmid DNA. Hum Gene Ther 1999; 10: 1735–1737.
Rossmanith W, Chabicovsky M, Herkner K, Schulte-Hermann R . Cellular gene dose and kinetics of gene expression in mouse livers transfected by high-volume tail-vein injection of naked DNA. DNA Cell Biol 2002; 21: 847–853.
Keravala A, Lee S, Thyagarajan B, Olivares EC, Gabrovsky VE, Woodard LE et al. Mutational derivatives of PhiC31 integrase with increased efficiency and specificity. Mol Ther 2009; 17: 112–120.
Suda T, Gao X, Stolz DB, Liu D . Structural impact of hydrodynamic injection on mouse liver. Gene Therapy 2007; 14: 129–137.
Chavez CL, Keravala A, Woodard LE, Hillman RT, Chu J, Calos MP . Longevity of phiC31 integrase in the mouse liver. (in preparation).
Bonner WM . Proximity and accessibility studies of histones in nuclei and free nucleosomes. Nucleic Acids Res 1978; 5: 71–85.
Whittaker GR, Helenius A . Nuclear import and export of viruses and virus genomes. Virology 1998; 246: 1–23.
Gallay P, Hope T, Chin D, Trono D . HIV-1 infection of nondividing cells through the recognition of integrase by the importin/karyopherin pathway. Proc Natl Acad Sci USA 1997; 94: 9825–9830.
Yoshino H, Hashizume K, Kobayashi E . Naked plasmid DNA transfer to the porcine liver using rapid injection with large volume. Gene Therapy 2006; 13: 1696–1702.
Aliño SF, Herrero MJ, Noguera I, Dasí F, Sánchez M . Pig liver gene therapy by noninvasive interventionist catheterism. Gene Therapy 2007; 14: 334–343.
Lambotte L, Saliez A, Triest S, Tagliaferri EM, Barker AP, Baranski AG . Control of rate and extent of the proliferative response after partial hepatectomy. Am J Physiol 1997; 273: G905–G912.
Budker VG, Subbotin VM, Budker T, Sebestyén MG, Zhang G, Wolff JA . Mechanism of plasmid delivery by hydrodynamic tail vein injection. II. Morphological studies. J Gene Med 2006; 8: 874–888.
Khorsandi SE, Bachellier P, Weber JC, Greget M, Jaeck D, Zacharoulis D et al. Minimally invasive and selective hydrodynamic gene therapy of liver segments in the pig and human. Cancer Gene Ther 2008; 15: 225–230.
Fabre JW, Grehan A, Whitehorne M, Sawyer GJ, Dong X, Salehi S et al. Hydrodynamic gene delivery to the pig liver via an isolated segment of the inferior vena cava. Gene Ther 2008; 15: 452–462.
Schulte-Hermann R . Initiation and promotion in hepatocarcinogenesis. Arch Toxicol 1987; 60: 179–181.
Yeikilis R, Gal S, Kopeiko N, Paizi M, Pines M, Braet F et al. Hydrodynamics based transfection in normal and fibrotic rats. World J Gastroenterol 2006; 12: 6149–6155.
Keravala A, Portlock JL, Nash JA, Vitrant DG, Robbins PD, Calos MP . PhiC31 integrase mediates integration in cultured synovial cells and enhances gene expression in rabbit joints. J Gene Med 2006; 8: 1008–1017.
Portlock JL, Keravala A, Bertoni C, Lee S, Rando TA, Calos MP . Long-term increase in mVEGF164 in mouse hindlimb muscle mediated by phage phiC31 integrase after nonviral DNA delivery. Hum Gene Ther 2006; 17: 871–876.
Acknowledgements
Many thanks to Julien Sage and members of his laboratory at Stanford for assistance with the liver-staining experiments. Thanks to Anne Brunet and members of her laboratory for assistance with their microscope. We appreciate the contributions of Christopher L Chavez to the conclusions made in this paper. PHS Grant Number CA09302, awarded by the National Cancer Institute, DHHS, supported LEW. RTH was supported by a medical research training fellowship from the Howard Hughes Medical Institute. This work was supported by NIH grant HL068112 to MPC.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Woodard, L., Hillman, R., Keravala, A. et al. Effect of nuclear localization and hydrodynamic delivery-induced cell division on φC31 integrase activity. Gene Ther 17, 217–226 (2010). https://doi.org/10.1038/gt.2009.136
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gt.2009.136
Keywords
This article is cited by
-
Global mapping of binding sites for phic31 integrase in transgenic maden-darby bovine kidney cells using ChIP-seq
Hereditas (2019)
-
Kidney-specific transposon-mediated gene transfer in vivo
Scientific Reports (2017)
-
Ultrasound-targeted hepatic delivery of factor IX in hemophiliac mice
Gene Therapy (2016)
-
Improved site-specific recombinase-based method to produce selectable marker- and vector-backbone-free transgenic cells
Scientific Reports (2014)